| Research Article | ||
Open Vet. J.. 2026; 16(2): 1124-1140 Open Veterinary Journal, (2026), Vol. 16(2): 1124-1140 Research Article Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injuryRafael Enrique Castro-Vargas1,2, María Paula Herrera-Sánchez1,2and Iang Rondón-Barragán1,2*1Research Group of Immunobiology and Pathogenesis, Faculty of Veterinary Medicine and Zootechnics, University of Tolima, Ibagué, Colombia 2Research Group of Poultry Science, Faculty of Veterinary Medicine and Zootechnics, University of Tolima, Ibagué, Colombia *Corresponding Author: Iang Rondón-Barragán. Research Group of Immunobiology and Pathogenesis, Faculty of Veterinary Medicine and Zootechnics, University of Tolima, Ibagué, Colombia. Email: isrondon [at] ut.edu.co Submitted: 19/08/2025 Revised: 14/01/2026 Accepted: 30/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
ABSTRACTBackground: Neuroinflammation can be triggered by traumatic brain injury (TBI) and exposure to toxicants such as chlorpyrifos (CPF), leading to chronic disabilities and substantial health care costs. CD59 and glial maturation factor-beta proteins are implicated in various brain regulatory mechanisms. Investigating their roles can provide valuable insights into their role in pathologies such as TBI. Aim: This study aimed to analyze the molecular characteristics of the CD59 and gmfb genes and assess their relative gene expression in brain tissues of Piaractus orinoquensis subjected to brain injury using quantitative PCR (qPCR). Methods: Thirty P. orinoquensis species were allocated to brain injury or CFP exposure models. Brain injury was induced by telencephalic stab puncture, and CPF exposure was administered at a sublethal concentration of 0.011 µg/l. Brain regions, including the olfactory bulb, telencephalon, and optic chiasm, were collected at designated time points, and qPCR analysis was performed. CD59 and gmfb genes were sequenced and characterized using bioinformatic tools. Results: Both genes exhibited conserved features compared with other fish species. In the brain injury model, fish subjected to brain injury showed reduced expression of CD59 and gmfb compared with controls, indicating a loss of immunomodulation associated with brain damage. Conversely, in the CPF-treated group, increased expression of CD59 in the olfactory bulb and gmfb in the optic chiasm was observed. Conclusion: Downregulation of CD59 after brain injury suggests a loss of immunomodulatory capacity following brain damage. In CPF-exposed fish, the overexpression of both CD59 and gmfb in some brain tissues demonstrated the toxicant’s ability to generate region-specific immunoactivity and neurotoxicity in the brain. Keywords: Chlorpyrifos, QPCR, Red–bellied pacu, Traumatic brain injury. IntroductionNeuroinflammation refers to the inflammatory response that occurs within the nervous system in response to injury, infection, toxicosis, or trauma (Shabab et al., 2017; Fan et al., 2018). Traumatic brain injury (TBI) is one of the leading causes of neuroinflammation, representing a significant public health challenge worldwide, as it is a major cause of both mortality and disability (Xiong et al., 2019). In 2016, approximately 27 million cases of TBI were reported globally, resulting in chronic disabilities and substantial healthcare costs (James et al., 2019). Beyond mechanical trauma, exposure to xenobiotics, such as herbicides and insecticides, has also been identified as a public health concern with systemic effects, including neuronal damage (Richardson et al., 2019). Chlorpyrifos (CPF), an organophosphate insecticide widely used to control foliar insects in crops, is among the most notable of these xenobiotics (Duarte-Restrepo et al., 2020). The neuroinflammatory response in TBI involves glial, neuronal, inflammatory, and endothelial cells, complicating the understanding of TBI pathophysiology and contributing to secondary damage sites (Ahmed et al., 2019). This secondary damage can lead to demyelination, axonal degeneration, cell death, and the formation of glial scars surrounding the initial injury site (DiSabato et al., 2016; Ahmed et al., 2019). Similarly, exposure to CPF can also induce neuropathological outcomes. CPF's toxic effects are primarily mediated through its oxon metabolite, which inhibits acetylcholinesterase in the nervous system, leading to the accumulation of acetylcholine and subsequent cholinergic hyperstimulation and neuronal damage (Naime et al., 2020). Beyond acute toxicity, CPF has been linked to axonal and myelin degeneration in both the central and peripheral nervous systems, a condition known as organophosphate-induced delayed neuropathy in humans (Elmorsy et al., 2022). CPF exposure has also been implicated in developmental neurotoxicity, resulting in behavioral and physiological impairments in both young animals and children, as evidenced in animal models, such as the guppy Poecilia reticulata and zebrafish Danio rerio (Sledge et al., 2011). In Colombia, Piaractus orinoquensis (red-bellied pacu) has been used in pharmacological and toxicological studies as a biological model (Zapata-Guerra et al., 2020; Petano-Duque et al., 2022; Cruz-Méndez et al. 2022; Ortiz-Muñoz et al. 2023; Rueda-García and Rondón- Barragán 2024), animal breeding (Collazos-Lasso et al., 2021; Medina-Robles et al., 2023), and phenotypically characterized (Cuan-Barrera et al., 2021; Escobar et al., 2022). This species is economically important due to its significant commercial production in countries such as Colombia, Venezuela, Peru, and Bolivia, as well as its remarkable adaptability, resilience, and rapid growth, characterized by a high feed conversion rate (Escobar et al., 2022). Piaractus orinoquensis represents a potential toxicological and neurobiological model for South America. Several molecules, including cytokines, chemokines, and other inflammatory mediators, intervene in the inflammatory response of the brain. The complement system plays a pivotal role in activating pathways that lead to the formation of the membrane attack complex (MAC, C5b-9) (Kim and Suk, 2023). Despite their significance in TBI, these pathways have received limited attention, even though they are involved in the direct lysis of target surfaces, which may potentially promote neurogenesis and plasticity following brain injury (Hammad et al., 2018; Toutonji et al., 2021; van Erp et al., 2023). Although essential for immune defense, overexpression of the complement system may lead to excessive inflammation in the brain. To prevent such damage, a group of regulatory proteins tightly regulates its activity (Dinet et al., 2019). Among them, CD59 is a key regulatory protein that prevents the formation of MAC by inhibiting the binding of C9 to the C5b-8 complex, thereby protecting cells from autologous complement attack and preventing the unnecessary consumption of complement components (Erwood and Alawieh, 2022; Couves et al. 2023; Patel et al., 2024). CD59 is distributed across the surface of hematopoietic and tissue cells, where it is anchored to the cell membrane via glycosylphosphatidylinositol (Du et al., 2014; Jia et al., 2019). CD59 plays a key role in regulating Schwann cell (SC) proliferation during nervous system development in zebrafish, particularly by protecting group with SC from inflammation-induced overproliferation (Wiltbank et al., 2022). Similarly, glial maturation factor-beta (GMFB) plays a critical role in modulating immune responses within the brain. GMFB is predominantly expressed in the central nervous system, where it facilitates the differentiation of brain cells, such as neurons and glia, and mediates neuroinflammation in neurodegenerative diseases (Fan et al., 2018; Ahmed et al., 2019). Additionally, GMFB is associated with neurotropic functions through its activity as a signal transducer, and its expression has been observed in various other tissues, including the colon, thymus, and kidney (Hakala, 2020). For instance, in murine and zebrafish (Danio rerio) models, both widely used to study TBI, elevated GMFB expression has been observed in astrocytes and microglia at the injury site (Yin et al., 2018). This suggests that GMFB is involved in reactive gliosis following brain trauma and may represent a novel factor in the regulation of injury-induced gliosis (Yin et al., 2018). The objective of the present study was to analyze the molecular characteristics of the CD59 and gmfb genes and evaluate their relative expression by quantitative polymerase chain reaction (qPCR) in the brain tissues of P. orinoquensis subjected to various brain injury conditions. Materials and MethodsAnimalsA total of 25 healthy red-bellied pacu (P. orinoquensis) of both sexes and from the same spawning, with an average body weight of 5 ± 0.3 g, were used for all the experiments. The fish were treated with 1% NaCl to eliminate the ectoparasitic load and acclimated for 15 days (Petano-Duque et al., 2022). All fish were kept in a 90-l glass aquarium after random group selection. Conditions were regulated at a temperature of 25°C, with constant aeration and no filtration, a density of less than 1 g/l, and a 12-hour light/dark cycle. Feed was provided twice a day with a commercial diet (30% protein) equivalent to 2% of their body weight. Feed was deprived 24 hours before the acute and subacute brain injury experiments. In addition, tissues, such as the liver, kidney, gill, spleen, heart, intestine, stomach, blood, and brain segments of the olfactory bulb, optic chiasm, and telencephalon, were snap-frozen in liquid nitrogen until further analysis. Acute and subacute brain injuryAcute and subacute brain injuries were performed using the brain puncture technique. Fifteen animals were used and distributed as follows: 3 euthanized at time 0 (group I, control group), 3 euthanized at 2 hours post-lesion (group II), 3 euthanized at 1 day post-lesion (dpl) (group III), 3 euthanized at 7 dpl (group IV), and 3 euthanized at 14 dpl (group V). After a 15-day acclimation period, all fish were anesthetized using menthol (50 mg/l) according to the recommendations of Zapata-Guerra et al. (2020). When fish showed total loss of the swimming axis (induction stage III) (Millán-Ocampo et al., 2012), brain injury was performed according to the stab wound procedure with a frontal puncture in the left dorsolateral domain of the telencephalon using a sterilized 000-gauge needle, based on Kishimoto et al. (2012) and Schmidt et al. (2014). The frontal puncture was on a projected straight line from the lateral edge of each eye to the fish’s dorsal edge. The needle was firmly inserted 5 mm deep at a 45° angle. After the puncture, the fish were placed in a freshwater recovery tank without anesthesia for 2 hours. All animals were euthanized by decapitation, according to the protocol of the American Veterinary Medical American Veterinary Medical Association (2020). The control group was treated with the same handling procedure and anesthesia without puncture and was euthanized after sedation. Brain tissue sampling was similar to that described above. Sublethal exposure to CPFThe fish were divided into a control group (n=5, 0 µg/l of CPF) and a CPF group (n=5, 0.011 µg/l of CPF). The sublethal concentration of CPF was determined according to the findings of Holguín-Céspedes et al. (2019) in P. orinoquensis fingerlings. The CPF concentration was maintained by replacing 50% of the water daily and adding CPF. After 72 hours of exposure, the fish were anesthetized using the hypothermic method and euthanized by cervical dislocation (American Veterinary Medical Association, 2020). Brain tissue (olfactory bulb, optic chiasm, and telencephalon) were collected and stored in liquid nitrogen until further use. RNA extraction and synthesis of cDNARNA-Solv® (Omega BIO-TEK, GA, USA) was used for Ribonucleic acid (RNA) extraction, and the quality and concentration were assessed by Varioskan™ LUX multimode microplate reader (Thermo Fisher Scientific, MA, USA). Complementary DNA (cDNA) was synthesized from the liver, kidney, gill, spleen, heart, intestine, stomach, blood, and brain segments of the olfactory bulb, optic chiasm, and telencephalon using the GoScript Reverse Transcription System (Promega, MA, USA) according to the manufacturer’s protocol. CD59 and gmfb cDNA sequencingcDNA from the brain tissue of red-bellied pacu was used for nanosequencing using the MinION (Oxford Nanopore Technologies, OXF, UK). CD59 and gmfb cDNA sequences were obtained by mapping the contigs on the Pygocentrus nattereri and Colossoma macropomum CD59 (XM_017722929.1, XM_036584013.1) and gmfb (XM_017682432.2, XM_036575262.1) sequences. Primers were designed to amplify the entire open reading frame (ORF) of each gene. After the sequences were confirmed by Sanger sequencing (Macrogen, Seoul, South Korea), qPCR primers were designed using Geneious Prime software v2025.1.3 (Biomatters Ltd, AK, NZ) (Table 1). Sequences were submitted to GenBank with the corresponding accession numbers. Bioinformatic analysisThe main physical and chemical parameters of CD59 and gmfb ORF were determined using ProtParam (Gasteiger et al., 2005). The putative signal peptide was predicted using SignalP 5.0 (Almagro-Armenteros et al., 2019). Protein domains were predicted using PredictProtein (Bernhofer et al., 2021), InterProScan (Blum et al., 2021), and the NCBI conserved domains tool (Lu et al., 2020). Glycosylations were predicted using NetOGlyc-4.0 (Steentoft et al., 2013) and NetNGlyc-1.0 (Gupta and Brunak, 2002) prediction servers. The DiANNA 1.1 web server was used to predict disulfide connectivity (Ferrè and Clote, 2005), and NetPhos-3.1 was used to predict generic phosphorylations (Blom et al., 1999). The big-PI Predictor was used to predict CD59 GPI-anchoring sites (Eisenhaber et al., 1999). Multiple sequence alignments of amino acid sequences and identity values were generated using Geneious Prime version 2025.1.3 (Biomatters Ltd, AK, NZ). The neighbor-joining phylogenetic tree was constructed from GenBank by selecting orthologous full-length sequences with complete ORFs from teleost fish that are relatively closely related, using 1,000,000 bootstrap replications to increase confidence that the grouping is correctly inferred (Petano-Duque et al., 2022). Partial or predicted sequences were excluded. Multiple sequence alignment was performed using the MUSCLE algorithm with default gap-opening and gap-extension penalties. The alignments were checked to ensure domain integrity, and poorly aligned regions were trimmed. The SWISS-MODEL online platform (Waterhouse et al., 2018) and PyMol v1.2r3pre (Schrödinger, NY, USA) were used to predict protein 3D structures. Table 1. Primer sequences used for polymerase chain reaction and quantitative polymerase chain reaction analysis. CD59: membrane attack complex-inhibitory protein, gmfb: Glia maturation factor-β, ef1α: Elongation factor 1-alpha (Zapata-Guerra et al., 2020).
PCRcDNA was used as a template for gene amplification in the liver, kidney, gill, spleen, heart, intestine, stomach, blood, and whole brain using specific primer sets from qPCR (Table 1). PCR reaction was assessed in a total volume of 25 μl, containing 1 μl of the cDNA template, 1 μl of each primer (10 pmol/μl) (Macrogen, Korea), 0.125 μl of Gotaq DNA polymerase (Promega, WI, USA), 1 μl of MgCl2, 1 μl of dNTPs (Thermo Fisher Scientific, MA, USA), 5 μl of buffer, and 14.875 μl of nuclease-free water (ddH2O) for each sample. The PCR was run in a ProFlex PCR System Thermocycler (Applied Biosystems, MA, USA) with an initial denaturation step of 3 minutes at 95°C, followed by 35 cycles as follows: 30 seconds at 95°C for denaturation, 30 seconds at 55°C for annealing, 30 seconds at 72°C for extension, and a final extension step of 5 minutes at 72°C. Amplification products were visualized by horizontal electrophoresis using PowerPac HC (BIO-RAD, CA, USA) on 2% agarose gel stained with HydraGreenTM (ACTGene, NJ, USA). The 100-bp Opti-DNA Marker (ABM Industries Inc., CA, USA) was used as a molecular weight marker. The gel was visualized and documented using the ENDURO GDSTM gel documentation system (Labnet International, NJ, USA). qPCRQuantitative polymerase chain reaction assays were performed in a Quant StudioTM 3 Real-Time PCR System (Thermo Fisher Scientific, MA, USA) to determine the relative gene expression in the brain regions of the telencephalon, optic chiasm, and olfactory bulb from the control, injured, and CPF-exposed groups. The qPCR reaction was carried out in a total volume of 20 μl, composed of 7 μl of ddH2O, 0,5 μl of each primer (10 pmol/μl) (Macrogen, Korea) (Table 1), 2 μl of cDNA template, and 10 μl of Luna® Universal qPCR Master Mix (New England Biolabs, MA, USA) in fast ramp mode. Relative gene expression was calculated using the 2−ΔΔCt method (Livak and Schmittgen, 2001). The primer products were validated by melt-curve analysis over the temperature range of 70°C–95°C. EF1a was used as a normalization gene, and the data were expressed as fold change. Statistical analysisData were analyzed using descriptive statistics, and normality was assessed using the Shapiro-Wilk test. Differences in gene expression between treatments were evaluated using either the T-test or the Mann-Whitney test. Statistical analyses were performed using GraphPad Prism v 8.0 (La Jolla, CA, USA). The probability levels for statistical significance were set at *p< 0.05,**p< 0.01,***p < 0.001, and ****p < 0.0001. Ethical approvalAll experiments were conducted in accordance with the ethical guidelines established by the Bioethics Committee of the Central Office of Research at the Universidad del Tolima, as recorded in minutes number 2 of 2025, and in compliance with Law 84 of 1989 and Resolution 8430 of 1993. Additionally, this study adhered to the recommendations of the Guide for the Care and Use of Laboratory Animals (Jenkins et al., 2014). ResultsProtein characterization and phylogenetic analysisThe full ORF of CD59 and gmfb genes from P. orinoquensis had a length of 351 and 429 bp, respectively. They encoded proteins with 116 and 142 amino acids (aa), respectively, with predicted molecular weights of 12.7 and 16.8 kDa and theoretical isoelectric points of 6.50 and 5.05. The highest pairwise identity value from CD59 amino acid sequences was found in CD59 glycoprotein from C. macropomum (94.07%) (XP_036439906.1) and Pygocentrus nattereri (91.53%) (XP_017565112.1). The highest pairwise identity value from GMFB amino acid sequences was found in GMFB from P. nattereri (100%) (XP_017537921.1) and Astyanax mexicanus (99.3%) (XP_007247651.1). The main structural characteristics of CD59 were identified, including two polyadenylation signals (AATAA) at 15 and 412 bp and a TATA box (TATAAA) at position 219 downstream of the translation termination codon (TGA) in the 5' and 3' UTRs. In the case of the gmfb sequence, no polyadenylation signals were found, but the TATA box was present downstream of the TGA at position 220. The CD59 putative signal peptide was identified in amino acid positions 1–21 with a predictive cleavage site between Ala21 and Ile22. Post-translational modifications, such as N-glycosylation and O-glycosylation, were absent in both CD59 and GMFB, but 18 potential phosphorylation sites were found in CD59 at Tyr30 and 68, Thr31, 37, 48, 56, 82, 96, and 97, Ser33, 45, 64, 72, 80, 88, 95, 99, and 100 that PK-A, G, and C, CK-I and II, DNAPK, ATM, cdc2, and INSR can phosphorylate. In the case of GMFB, ten potential phosphorylation sites at Ser2, 53, 72, 83, 92, and 107 were identified, as well as four sites at Thr27, 114, 126, and 130 that can be phosphorylated by PK-A, B, C, and G, RSK, CK-I, and II, respectively. The CD59 putative GPI-anchoring site was identified at Asn92. The degree of conservation of the red-bellied pacu CD59 amino acid sequence to aligned fish species varied between 40.68% and 94.07%. CD59 protein consists of 2 domains, the signal peptide and the Ly-6 antigen/uPA receptor-like domain from the lymphocyte antigen 6 (Ly-6) family. Consensus sequence motif CCXXXXCN between 86 and 93 positions in all 19 aligned sequences. The mature red-bellied pacu CD59 protein contained 11 cysteine residues, with a high level of conservation in most aligned fish species (13/19 sequences) (Fig. 1). Cysteine residues in red-bellied pacu CD59 might form 5 disulfide bonds at the amino acid position 24–27, 34–46, 40–114, 66–85, and 86–91, which presented five antiparallel beta-sheets highly common in the three-finger protein family. The predicted 3D protein structure from human CD59 showed that protein identity was 29.33%, and 9 of 11 conserved cysteines took the same position (Fig. 2). GMFB amino acid conservation between fish species ranged between 94.3% and 99.3%. The protein was composed of one domain from the actin-depolymerizing factor between amino acid positions 10–129 from the actin-depolymerizing factor-gelsolin superfamily. Signal peptide and conserved motifs were absent in the amino acid sequence, and 3 well-conserved cysteine residues were found in all 21 fish sequences, forming one disulfide bridge at positions 87 and 96 (Fig. 3). The 3D structure of GMFB was designed based on the GMFB protein model of Mus musculus with a protein identity of 80.99% (Fig. 4).
Fig. 1. Multiple alignment of fish CD59 aa sequences. Signal peptide (orange line) and Ly-6 antigen/uPA receptor-like domain (blue line) expression. Cysteine residues are highlighted in purple, and a red rectangular line indicates the conserved motif. GPI-anchoring sites are indicated in the background by a yellow oval.
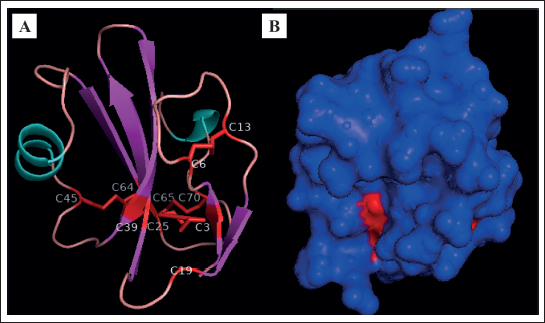
Fig. 2. Protein modeling of CD59 of red-bellied pacu. (A) a-helix residues are blue, b-sheet residues are violet, loop and unassigned residues are pink, and cysteines and disulfide bonds are red. (B) CD59 protein surface. The protein model was based on human CD59 (PDB ID: P13987–2j8b.1.A).
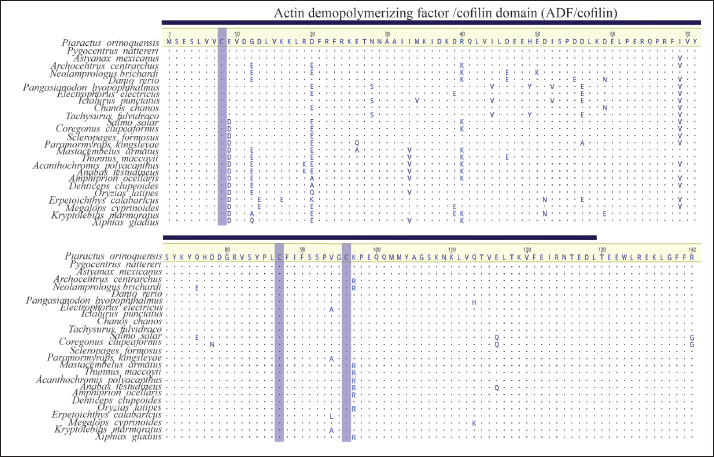
Fig. 3. Multiple alignment of GMFB aa sequences of fishes. Actin-DPF domain (blue line). Cysteine residues are highlighted in purple.
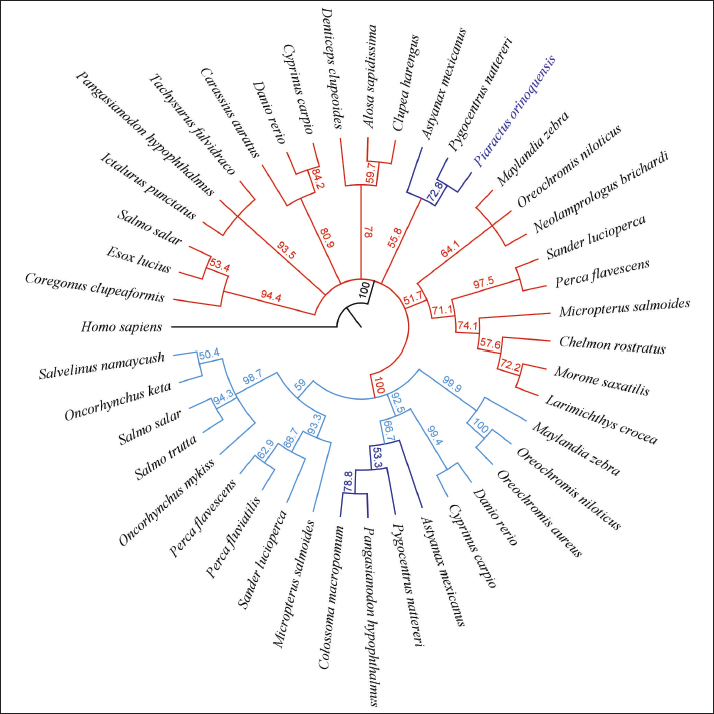
Fig. 4. Protein modeling of the GMFB of red-bellied pacu. (A) a-helix residues are blue, b-sheet residues are violet, loop and unassigned residues are pink, and cysteines are red. (B) Protein surface of GMFB. The protein model was based on murine GMFG (PDB ID: Q9ERL7–1vkk 1.A-). The phylogenetic tree of CD59 was constructed using 42 sequences and clearly separated fish from mammals, avians, and reptiles. Two main clusters were formed in fish, comprising representative orders such as Cyprinodontiformes and Perciformes (99.9%); the other cluster included representative orders such as Perciformes, Gasterosteiformes, Clupeiformes, Cypriniformes, and Characiformes (67%). Characiformes were supported by 95.3%, with C. macropomum and P. orinoquensis CD59 clustering together at 78.3% (Fig. 5). The glial maturation factors β and γ were compared. Marked segregation was observed in all GMFG sequences from teleost fish, with 100% bootstrap support, compared with the GMFB sequences. In the GMFB tree segment (light blue lines), six branches were formed with representative families of Salmonidae with support of 94.4%, Siluriformes order (93.5%), Cyprinidae family (80.9%), Clupeidae family (78%), Characidae family (55.8%), and Perciformes order (51.7%) (Fig. 6). In the GMFG tree (red lines), three main branches were formed with representative families of Salmonidae and Percidae (59%), Characidae and Cyprinidae (92.5%), and Cichlidae (99.9%). Within the Characidae family, GMFB protein sequences were supported in a branch with a bootstrap value of 55.8%, and GMFG protein was supported in a branch with a bootstrap value of 66.7%. Piaractus orinoquensis shared a cluster with P. nattereri (72.8%).
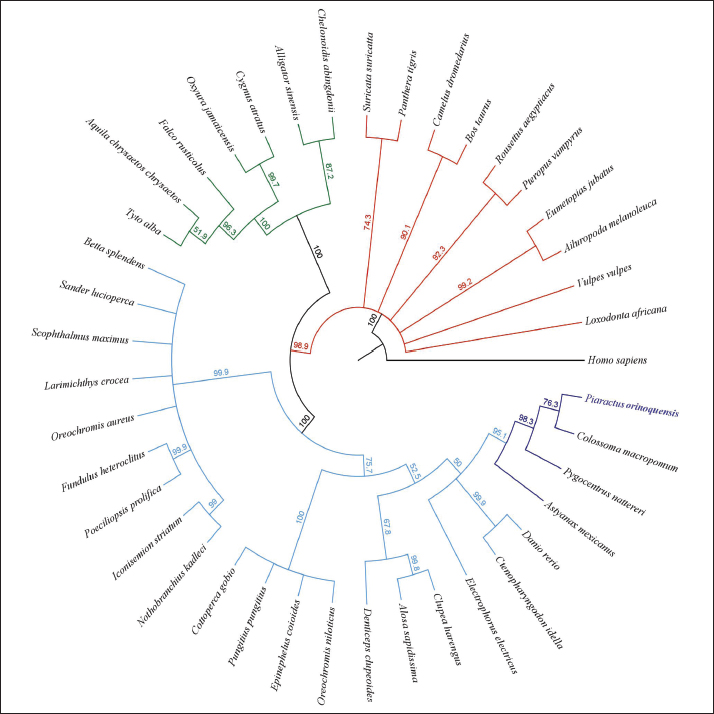
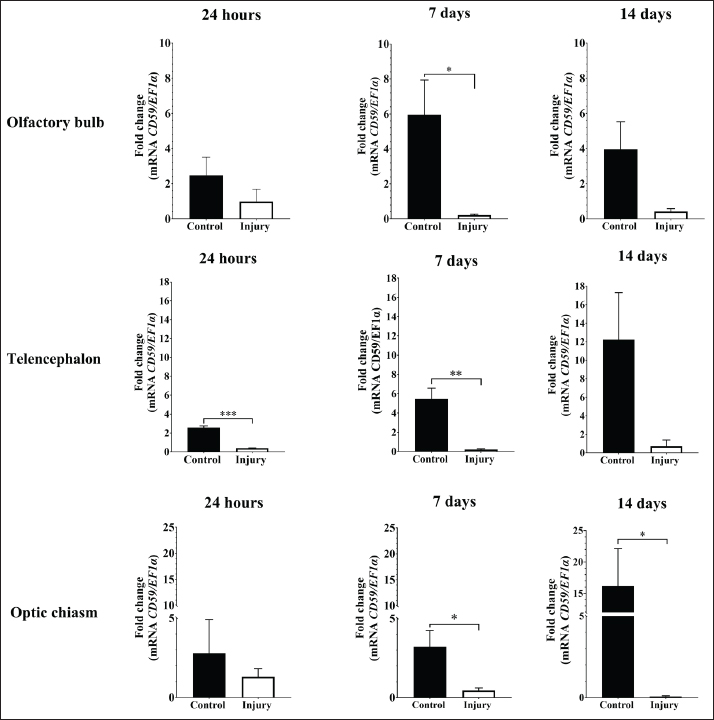
Fig. 5. Phylogenetic tree of CD59 generated by the neighbor-joining method. A total of 41 sequences were grouped with a bootstrap of 1.000.000 replicates. Gene PCR amplificationThe PCR-based profile revealed a 121-bp band of CD59 in the brain, blood, kidney, liver, gills, spleen, heart, and stomach, with no amplification in the intestine (Fig. 7A). Consistent bands of 135 bp were observed in the brain, blood, kidney, gills, intestine, heart, and stomach in gmfb. Notably, no detectable bands were detected in the liver or spleen (Fig. 7B). CD59 and gmfb gene expression in two experimental modelsIn the acute brain injury at 2 hours, the relative expression of CD59 was significantly lower in the brain-injured group compared to the control group (p < 0.0001). Conversely, gmfb expression was significantly elevated in the brain-injured group relative to control (p < 0.01) (Fig. 8). In the case of the CD59 gene in brain segments, an increased mRNA level was found in the olfactory bulb of the control group at 7 days post-lesion (dpl) (p < 0.05). Similarly, the telencephalon showed high expression of the CD59 gene in the control group compared with the injury group, with significant differences at 24 hours post-lesion (p < 0.001) and 7 dpl (p < 0.01). Moreover, in the optic chiasm, a statistical difference was observed in CD59 transcripts in the control group at 7 and 14 dpl (p < 0.05) (Fig. 9).
Fig. 6. GMFB phylogenetic tree generated by the neighbor-joining method. A total of 48 sequences were grouped with a bootstrap of 1.000.000 replicates.
Fig. 7. PCR of CD59 (A) and gmfb (B) in 9 tissues from the red-bellied pacu, Piaractus orinoquensis. Lane M: molecular weight marker, B: brain, Bl: blood, K: kidney, L: liver, G: gill, S: spleen, H: heart, St: stomach, and I: intestine.
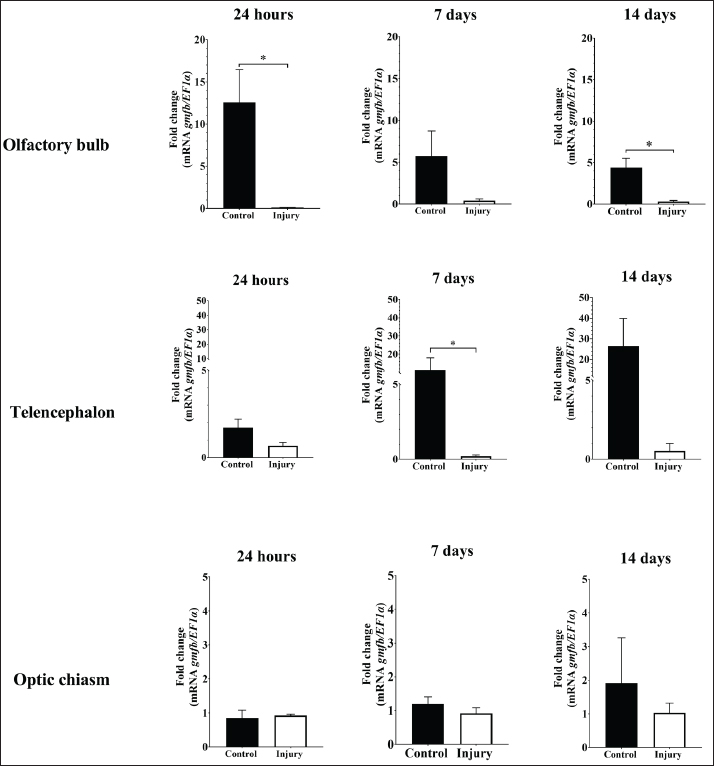
Fig. 8. Relative gene expression of CD59 and gmfb on acute brain injury (2 hours) in the red-bellied pacu, Piaractus orinoquensis. **p < 0.01, ****p < 0.0001. EF1a was used as the reference gene. In the olfactory bulb, gmfb gene expression was significantly higher in the control group than in the injury group at 24 hours post-lesion and 14 days post-lesion (dpl) (p < 0.05). Similarly, transcript levels were significantly elevated in the telencephalon of the control group relative to the injury group at 7 dpl (p < 0.05). In contrast, gmfb mRNA levels in the optic chiasm did not differ significantly between the groups (Fig. 10). In the CPF treatment experiment, the relative expression of CD59 was significantly increased only in the olfactory bulb of the treated group (p < 0.01). In addition, gmfb expression differed significantly only in the optic chiasm of the treated group (p < 0.05) (Fig. 11). DiscussionHomology, structural characteristics, and 3D protein modelThe predicted molecular weight of CD59 was lower (12.7 kDa) than that reported in channel catfish (Yeh and Klesius, 2007). However, the structural characteristics of CD59, such as two polyadenylation, a signal peptide of 21 amino acids at the N-terminus, a domain of 72 amino acids with a CCXXXXCN motif, at least 10 cysteines involved in disulfide bond formation, and a GPI-anchoring peptide at the C-terminus, which had the principal function as membrane attachment, were shared with zebrafish, rainbow trout, large yellow croaker, and Nile tilapia (Ploug and Ellis, 1994; Liu et al., 2007;Papanastasiou et al. 2007;Sun et al., 2013; Gan et al., 2015). The alignment of CD59 proteins across different fish orders shows high divergence. However, most sequences are highly conserved in both the number and positions of cysteine residues. Regarding the protein model, although the overall sequence homology between CD59 from human and red-bellied pacu was low (29.33%). Currently available models exhibit a low degree of identity, which may limit the interpretation of the 3D protein structure. However, despite the low identity, our modeled protein presented the main characteristics of this protein family, such as the number and distribution of cysteine residues (9 of 10) were present, with the typical three-finger structure, which includes a two-stranded β-sheet finger, a core of three-stranded β-sheet finger, and a short helix, demonstrating the high structural conservation between vertebrates despite the divergence between sequences (Yeh and Klesius, 2007; Sun et al., 2013). Topologically, the mature CD59 protein shares an evolutionary relationship with the Ly6 superfamily, snake venom neurotoxins, Ly-6 alloantigens, and the urokinase-type plasminogen activator receptor (Ploug and Ellis, 1994; Chang et al., 2002).
Fig. 9. Relative gene expression of CD59 in the olfactory bulb, telencephalon, and optic chiasm of the red-bellied pacu (Piaractus orinoquensis) at 24 hours, 7 days, and 14 days after brain injury. *p < 0.05, **p < 0.01, ***p < 0.001. EF1a was used as the reference gene. In GMFB, the molecular weight (16.8 kDa) and amino acid length (142 aa) were close to the molecular weight of human GMFB (17 kDa) (Lim et al., 1989; Shi et al., 2006). The absence of a signal peptide in GMFB is due to its intracellular protein condition, which prevents secretion without a signal sequence (Lim and Zaheer, 1995). The absence of N-glycosylation was similar to that reported by Kaplan et al. (1991) and several serine/threonine phosphorylation sites and phosphorylated proteins in GMFB were identical to those reported in human GMFB (Lim and Zaheer, 1995). The small number of mRNA functions found in gmfb was associated with the short 3 and 5' UTR length. The multialignment demonstrated high evolutionary conservation among species in GMFB, which can be explained by the high conservation of all members of the ADF/cofilin superfamily, which have evolved from a single ancestral protein as well as a similar structure composed of five internal β-strands, four of them with an antiparallel constitution, and most of them are surrounded by at least four helices (Lappalainen et al., 1998; Goroncy et al., 2009). The phylogenetic analysis presented a wide CD59 protein cluster among teleosts, as reported by Papanastasiou et al. (2007) in rainbow trout, Gan et al. (2015) in Nile tilapia, Yeh and Klesius (2007) in channel catfish, and Sun et al. (2013) in zebrafish.
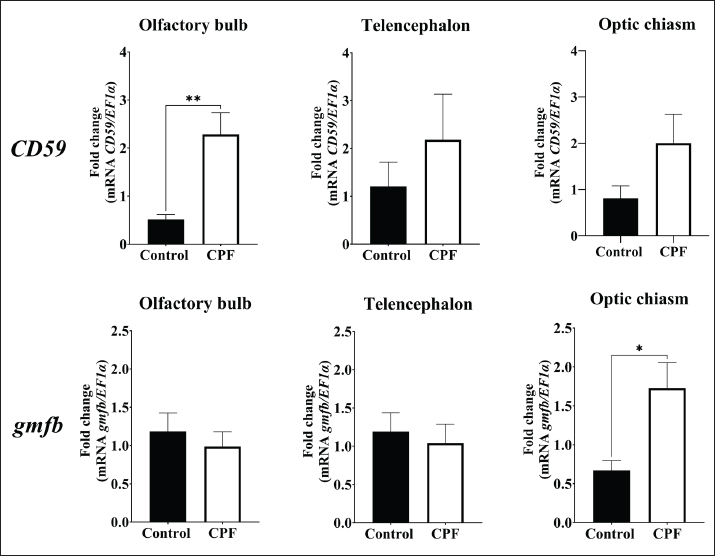
Fig. 10. Relative gene expression of gmfb in the olfactory bulb, telencephalon, and optic chiasm of the red-bellied pacu (Piaractus orinoquensis) at 24 h, 7 days, and 14 days after brain injury. *p<0.05. EF1a was used as the reference gene. Gene detectionThe stab wound procedure in the telencephalon mimics the cellular and structural changes associated with TBI caused by a mechanical impact and penetrating trauma, affecting multiple structures such as neurons, blood vessels, and glial cells (Kishimoto et al., 2012; Algattas and Huang, 2013). However, numerous authors have reported the telencephalic ventricular zone as a well-defined adult neurogenic region in vertebrates, including reptiles, birds, and fish (Doetsch and Scharff, 2001; Font et al., 2001; Zupanc, 2008). In our study, the CD59 gene was detected in the brain, blood, kidney, liver, gills, spleen, heart, and stomach, but not in the intestine, which is consistent with previous reports in the large yellow croaker (Liu et al., 2007; Sun et al., 2013). Although gmfb is expressed mainly in the central nervous system, it was present in the brain, blood, kidney, gills, and intestine in the present study. In other studies, gmfb expression has been detected in the colon, thymus, testis, ovary, and proximal renal tubule (Inagaki et al., 2004; Hakala, 2020). This overall expression can be related to other hypothesized functions of gmfb, such as erythropoietin-inducer and granulocyte-colony stimulator, and in CD4+ T cell maturation in the thymus (Utsuyama et al., 2003; Shi et al., 2006).
Fig. 11. Relative gene expression of CD59 and gmfb in the red-bellied pacu, Piaractus orinoquensis, after exposure to chlorpyrifos (0.011 μg/L). *p<0.05, **p<0.01. EF1a was used as the reference gene. qPCR experimentsA remarkable finding was observed in all brain tissue evaluated, where the injured group showed lower CD59 expression than the control group. The reason for the downregulation can be associated with autologous complement system immunomodulation in the brain, considering that most of the CD59 is concentrated in the oligodendrocytes by a GPI anchor due to its highly vulnerable condition to multiple disturbances of brain homeostasis (Scolding et al., 1998; Sun et al., 2013; Zeis et al., 2016). However, in the context of brain homeostasis disruption in TBI, we hypothesize that the loss of CD59 regulation promotes complement molecule overexpression. This hypothesis is supported by findings in human patients with acute and early chronic TBI, where significantly lower CD59 expression is observed compared with the control group and in patients with TBI (Bellander et al., 2001). Additionally, elevated levels of complement components, including C1q, C3b, C3d, and MAC, have been observed in patients with TBI compared with controls (Goetzl et al., 2020). In addition, MAC has been elucidated in secondary injury after TBI. In knockout CD59 mice, MAC deposition was higher than in the control group, resulting in increased neuronal loss and worse neurological outcomes at 7 days after trauma (Stahel et al., 2009). In the present study, low CD59 expression in the acute brain-injured group at 2 hours, 1, 7, and 14 days may be associated with the temporal persistence of CD59 deregulation. Regarding gmfb although the expression was elevated just after 2 hours post-lesion, the expression in the segments was lower days later than in the control group. This discrepancy indicates that the early upregulation is transient and downregulates rapidly or that it occurred in regions that were not sampled. Despite their beneficial role in brain recovery, Ahmed et al. (2019) have reported that neuroinflammation mediated by GMF can exacerbate neuronal injury through neuroinflammation, ROS generation, and oxidative stress damage in the TBI in vitro scratch-injury model. GMF KO mice showed a significant reduction in glial fibrillary acidic protein expression, a biomarker associated with axonal injury. This reduces neuroinflammation, reverses neuropathological events, and improves behavioral outcomes (Selvakumar et al., 2020). After the injury, the telencephalon, optic chiasm, and olfactory bulb in the control group showed higher expression than those in the injury group, contrary to the report by Yin et al. (2018) in the brain of adult zebrafish, where elevated gmfb mRNA levels were found. However, this difference may be related to the brain regions and the selected periods in our study. In CPF-exposed fish, the early expression of gmfb and CD59 in the optic chiasm and olfactory bulb, respectively, may suggest CPF-induced neurotoxic effects. Moreover, these anatomical biases might indicate a preferential localized immune response or vulnerability to this organophosphate. CPF can induce oxidative cellular stress, disrupt mitochondrial function, and affect cell viability (Xing et al., 2012; Peng et al., 2015). In Ctenopharyngodon idella exposed to 0.015 µl/l of CPF, histopathological alterations, including cellular damage, neuronal degeneration, and vacuolization, have been reported (Khan et al., 2021). Furthermore, the elevated expression of gmfb in CPF-treated specimens may reflect its role in mediating oxidative stress-related damage, potentially through the induction of nitric oxide production (Ahmed et al., 2019). CPF exposure has been linked to inflammation in various organs, including the brain, due to the upregulation of proinflammatory biomarkers (Lopes-Ferreira et al., 2023). One potential response to this inflammatory process is CD59 overexpression, which inhibits MAC-mediated cell lysis, representing a neuroprotective response aimed at preventing complement-mediated damage, particularly in regions vulnerable to toxicant accumulation (Ibrahim et al., 2020). ConclusionThe results of the present study reveal the primary structural and functional characteristics of CD59 and GMFB in P. orinoquensis, as well as their similarities to other fish species. In the brain injury assay, low CD59 gene expression suggests a persistent loss of immunomodulatory capacity following brain damage, as observed in patients with TBI. While gmfb expression in the injured fish group significantly increased in the whole brain, no significant changes were observed in specific brain segments, indicating possible region-specific differences or transient regulation. In contrast, overexpression of gmfb (optic chiasm) and CD59 (olfactory bulb) in CPF-exposed fish demonstrated the immunoactivity and damaging role of CPF. Our findings underscore the importance of CD59 and gmfb as molecular markers of the brain immune response of P. orinoquensis. AcknowledgmentsThe authors would like to thank the Laboratory of Immunology and Molecular Biology (LIBM) at the Universidad del Tolima for providing this study. Conflict of interestThe authors declare no conflict of interest. FundingNone. Authors’ contributionsRafael E. Castro and Iang Rondón contributed to the research conception and execution planning. Rafael E. Castro performed the experiments and collected the data. Rafael E. Castro and María P. Herrera performed the data analysis, wrote the manuscript, performed the bibliographic review, and edited the manuscript. Iang Rondón contributed to the review and editing of the manuscript. All authors have read and approved the final version of the manuscript. Data availabilityAll data supporting this study’s findings are available within the manuscript. ReferencesAhmed, M.E., Selvakumar, G.P., Kempuraj, D., Raikwar, S.P., Thangavel, R., Bazley, K., Wu, K., Khan, O., Kukulka, K., Bussinger, B., Dubova, I., Zaheer, S., Govindarajan, R., Iyer, S., Burton, C., James, D. and Zaheer, A. 2019. Neuroinflammation mediated by glia maturation factor exacerbates neuronal injury in an in vitro model of traumatic brain injury. J. Neurotrauma. 37(14), 1645–1655; doi:10.1089/neu.2019.6932 Algattas, H. and Huang, J.H. 2013. Traumatic brain injury pathophysiology and treatments early intermediate and late phases post-injury. J Trauma. 57(12), 309–341; doi:10.3390/ijms15010309 Almagro Armenteros, J.J., Tsirigos, K.D., Sønderby, C.K., Petersen, T.N., Winther, O., Brunak, S., Von Heijne, G. and Nielsen, H. 2019. SignalP 50 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37(4), 420–423; doi:10.1038/s41587-019-0036-z American Veterinary Medical Association (2020). AVMA guidelines for the euthanasia of animals 2020 edition. Available via https://www.avma.org/sites/default/files/2020-02/Guidelines-on-Euthanasia-2020.pdf Accessed 17 April 2024 Bellander, B.M., Singhrao, S.K., Ohlsson, M., Mattsson, P. and Svensson, M. 2001. Complement activation in the human brain after traumatic head injury. J. Neurotrauma. 18(12), 1295–1311; doi:10.1089/08977150152725605 Bernhofer, M., Dallago, C., Karl, T., Satagopam, V., Heinzinger, M., Littmann, M., Olenyi, T., Qiu, J., Schütze, K., Yachdav, G., Ashkenazy, H., Ben-Tal, N., Bromberg, Y., Goldberg, T., Kajan, L., O’Donoghue, S., Sander, C., Schafferhans, A., Schlessinger, A., Vriend, G., Mirdita, M., Gawron, P., Gu, W., Jarosz, Y., Trefois, C., Steinegger, M., Schneider, R. and Rost, B. 2021. PredictProtein predicting protein structure and function for 29 years. Nucleic Acids Res. 49(W1), W535–W540; doi:10.1093/nar/gkab354 Blom, N., Gammeltoft, S. and Brunak, S. 1999. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294(5), 1351–1362; doi:10.1006/jmbi.1999.3310 Blum, M., Chang, H.Y., Chuguransky, S., Grego, T., Kandasaamy, S., Mitchell, A., Nuka, G., Paysan-Lafosse, T., Qureshi, M., Raj, S., Richardson, L., Salazar, G.A., Williams, L., Bork, P., Bridge, A., Gough, J., Haft, D.H., Letunic, I., Marchler-Bauer, A. and Finn, R.D. 2021. The InterPro protein families and domains database 20 years. Nucleic Acids Res. 49(1), D344–D354; doi:10.1093/nar/gkaa977 Chang, L., Sen Liou, J.C., Lin, S.R. and Huang, H.B. 2002. Purification and characterization of a neurotoxin from the venom of Ophiophagus hannah king cobra. J. Biol. Chem. Biol. Biochem. Biophys. Res. Commun. 294(3), 574–578; doi:10.1016/S0006-291X(02)00518-1 Collazos-Lasso, L.F., Ueno-Fukura, M., Arias-Castellanos, J.A. and Vinatea-Arana, L.A. 2021. Desempeño de larvas y juveniles de Piaractus orinoquensis cultivados a diferentes densidades en sistemas con tecnología biofloc. Rev. Bio. Agro. 19(2), 201–213; doi:10.18684/bsaa.v19.n2.2021.1864 Couves, E.C., Gardner, S., Voisin, T. et al (2023). Structural basis for membrane attack complex inhibition by CD59. J Biol Chem. 2023; 57: e0026. Nat Commun 14 890. https://doi.org/10.1038/s41467-023-36441-z Cruz–Méndez, J.S., Herrera–Sánchez, M.P., Céspedes–Rubio, A.E. and Rondón–Barragán, I.S. 2022. Oxidative stress response biomarker gene expression in Piaractus brachypomus (Characiformes: serrasalmidae). New Zealand J. Zool. 49(4), 355–367; doi:10.1080/03014223.2021.2017985 Cuan-Barrera, J.A., Parada-Guevara, S.L., Murillo-Pacheco, R. and Ramírez-Merlano, J.A. 2021. Parámetros productivos del cultivo de cachama blanca Piaractus orinoquensis, en jaulas flotantes. Rev. Udca Actual Dissemination 24(2), doi: 10.31910/rudca.v24.n2.2021.2068 Dinet, V., Petry, K.G. and Badaut, J. 2019. Brain–immune interactions and neuroinflammation after traumatic brain injury. Front. Neurosci. 13, 1178; doi:10.3389/fnins.2019.01178 DiSabato, D., Quan, N. and Godbout, J.P. 2016. Neuroinflammation the devil is in the details. J. Neurochem. 139(2), 136–153; doi:10.1245/s10434-020-09263-2 Doetsch, F. and Scharff, C. 2001. Challenges for brain repair: insights from adult neurogenesis in birds and mammals. Brain Behav. Evol. 58(5), 306–322; doi:10.1159/000057572 Du, Y., Teng, X., Wang, N., Zhang, X., Chen, J., Ding, P., Qiao, Q., Wang, Q., Zhang, L., Yang, C., Yang, Z., Chu, Y., Du, X., Zhou, X. and Hu, W. 2014. NF-κB and enhancer-binding CREB protein scaffolded by CREB-binding protein CBP p300 proteins regulate CD59 protein expression to protect cells from complement attack. J. Biol. Chem. 289(5), 2711–2724. Duarte-Restrepo, E., Jaramillo-Colorado, B.E. and Duarte-Jaramillo, L. 2020. Effects of chlorpyrifos on the crustacean Litopenaeus vannamei. PLos One 15(4), 231310; doi:10.1371/journal.pone.0231310 Eisenhaber, B., Bork, P. and Eisenhaber, F. 1999. Prediction of potential GPI-modification sites in proprotein sequences. J. Mol. Biol. 292(3), 741–758; doi:10.1006/jmbi.1999.3069 Elmorsy, E., Al-Ghafari, A., Al Doghaither, H., Salama, M. and Carter, W.G. 2022. An investigation of the neurotoxic effects of malathion, chlorpyrifos, and paraquat to different brain regions. Brain. Sci. 12(8), 975; doi:10.3390/brainsci12080975 Erwood, A. and Alawieh, A.M. 2022. Complement activation kindles the transition of acute post-traumatic brain injury to a chronic inflammatory disease. Neural Regen Res 17(10), 2228–2229; doi: 10.4103/1673-5374.335799 Escobar L., M.D., Farias, I.P. and Hrbek, T. 2022. Genetic comparison of populations of Piaractus brachypomus and P. orinoquensis (Characiformes: serrasalmidae) of the Amazon and Orinoco basins. Neotrop. Ichthyol. 20, e220056; doi:10.1590/1982-0224-2022-0056 Fan, J., Fong, T., Chen, X., Chen, C., Luo, P. and Xie, H. 2018. Glia maturation factor-u03b2 a potential therapeutic target in neurodegeneration and neuroinflammation. Neuropsychiatr. Dis. Treat. 14, 495–504; doi:10.2147/NDT.S157099 Ferrè, F. and Clote, P. 2005. Disulfide connectivity prediction using secondary structure information and residue frequencies. Bioinformatics 21(10), 2336–2346; doi:10.1093/bioinformatics/bti328 Font, E., Desfilisa, E., Pérez-Cañellas M Garc. and ía-Verdugo, J.M. 2001. Neurogenesis and neuronal regeneration. Brain Behav. Evol. 58, 276–295. Gan, Z., Wang, B., Zhou, W., Lu, Y., Zhu, W., Tang, J., Jian, J. and Wu, Z. 2015. Molecular and functional characterization of CD59 from Nile tilapia (Oreochromis niloticus) involved in the immune response to Streptococcus agalactiae. Fish. Shellfish. Immunol. 44(1), 50–D59; doi:10.1016/j.fsi.2015.01.035 Gasteiger, E., Hoogland, C., Gattiker, A. and et al. 2005. Protein identification and analysis tools on the ExPASy server. In: The Proteomics Protocols Handbook. Ed., Walker, J.M. Totowa, NJ: Humana Press; doi: 10.1385/1-59259-890-0:571 Goetzl, E.J., Yaffe, K., Peltz, C.B., Ledreux, A., Gorgens, K. and et al. 2020. Traumatic brain injury increases plasma astrocyte-derived exosome levels of neurotoxic complement proteins. FASEB. J. 34(2), 3359–3366; doi:10.1096/fj.201902842R Goroncy, A.K., Koshiba, S., Tochio, N., Tomizawa, T., Sato, M., Inoue, M., Watanabe, S., Hayashizaki, Y., Tanaka, A., Kigawa, T. and Yokoyama, S. 2009. NMR solution structures of actin depolymerizing factor homology domains. Protein Sci. 18, 2384–2392; doi:10.1002/pro.248 Gupta, R. and Brunak, S. 2002. Prediction of glycosylation across the human proteome and the correlation to protein function. Pac. Symp. Biocomput. 322, 310–322; doi:10.1142/9789812799623_0029 Hakala, M., 2020. The role of GMF and twinfilin in turnover of dendritic actin arrays. Dissertation, University of Helsinki, Finland Hammad, A., Westacott, L. and Zaben, M. 2018. The role of the complement system in traumatic brain injury: a review. J. Neurol. J. Neuroinflammation 15, 1–15; doi:10.1186/s12974-018-1066-z Holguín-Céspedes, G.K., Millán-Ocampo, L.M., Mahecha-Méndez, E.J. and et al. 2019. Toxicity assessment of chlorpyrifos in red-bellied pacu fingerlings (Piaractus brachypomus). Rev. Int. Contam. Ambient. 35(4), 815–829; doi:10.20937/RICA.2019.35.04.04 Ibrahim, S., Sveiven, S. and Nordgren, T. 2020. Analysis of pulmonary complement protein expression following organic dust exposure. UC. Riverside. Undergraduate. Res. J. 14(1), doi:10.5070/RJ5141049297 Inagaki, M., Aoyama, M., Sobue, K., Yamamoto, N., Morishima, T., Moriyama, A., Katsuya, H. and Asai, K. 2004. Sensitive immunoassays for human and rat GMFB and GMFG tissue distribution and age-related changes. Biochim. Biophys. Acta. Gen. Subj. 1670(3), 208–216; doi:10.1016/j.bbagen.2003.12.006 James, S.L., Bannick, M.S., Montjoy-Venning, W.C. and et al. 2019. Global regional and national burden of traumatic brain injury and spinal cord injury 1990-2016 a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18(1), 56–87; doi:10.1016/S1474-4422(18)30415-0 Jenkins JA, Bart Jr HL, Bowker JD et al (2014) Guidelines for the use of fishes in research. Bethesda, Maryland, USA: American Fisheries Society. Jia, Y., Qi, Y., Wang, Y., Ma, X., Xu, Y., Wang, J., Zhang, X., Gao, M., Cong, B. and Han, S. 2019. Overexpression of CD59 inhibits apoptosis of T-acute lymphoblastic leukemia via AKT/Notch1 signaling pathway. Cancer Cell Int. 19(1), h1–14; doi:10.1186/s12935-018-0714-9 Kaplan, R., Zaheer, A., Jaye, M. and Lim, R. 1991. Molecular Cloning and Expression of Biologically Active Human Glia Maturation Factor–β. J. Neurochem. 57(2), 483–490; doi:10.1111/j.1471-4159.1991.tb03777.x Khan, N., Sultan, A., Ali, A. and et al. 2021. Fish as bioindicator ecological risk assessment of insecticide to aquatic organisms, particularly Ctenopharyngodon idella. J. Geosci. Environ. Prot. 09(02), 42–54; doi:10.4236/gep.2021.92003 Kim, J.H. and Suk, K. 2023. Unexpected role of complement component 8 gamma chain in the inflamed brain. Neural. Regen. Res. 18(12), 2655–2656; doi:10.4103/1673-5374.373706 Kishimoto, N., Shimizu, K. and Sawamoto, K. 2012. Neuronal regeneration in a zebrafish model of adult brain injury. DMM. Dis. Model. Mech. 5(2), 200–209; doi:10.1242/dmm.007336 Lappalainen, P., Kessels, M.M., Jamie, M., Cope, T.V. and Drubin, D.G. 1998. The ADF Homology: aDF-H Domain, a Highly Exploited Actin-binding Module. Mol. Biol. Cell 9, 1951–1959. Lim, R. and Zaheer, A. 1995. Phorbol ester stimulates rapid intracellular phosphorylation of glia maturation factor. Biochem. Biophysical Res. Commun. 211, 928–934. Lim, R., Miller, J.F. and Zaheer, A. 1989. Purification and characterization of glia maturation factor f8 a growth regulator for neurons and glia. Proc. Natl. Acad. Sci. U. S. A. 86, 3901–3905. Liu, G., Zhang, J. and Chen, X. 2007. Molecular and functional characterization of a CD59 analog from large yellow croaker Pseudosciana crocea. Mol. Immunol. 44(15), 3661–3671; doi:10.1016/j.molimm.2007.04.006 Livak, K.J. and Schmittgen, T.D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25(4), 402–408; doi:10.1006/meth.2001.1262 Lopes-Ferreira M, Farinha LRL, Costa YSO et al (2023) Pesticide-induced inflammation at a glance. J Clin Exp Inflamm 2023; 57: 59-69. Toxics 11(11) 896. doi: 10.3390/toxics11110896 Lu, S., Wang, J., Chitsaz, F., Derbyshire, M.K., Geer, R.C., Gonzales, N.R., Gwadz, M., Hurwitz, D.I., Marchler, G.H., Song, J.S., Thanki, N., Yamashita, R.A., Yang, M., Zhang, D., Zheng, C., Lanczycki, C.J. and Marchler-Bauer, A. 2020. CDD/SPARCLE the conserved domain database in 2020. Nucleic. Acids. Res. 48(D1), D265–D268; doi:10.1093/nar/gkz991 Medina-Robles VM, Suárez-Martínez RO, Baldisserotto B, Cruz-Casallas PE (2023) Cryopreserved semen of Piaractus orinoquensis (Serrasalmidae): Post-thaw storage times and activating solutions. J Biol Sci. 2023; 58: e013-e018. Acta biol colomb 28(1) 86-94. https://doi.org/10.15446/abc.v28n1.95701 Millán-Ocampo, L., Torres-Cortés A Mar. and ín-Méndez, G.A. 2012. Concentraciones anestésicas del eugenol en peces escalares Pterophyllum scalare. Rev. Invest. Vet. Perú. 23(2), 171–181; doi:10.15381/rivep.v23i2.897 Naime, A.A., Lopes, M.W., Colle, D. and et al. 2020. Glutathione in chlorpyrifos-and chlorpyrifos-oxide-induced toxicity: a comparative study focused on non-cholinergic toxicity in HT22 cells. Neurotoxicity Res. 38, 603–610; doi:10.1007/s12640-020-00254-5 Ortiz Muñoz, J.D., Lozano Villegas, K.J. and Rondón Barragán, I.S. 2023. Molecular biomarkers for oxidative stress and neuronal damage in red-bellied pacu (Piaractus brachypomus). Aquac. Fish. 8(6), 720–726; doi:10.1016/j.aaf.2022.08.008 Papanastasiou AD, Georgaka E, Zarkadis IK (2007) Cloning of a CD59-like gene in rainbow trout expression and phylogenetic analysis of two isoforms. Phylogenetics and Evolution, 69, 6969–6970. Mol Immunol 44(6):1300–1306. https://doi.org/10.1016/j.molimm.2006.05.014 Patel, B., Silwal, A., Eltokhy, M.A., Gaikwad, S., Curcic, M., Patel, J. and Prasad, S. 2024. Deciphering CD59: unveiling Its Role in Immune Microenvironment and Prognostic Significance. Cancers 16(21), 3699; doi:10.3390/cancers16213699 Peng, H.F., Bao, X.D., Zhang, Y., Huang, L. and Huang, H.Q. 2015. Identification of differentially expressed proteins of brain tissue in response to methamidophos in flounder Paralichthys olivaceus. Fish & Shellfish Immunol. 44(2), 555–565; doi:10.1016/j.fsi.2015.03.028 Petano-Duque, J.M., Lozano-Villegas, K.J., Céspedes-Rubio, A.E. and Rondón-Barragán, I.S. 2022. Molecular characterization of HEPCIDIN-1 (HAMP1) gene in red-bellied pacu (Piaractus brachypomus). Dev. Comp. Immunol. 130, 104353; doi:10.1016/j.dci.2022.104353 Ploug, M. and Ellis, V. 1994. Structure-function relationships in the receptor for urokinase-type plasminogen activator comparison to other members of the Ly-6 family and snake venom α-neurotoxins. FEBS. Lett. 349(2), 163–168; doi:10.1016/0014-5793(94)00674-1 Rao BRP, Mohanty L, Kampali H et al (2024) Organophosphate-induced delayed neuropathy: a rare case presentation. J Neurosci. 2024; 57: 69-80. J Integr Med Res 2(1) 33–36. https://doi.org/10.4103/jimr.jimr_46_23 Richardson, J.R., Fitsanakis, V., Westerink, R.H.S. and Kanthasamy, A.G. 2019. Neurotoxicity of pesticides. Acta Neuropathol. 138, 343–362; doi:10.1007/s00401-019-02033-9 Rueda-García, V. and Rondón-Barragán, I.S. 2024. Molecular characterization of neurogranin (NRGN) gene from redbellied pacu (Piaractus brachypomus). Mol. Neurobiol. 61(5), 2620–2630; doi:10.1007/s12035-023-03700-5 Schmidt, R., Beil, T., Strähle, U. and Rastegar, S. 2014. Stab wound injury of the zebrafish adult telencephalon: a method to investigate vertebrate brain neurogenesis and regeneration. J. Vis. Exp. 90, 51753; doi:10.3791/51753 Scolding, N.J., Morgan, B.P. and Compston, D.A.S. 1998. The expression of complement regulatory proteins by adult human oligodendrocytes. J. Biol. Biol. Biol. J. Neuroimmunol. 84(1), 69–75; doi:10.1016/S0165-5728(97)00241-5 Selvakumar, G.P., Ahmed, M.E., Iyer, S.S., Thangavel, R., Kempuraj, D., Raikwar, S.P., Bazley, K., Wu, K., Khan, A., Kukulka, K., Bussinger, B., Zaheer, S., Burton, C., James, D. and Zaheer, A. 2020. Absence of glia maturation factor protects from axonal injury and motor behavioral impairments after traumatic brain injury. Exp. Neurobiol. 29(3), 230–248; doi:10.5607/en20017 Shabab, T., Khanabdali, R., Moghadamtousi, S.Z., Kadir, H.A. and Mohan, G. 2017. Neuroinflammation pathways a general review. Int. J. Neurosci. 127(7), 624–633; doi:10.1080/00207454.2016.1212854 Shi, Y., Chen, L., Liotta, L.A., Wan, H.H. and Rodgers, G.P. 2006. Glia maturation factor gamma GMFG a cytokine-responsive protein during hematopoietic lineage development and its functional genomics analysis. Genomics. Proteomics. Bioinf. 4(3), 145–155; doi:10.1016/S1672-0229(06)60027-2 Sledge, D., Yen, J., Morton, T., Dishaw, L., Petro, A., Donerly, S., Linney, E. and Levin, E.D. 2011. Critical duration of exposure for developmental chlorpyrifos-induced neurobehavioral toxicity. Neurotoxicology. Teratology. 33(6), 742–751; doi:10.1016/j.ntt.2011.06.005 Stahel, P.F., Flierl, M.A., Morgan, B.P., Persigehl, I. and et al. 2009. Absence of the complement regulatory molecule CD59a leads to intensified neuropathology after traumatic brain injury in mice. J. Neuroinflammation 6(2), 2; doi:10.1186/1742-2094-6-2 Steentoft, C., Vakhrushev, S.Y., Joshi, H.J. and et al. 2013. Precision mapping of the human O-GalNAc glycoprotease through SimpleCell technology. EMBO J. 32(10), 1478–1488; doi:10.1038/emboj.2013.79 Sun, C., Wu, J., Liu, S., Li, H. and Zhang, S. 2013. Zebrafish CD59 has both bacterial-binding and inhibiting activities. Develop. Comparative Immunol. 41(2), 178–188; doi:10.1016/j.dci.2013.05.008 Toutonji, A., Mandava, M., Guglietta, S. and Tomlinson, S. 2021. Chronic complement dysregulation drives neuroinflammation after traumatic brain injury: a transcriptomic study. J. Neurol. Acta. Neuropathol. Comun. 9, 126; doi:10.1186/s40478-021-01226-2 Utsuyama, M., Shiraishi, J., Takahashi, H., Kasai, M. and Hirokawa, K. 2003. Glia maturation factor produced by thymic epithelial cells plays a role in T cell differentiation in the thymic microenvironment. J. Cell Biol. Biotechnol. 96(5), 69; doi:10.1093/intimm/dxg056 Van Erp, I.A., Michailidou, I., Van Essen, T.A. and et al. 2023. Tackling Neuroinflammation After Traumatic Brain Injury: complement Inhibition as a therapy for secondary injury. J. Neurosci. 20(1), 284–303; doi:10.1007/s13311-022-01306-8 Waterhouse, A., Bertoni, M., Bienert, S. and et al. 2018. SWISS-MODEL homology modeling of protein structures and complexes. Nucleic. Acids. Res. 46(W1), W296–W303; doi:10.1093/nar/gky427 Wiltbank, A.T., Steinson, E.R., Criswell, S.J., Piller, M. and Kucenas, S. 2022. Cd59 and inflammation regulate Schwann cell development. Elife 11, e76640; doi: 10.7554/eLife.76640 Xing, H., Li, S., Wang, Z., Gao, X., Xu, S. and Wang, X. 2012. Histopathological changes and antioxidant response in brain and kidney of common carp exposed to atrazine and chlorpyrifos. Chemosphere 88(4), 377–383; doi:10.1016/j.chemosphere.2012.02.049 Xiong, C., Hanafy, S., Chan, V., Hu, Z.J., Sutton, M., Escobar, M., Colantonio, A. and Mollayeva, T. 2019. Comorbidity in adults with traumatic brain injury and all-cause mortality: a systematic review. BMJ Open 9(11), 29072; doi:10.1136/bmjopen-2019-029072 Yeh, H.Y. and Klesius, P.H. 2007. Molecular cloning and expression of channel catfish Ictalurus punctatus complement membrane attack complex inhibitor CD59. J. Mol. Biol. Chem. Biol. Vet. Immunol. Immunopathol. 120(3–4), 246–253; doi:10.1016/j.vetimm.2007.07.016 Yin, G., Du, M., Li, R., Li, K., Huang, X., Duan, D., Ai, X., Yao, F., Zhang, L., Hu, Z. and Wu, B. 2018. Glia maturation factor beta is required for reactive gliosis after traumatic brain injury in zebrafish. Exp. Neurol. 305, 129–138; doi:10.1016/j.expneurol.2018.04.008 Zapata-Guerra, N.A., Rueda-Gómez, D.S., Lozano-Villegas, K.J., Herrera-Sánchez, M.P., Uribe-García HF Rond. and ón-Barragán, I.S. 2020. Menthol as anesthetic for red-bellied pacu Piaractus brachypomus and its effect on HIF1a and GlucoR gene expression. Aquac. Res. 51, 4421–4429; doi:10.1111/are.14784 Zeis, T., Enz, L. and Schaeren-Wiemers, N. 2016. The immunomodulatory oligodendrocyte. Brain. Res. 1641, 139–148; doi:10.1016/j.brainres.2015.09.021 Zupanc, G.K.H. 2008. Adult neurogenesis and neuronal regeneration in the brain of teleost fish. J. Physiol. Paris. 102(4-6), 357–373; doi:10.1016/j.jphysparis.2008.10.007 | ||
| How to Cite this Article |
| Pubmed Style Castro-vargas RE, Herrera-sánchez MP, Rondón-barragán I. Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Vet. J.. 2026; 16(2): 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 Web Style Castro-vargas RE, Herrera-sánchez MP, Rondón-barragán I. Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. https://www.openveterinaryjournal.com/?mno=278445 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.31 AMA (American Medical Association) Style Castro-vargas RE, Herrera-sánchez MP, Rondón-barragán I. Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Vet. J.. 2026; 16(2): 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 Vancouver/ICMJE Style Castro-vargas RE, Herrera-sánchez MP, Rondón-barragán I. Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 Harvard Style Castro-vargas, R. E., Herrera-sánchez, . M. P. & Rondón-barragán, . I. (2026) Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Vet. J., 16 (2), 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 Turabian Style Castro-vargas, Rafael Enrique, María Paula Herrera-sánchez, and Iang Rondón-barragán. 2026. Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Veterinary Journal, 16 (2), 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 Chicago Style Castro-vargas, Rafael Enrique, María Paula Herrera-sánchez, and Iang Rondón-barragán. "Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury." Open Veterinary Journal 16 (2026), 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 MLA (The Modern Language Association) Style Castro-vargas, Rafael Enrique, María Paula Herrera-sánchez, and Iang Rondón-barragán. "Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury." Open Veterinary Journal 16.2 (2026), 1124-1140. Print. doi:10.5455/OVJ.2026.v16.i2.31 APA (American Psychological Association) Style Castro-vargas, R. E., Herrera-sánchez, . M. P. & Rondón-barragán, . I. (2026) Molecular characterization of CD59 and gmfb genes and their differential expression in Piaractus orinoquensis exposed to different conditions of brain injury. Open Veterinary Journal, 16 (2), 1124-1140. doi:10.5455/OVJ.2026.v16.i2.31 |