| Research Article | ||
Open Vet. J.. 2026; 16(2): 817-823
Open Veterinary Journal, (2026), Vol. 16(2): 817-823 Research Article Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first libyan record of Squalus megalopsOsama M. El-barasi1*, Ahmad M. Azab2, Mariem Feki3, Esmail Shakman4 and Mohamed A. M. El-Tabakh21Department of Zoology, Faculty of Science, Derna University, Derna, Libya 2Department of Zoology, Faculty of Science, Al-Azhar University, Cairo, Egypt 3Project Officer , WWF North Africa, EL Menzah,Tunis 4Department of Zoology, Faculty of Science, University of Tripoli, Tripoli, Libya *Corresponding Author: Osama M. El-barasi. Zoology Department, Faculty of Science, Derna University, Libya. E-Mail: o.mofftah [at] uod.edu.ly Submitted: 20/10/2025 Revised: 30/12/2025 Accepted: 12/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
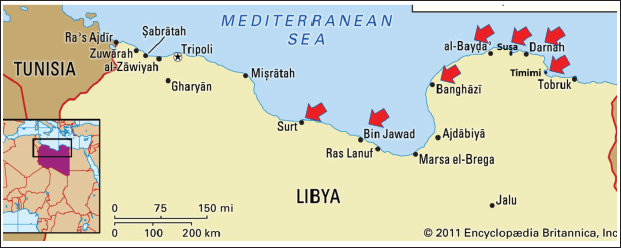
ABSTRACTBackground: A genetic analysis was conducted to identify and classify sharks from the families Squalidae and Squatinidae at the Eastern Libyan coast of the Mediterranean Sea. Aim: This study was based on COI gene barcoding, and the sequences obtained were matched with global reference databases to ascertain their taxonomic identity and genetic diversity ratio. Methods: The sequences were then deposited in the GenBank database through the BankIt submission system, where the following accession numbers were assigned: Squalus mitsukurii (PV717998), Squalus megalops (PV718644), Squatina oculata (PV716693), Squatina squatina (PV738303), and Squatina aculeata (PV719649). The phylogenetic evolution tree was constructed by applying the neighbor-joining method (Chromas Pro v1.5 Beta). Results: Squalus mitsukurii was the first confirmed genetic record of the Mediterranean Sea, whereas S. megalops was the first confirmed genetic record of the Libyan coast. In addition, S. oculata, S. squatina, and S. aculeata were genetically confirmed in Libyan waters for the first time. Haplotype network analysis revealed a clear distinction of genetic distance among the species, with little variation among the species. Nucleotide diversity and Tajima D were computed, and both showed a strong deviation from neutrality, which could have been caused by either selective pressures or past demographic changes. Conclusion: Collectively, these findings contribute significantly to the ichthyofaunal diversity of Libya and the Mediterranean Sea in general, and they further demonstrate the power of DNA barcoding as a means of recording and preserving shark diversity. Keywords: Squalus mitsukurii, Squalus megalops, Mediterranean Sea, Libya, DNA barcoding. IntroductionThe Libyan coastline spans almost 2,000 km, making it the longest stretch among the Mediterranean coastlines, covering approximately 3.9% of its southern border (Zupanovic and El-Buni, 1982). Sharks are the top trophic level in any marine ecosystem, where they function as key regulators in ensuring ecological stability (Cortes, 1999). Although ecologically significant, elasmobranch species are largely typified by low growth rate, delayed maturation, and low fecundity compared with selfish (Cortés and E, 2004). These biological characteristics lead to their low capacity to withstand fishing pressure and lead to naturally low rates of population growth (Smith et al., 2009). The lack of basic information on the majority of shark and ray species is increasingly becoming a problem for fisheries managers and scientists worldwide. One of the factors that adversely impacts data collection is the misidentification of many species in the chondrichthyans and their body parts. When a key diagnostic structure has been destroyed or lost, morphologically very similar species may prove difficult to identify and poorly prepared and/or defaced specimens (Holmes et al., 2009; El-Tabakh et al., 2024a; Al-Khatib et al., 2025). A proposed solution is the possibility of a fast and precise species identification system, i.e., DNA barcoding, that could sequence a region of the mitochondrial cytochrome C oxidase 1 gene (COX1), which has the potential to identify any stage of its life. In the case of sharks and rays, it may offer a convenient means of inferring whether the sharks are harvested legally or illegally, and the shark fin perhaps harvested for the shark fin trade, which is rather lucrative (Ward et al., 2008; Moftah et al., 2011; Al-Khatib et al., 2022). Moreover, DNA barcoding of samples can provide a good identification of those species even when the specimen is incomplete or when the species under consideration has demonstrated the occurrence of cryptic variability (Vella et al., 2017; Afifi et al., 2023; El-Tabakh et al., 2024b; Afifi et al., 2024). Regardless of the ecological significance of the sharks, the Libyan cost of the eastern and central Mediterranean is one of the least examined territories on the genetic level, which creates significant uncertainty in terms of species identification and diversity. Thus, this study aimed to offer the initial molecular characterization of Squalidae and Squatinidae family sharks along the Libyan coast using mitochondrial DNA barcoding. In this way, a significant information gap will be addressed in the region and provide much-needed baseline data on the genetic diversity and distribution of these species. Materials and MethodsSamples collectionA total of 23 shark individuals from 5 species (2 species belonging to the family Squalidae and 3 species belonging to the Squatinidae family) were collected from fish markets in key landing sites, including At-Tamimi, Derna, Susa, Al-Bayda, Benghazi, Bin Jawad, and Sirte, between November 2023 and November 2024 (Fig. 1). Sampling sites are the key shark landing zones in the area. The species were morphologically identified according to Compagno (2001). Tissue samples were taken from each shark and preserved in 70% ethanol at −20 °C for further molecular analyses. DNA extraction and analysesSubsamples of the tissue samples were extracted from the collected specimens, stored in 70% ethanol, and refrigerated at −20 C. The Qiagen DNeasy Blood and Tissue Kit, under the standard spin-column protocol, was used to extract genomic DNA. Tissues were lysed using ATL buffer and Proteinase K at 56°C, bound, washed using AW1/AW2 buffers, and finally eluted in 70 AE buffer. The extracted DNA was kept at −20°C, and its concentration and purity were studied using a NanoDrop 2000. To monitor potential contamination, negative controls were included during both DNA extraction and PCR. The 50 regions of the cytochrome C oxidase 1 (COXI) gene in the mitochondrial DNA were amplified in 655 bp using designed primers (Ward et al., 2005): FishF1 (5’-TCAACCAACCACAAAGACATTGGCAC-3’) FishR1 (5’-TAGACTTCTGGGTGGCCAAAGAATCA-3’) The PCR reaction (25 µl) contained 5.5 µl water (nuclease free) 12.5 µl Green master mix, 1 µl primer of each, and 5 µl of genomic DNA. Then, it was amplified in a thermal cycler under the following conditions: the initial denaturation step was performed at 95°C with 2 minutes, followed by 35 cycles of 30 seconds at 94°C, 30 seconds at 54°C, and 1 minutes at 72°C, and a final extension of 10 minutes at 72 °C. Amplification of products was performed, and the PCR products were stored at 4 °C. Before sequencing, amplicons were subconfirmed on 1.5%, quantitated, and purified. The Sanger method was used to perform sequencing in both directions, and the ends with poor sequences were lopped off, and the sequence aligned. Forward and reverse sequencing of both strands was performed. Sequencing was 105 performed using the Sanger method on an ABI capillary sequencer. The sequence data were assembled using chromas Pro 1.5 beta (Technelysium Pty., Australia). The ambiguous bases and low-quality ends were manually trimmed before alignment. The newly mapped COXI sequences were compared with the initial gathering in GenBank by BLAST (http://blast.ncbi.nlm.nih.gov/Blast. cgi) to determine the species. BLAST searches were performed using an E-value threshold of 1 × 10−5, a minimum identity of ≥98%, query coverage ≥95%, and sequences producing more than 2% gaps were excluded from downstream analysis.
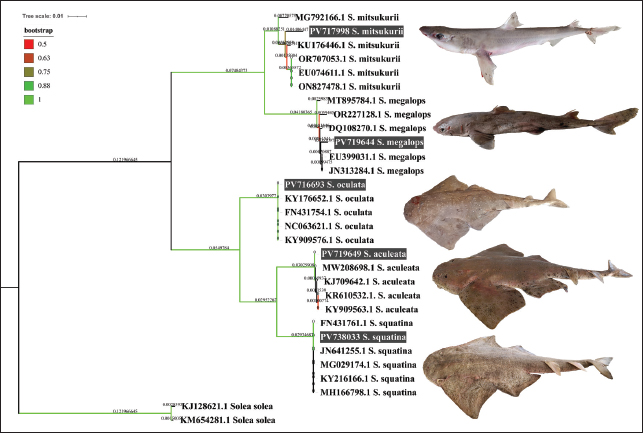
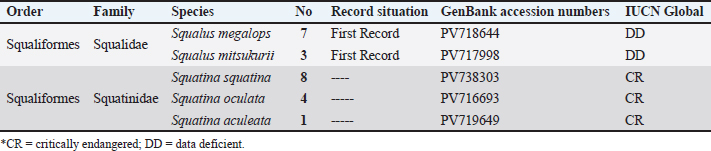
Fig. 1. Map showing the land markets along the eastern Libya coast of the Mediterranean Sea. The sequences were successfully deposited in the GenBank database through Bank It and given accession numbers. All GenBank submissions included publicly accessible metadata, including sampling locality and specimen information, and were verified to ensure correct linkage within the GenBank database. To calculate the genetic divergence of the genes by using the Tamura-Nei method (Tamura and Nei, 1993), multiple sequence alignment was conducted using MEGA 11.0 and MUSCLE. It was analyzed with the neighbor-joining (NJ) algorithm (Tamura, 1992) in terms of Tamura 3-parameter model, computed by bootstrap replicates (1000) using MEGA software V 11.0.13 (Tamura et al., 2021). iTOL was used to visualize the trees (Letunic and Bork, 2021). Two other tools, DnaSP v6.12.03 and PopArt v3.0, were also used to draw a minimum spanning network that shows the haplotype diversity and divergence. Following the elimination of low-quality ends, a 464-bp alignment was achieved using MUSCLE with default settings. This methodology is in line with the elasmobranch barcoding studies mentioned in the Introduction. Ethical approvalThe National Subcommittee for Bioethics College of Medical Technology, Derna, Ministry of Technical and Vocational Education, Libya, granted ethical approval for this research (approved on September 09, 2024). ResultsThe collected shark species have quite considerable global IUCN status according to Table 1. Squalus megalops and Squalus mitsukurii are two Squalidae species that are registered herein as the first records, with the former being the first record in the Mediterranean Sea and the latter being the first record along the Libyan coast, and appeared in the global assessment as DD. A review of all available records for Libya and the Mediterranean, which revealed no prior reports of this species, supports this conclusion. Three species of the family Squatinidae (Squatina squatina, Squatina oculata, and Squatina aculeata) are listed as critically endangered (CR) on the IUCN Red List. An evolutionary tree of the taxa under investigation was estimated by a neighbor-joining (NJ) method (Saitou and Nei, 1987) with evolutionary distances computed by the Tamura-Nei method (Tamura and Nei, 1993) and its value denoted in the units of the number of base substitutions per site. The phylogenetic evolution tree included 30 nucleotide sequence sets containing the 1st, 2nd, and synonymous 3rd codon positions and noncoding sites, respectively, with the ambiguous sites removed using pairwise deletion, yielding a total of 464 positions. The best phylogeny (Fig. 2) was obtained with a bootstrap of 1,000 replicates, with branch color assigned to the bootstrap value (Felsenstein, 1985). Reference COI sequences from related species were used as the context for rooting, and branches with bootstrap replicates below 50% were collapsed. Table 1. IUCN condition of the collected sharks.
Fig. 2. Neighbor-joining phylogenetic evolution.
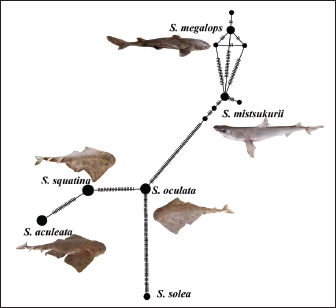
Fig. 3. Minimum haplotype spanning network. The circle size is proportional to the haplotype frequency, and each line connecting the circles represent one mutational step separating the haplotypes. The phylogenetic reconstruction showed separate clustering groups after the species designation. The genetic identification of the obtained sharks was performed by PCR and sequencing. The sequences were successfully deposited in the GenBank database through Bank It and given accession numbers. S. mitsukurii (PV717998), marking the first genetic note about the species in the Mediterranean Sea, clustering with strong bootstrap support with S. mitsukurii specimens representing other locations within the world, which may reflect a wider distribution than previously documented; S. megalops (PV718644), representing the first genetic evidence of the potential existence of the species on the Libyan coast; S. oculata (PV716693), S. squatina (PV738303), and S. aculeata (PV719649), representing the angel shark species, and are the first genetic report of those species in Libya, yet the demonstration that they clustered within strongly supported clades substantiates their molecular identification. A minimum haplotype-spanning network (Fig. 3) was constructed in PopArt v3.0 to illustrate the haplotype distribution. The genetic indices reported in this study (π, number of segregating sites, and Tajima’s D) are directly associated with the haplotype network analysis. Each species in the network displayed divergent haplotypes with small intraspecific differences, indicating measurable haplotype diversity within the same species. The COI alignment includes multiple Squatina and Squalus species from two families, with a teleost outgroup (Solea solea). The nucleotide diversity (DNA diversity, pi) across the dataset was 2,377.95, estimated at 118 segregating sites (of which 115 were informative to parsimony analysis). Tajima D=36128.9 with p=0 when the condition D ≥ 36128.9 was observed, which clearly shows a significant deviation in the possibility of neutrality, which could be a form of selective pressures or the opposite, which could be the historical demographic changes. These combined phylogenetic and haplotype analyses strengthen the genetic distinctiveness of the newly documented S. mitsukurii and S. megalops specimens, which are the first molecular evidence of Libyan Squatina spp., and provide important access to the ichthyofaunal records of the Mediterranean Sea and Libyan waters (Table 1). DiscussionThis is the first genetic (and subsequently genetic) record of Squalus mitsukurii in the Mediterranean Sea and a significant finding within the complicated taxonomic history of this species. A recent systematic study has shown that Squalus mitsukurii belongs to a species complex that includes Squalus blainvillei (Ward et al., 2007; Veríssimo et al., 2017). Such taxonomic complexity has led to extensive reclassification of its geographic distribution, and many prior historical records beyond the North Pacific Ocean have been nullified after reexamination using molecular techniques. Among these revisions are those of the South Pacific (Duffy and Last, 2007; Last et al., 2007), the South Atlantic and the western Indian Oceans (Viana et al., 2016 and 2017), and the north-eastern and the north-western Atlantic (Verisimo et al., 2017; Pfleger et al., 2018). In this context, the molecular assay of COXI sequences in the current study revealed evident grouping of the Libyan specimen of S. mitsukurii with respect to other conspecific sequences at other localities spread over the rest of the world, followed by a high bootstrap value in the phylogeny tree. This genetic affinity does not necessarily imply a natural range expansion. Although the exact cause cannot be confirmed, similar distributional shifts have been documented for several elasmobranch species in the Mediterranean, suggesting that natural range dynamics driven by environmental changes may play a role, which could be aided by environmental or oceanographic processes, but also a non-natural introduction, possibly associated with anthropogenic ones (e.g., maritime transport, inter-state fishing). Such a record is especially significant because little genetic data was available on the chondrichthyan species on the Libyan coast. DNA barcoding was most important in providing the correct identification of other morphologically similar elasmobranch fish, as postulated earlier, to overcome the problem of misidentification, which has interfered with biodiversity surveys in the area (Ward et al., 2005; Holmes et al., 2009). These findings are consistent with those of recent Mediterranean barcoding work, including that of El-Tabakh Et Al. (2024)who similarly demonstrated the effectiveness of COI markers in resolving taxonomic uncertainty among closely related elasmobranch species. Not only does the molecular confirmation of the presented paper improve the accuracy of the local faunal inventories, but it also creates a nice reference point to monitor and future conservation efforts. In the case of Squalus megalops, not previously reported in Libyan waters, the results conformed to the local observations of Marouani et al. (2023)in Tunisian waters and observations of Shakman et al. (2023)on S. blainville, wherein taxonomic uncertainty in the genus Squalus was discussed. In our dataset, the two species were clearly separated, showing distinct COI haplotypes and consistent interspecific divergence, supporting their molecular differentiation. These practical difficulties have been greatly influenced by the high morphological similarity among congeners, along with ongoing synonymous usage, often in the absence of distinctive morphological characteristics. In particular, until very recently, no documented report could confirm the presence of S. megalops in Libyan waters. However, this species had been previously recorded in the Egyptian Mediterranean waters by Shaban and El-Tabakh (2019) who identified it based on pectoral fin morphology and morphometric analysis. In contrast, more recent molecular studies, including those conducted in the Arca Gulf and Strait of Sicily regions, where the Libyan coast is located (Ferrari et al., 2021), did not provide genetic evidence for the occurrence of S. megalops in the Mediterranean Sea. Therefore, our records may reflect related but genetically undocumented species, such as S. blainvillei, or morphologically similar specimens yet to be identified at the genetic level. In this regard, the present work has provided the first confirmed genetic evidence of S. megalops in Libya, a contribution that is urgently needed to fill the knowledge gap regarding its distribution in the Eastern Mediterranean. This also confirmed that better biogeographic assessments and targeted conservation strategies for threatened species in the region can be achieved through the integration of both molecular and morphological data, thereby reducing the likelihood of misidentification. This study is the first genetic confirmation of three angel shark species (S. aculeata, S. oculata, and S. squatina) in Libyan waters. Although these species had been morphologically identified (e.g., S. aculeata in Zupanovic and El-Buni (1982) S. oculata in Lamboeuf et al. (1995) and S. squatina in Dauphin (1966) none of these had been genetically confirmed before the present study. Because Squatina species are quite morphologically similar, morphology-based species-level identification is bound to be made with uncertainty. This study improved upon those results by providing the first molecular evidence of the occurrence of these species in Libyan waters, thus addressing a relevant knowledge gap in the local elasmobranch diversity and improving the effectiveness of existing distribution information critical to exemplary conservation planning. The use of COXI-based DNA barcoding in the present study addresses such uncertainties and identifies the accurate taxonomic position of each of the species, which is supported by defined clades in the phylogenetic tree. This is the first confirmed genetic documentation of the presence of these angel shark species in Libyan waters, and this will enhance the credibility of regional studies on the region’s biodiversity. This kind of confirmation is given even more weight on the conservation front, since all three species are classified as threatened under the IUCN Red List, and precise distribution information is important in implementing management and protection measures. ConclusionIn Libya, this study is the first to use genetic barcoding methodology to identify shark species, a breakthrough in marine biodiversity studies in the region. Squalus mitsukurii has been genetically identified and forms the first record of the whole Mediterranean Sea. Squalus megalops was the first recorded species along the Libyan coast. In addition, the genetic analysis of S. aculeate, S. oculata, and S. squatina in Libyan waters formed a strong baseline for future taxonomic and conservation research. Such success demonstrated the importance of molecular tools in assessing species identification within taxonomically difficult groups, especially where morphologically similar taxonomic groups are involved, and the need to incorporate genetic analyses in fisheries monitoring and biodiversity estimation programs in Libya. Through the generation and deposition of new sequences of COXI genes in globally accessible databases, the study will contribute to a rich genetic resource across international databases and could also strengthen the scientific basis of future successful conservation of vulnerable and little-known elasmobranch species in the region. Future work should expand sampling coverage across additional regions of the Libyan coast and incorporate multi-locus or genomic markers to better resolve the population structure and long-term dynamics of these shark species. AcknowledgmentsThis research was partly supported by Research England. SL received funding from WWF-NL, whereas LM, SD, and VM were supported by WWF-UK. Additional financial support was provided by the National Center for Research and Consultation at the University of Derna under a cooperation agreement between the University and WWF North Africa. Conflict of interestsThe author(s) declare(s) that there are no conflicts of interest. Authors’ contributionsConceptualization, Ahmad M. Azab; Osama M. El-barasi; Databarasi;Data curation, Mohamed A. M. El-Tabakh; Osama M. El-barasi; Formal analysis, Mohamed A. M. El-Tabakh; Osama M. El-barasi; Funding acquisition, Mariem Feki; Investigation, Mohamed A.M. El Tabakh; Osama M. El-barasi Review & editing, Ahmad M. Azab; Mohamed A. M. El-Tabakh; Osama M. El-barasi; Mariem Feki & Esmail Shakman. All authors have read and agreed to the publication of this manuscript. FundingThis study was funded by IKEA via WWF’s forest practice and was undertaken in partnership with WCS. Data availabilityThe datasets used and analyzed in this study are available from the corresponding author on reasonable request. ReferencesAfifi, M.A.M., Azab, A.M., Ali, E., Ghazy, A. and El-Tabakh, M.A.M. 2024. DNA barcoding, phylogeography and evolutionary dynamics of Chrysichthys auratus. Gene 917, 148448. Afifi, M.A.M., Sarhan, M., Khalaf-Allah, H.M.M., Alabssawy, A.N., Abbas, M.M.M., Abdel-Hamid, F. and El-Tabakh, M.A.M. 2023. DNA barcoding supports life stages of wrasse fish, Novaculichthys taeniourus (Family Labridae) in the Hurghada Red Sea, Egypt. Zoologischer. Anzeiger. 304, 84–93. Al-khatib Et Al., Z.I.M. 2022. DNA barcoding and newly taxonomical traits for the shark taxonomy: a new approach. Egypt. J. Aquatic Biol. Fisheries 26(6), 1133–1144. Al-Khatib, Z.I.M., Azab, A.M., El-Tabakh, M.A.M. and Afifi, M.A.M. 2025. Quantitative morphometric analysis of dorsal fin for species identification in sharks (Family: carcharhinidae) from the Suez Gulf, Red Sea, Egypt. Medit. Aquac. J. 12(1), 13–27. Compagno, L. J. V. 2001. Sharks of the World: An Annotated and Illustrated Catalogue of Shark Species Known to Date. Vol. 1. Food and Agriculture Organization of the United Nations (FAO), Rome, Italy. Cortes, E. 1999. Standardized diet compositions and trophic levels of sharks. ICES J. Mar. Sci. 56(5), 707–717. Cortés, E., 2004. Life history patterns, demography, and population dynamics. Biology of sharks and their relatives, pp. 449–469. Dauphin, J.C. 1966. French Provincial Centers of Documentation and Research on Africa. Afr. Stud. Rev. 9(3), 48–65. Duffy, C.A.J., and Last, P.R., 2007. Redescription of the northern spiny dogfish Squalus griffini Phillipps, 1931 from New Zealand. Descriptions of new dogfishes of the genus Squalus, pp. 91–101. El-Tabakh, M.A., Azab, A.M., Khalaf-Allah, H.M., Sarhan, M., Abdelwarith, M.S., Besar, M.H., Zeina, A.F., Alabssawy, A.N., Bashar, M.A., Abo-Taleb, H.A. and Attallah, M.A., 2024. Unveiling diversity in shark fin characteristics: A comparative study across order Carcharhiniforms. Egyptian J. Aquatic Biol. Fisheries, 28(2), 235–246. El-Tabakh, E.T. 2024b. DNA barcoding for the crayfish Procambarus clarkii (Girard, 1852) from the River Nile, Egypt. Egypt. J. Aquatic Biol. Fisheries 28(6), 985–997. Felsenstein, J. 1985. Confidence limits on phylogenies: an approach using the bootstrap. evolution 39(4), 783–791. Ferrari, A., Di Crescenzo, S., Cariani, A., Crobe, V., Benvenuto, A., Piattoni, F., Mancusi, C., Bonnici, L., Bonello, J.J., Schembri, P.J., Serena, F., Massi, D., Titone, A. and Tinti, F. 2021. Puzzling over spurdogs: molecular taxonomy assessment of the Squalus species in the Strait of Sicily. Eur. Zoological J. 88(1), 181–190. Holmes, B.H., Steinke, D. and Ward, R.D. 2009. Identification of shark and ray fins using DNA barcoding. Fisheries Res. 95(2-3), 280–288. Lamboeuf, M., Ben Abdallah, A., Zgozi, S., Nafati, A., Amer, A. and Abdulbari, R. 1995. Libyan marine resource assessment: trawl survey results, 1993–94. Last, P.R., White, W.T. and Motomura, H., 2007. Part 6—A description of Squalus chloroculus sp. nov., a new spurdog from southern Australia, and the resurrection of S. montalbani Whitley. CSIRO Marine Atmos Res Paper, 14, pp. 55–69. Letunic, I. and Bork, P. 2021. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49(W1), W293–W296. Marouani, S., Karaa, S. and Jarboui, O., 2023. Overview of the genus Squalus in the Mediterranean Sea. In: Sharks – Past, Present and Future.London, United Kingdom: IntechOpen, pp. 1–27. https://doi.org/10.5772/intechopen.108977 wMoftah, M., Abdel Aziz, S.H., Elramah, S. and Favereaux, A. 2011. Classification of sharks in the Egyptian Mediterranean waters using morphological and DNA barcoding approaches. PLos One 6(11), e27001. Pfleger, M.O., Grubbs, R.D., Cotton, C.F. and Daly-Engel, T.S. 2018. Squalus clarkae sp. nov., a new dogfish shark from the Northwest Atlantic and Gulf of Mexico, with comments on the Squalus mitsukurii species complex. Zootaxa 4444(2), 101–119. Saitou, N. and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4(4), 406–425. Shakman, E., Siafenasar, A., Etayeb, K., Shefern, A., Elmgwashi, A., Al Hajaji, M., Bek Benghazi, N., Ben Abdalha, A., Aissi, M. and Serena, F. 2023. National inventory and status of Chondrichthyes in the South Mediterranean Sea (Libyan coast). Biodiversity J. 14(3), 459–480. Shaban, W.M., and El-Tabakh, M.A.M., 2019. New records, conservation status and pectoral fin description of eight shark species in the Egyptian Mediterranean waters. Egyptian J. Aquat. Biol. Fisher. 23(4), 503–519. Smith, M.A., Fernandez-Triana, J., Roughley, R. and Hebert, P.D.N. 2009. DNA barcode accumulation curves for understudied taxa and areas. Mol. Ecol. Resour. 9, 208–216. Tamura, K. and Nei, M. 1993. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10(3), 512–526. Tamura, K., Stecher, G. and Kumar, S. 2021. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38(7), 3022–3027. Tamura, K. 1992. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+ C-content biases. Mol. Biol. Evol. 9(4), 678–687. Vella, A., Vella, N. and Schembri, S. 2017. A molecular approach towards taxonomic identification of elasmobranch species from Maltese fisheries landings. Mar. Genomics. 36, 17–23. Veríssimo, A., Zaera-Perez, D., Leslie, R., Iglésias, S.P., Séret, B., Grigoriou, P., Sterioti, A., Gubili, C., Barría, C., Duffy, C. and Hernández, S., 2017. Molecular diversity and distribution of eastern Atlantic and Mediterranean dogfishes Squalus highlight taxonomic issues in the genus. Zoologica scripta, 46(4), 414–428. Viana, S.T.D.F., Carvalho, M.R.D. and Gomes, U.L. 2016. Taxonomy and morphology of species of the genus Squalus Linnaeus, 1758 from the Southwestern Atlantic Ocean (Chondrichthyes: squaliformes: Squalidae). Zootaxa 4133(1), 1–89. Viana, S.T.D.F.L., De Carvalho, M.R. and Ebert, D.A. 2017. Squalus bassi sp. nov., a new long-snouted spurdog (Chondrichthyes: squaliformes: Squalidae) from the Agulhas Bank. J. Fish. Biol. 91(4), 1178–1207. Ward, R.D., Holmes, B.H., White, W.T. and Last, P.R. 2008. DNA barcoding Australasian chondrichthyans: results and potential uses in conservation. Mar. Freshwater. Res. 59(1), 57–71. Ward, R.D., Holmes, B.H., Zemlak, T.S., Smith, P.J., Last, P., White, W. and Pogonoski, J., 2007. Part 12—DNA barcoding discriminates spurdogs of the genus Squalus. Descriptions of new dogfishes of the genus Squalus (Squaloidea: Squalidae), pp. 117–130. Ward, R.D., Zemlak, T.S., Innes, B.H., Last, P.R. and Hebert, P.D. 2005. DNA barcoding Australia’s fish species. Phil. Trans. Roy. Soc. B. Biol. Sci. 360(1462), 1847–1857. Zupanovic, S. and El-Buni, A.A., 1982. A contribution to demersal fish studies off the Libyan coast. Bulletin of the Marine Research Centre, Tripoli, Libya, 3, pp. 78–122. | ||
| How to Cite this Article |
| Pubmed Style El-barasi OM, Azab AM, Feki M, Shakman E, El-tabakh MAM. Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Vet. J.. 2026; 16(2): 817-823. doi:10.5455/OVJ.2026.v16.i2.5 Web Style El-barasi OM, Azab AM, Feki M, Shakman E, El-tabakh MAM. Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. https://www.openveterinaryjournal.com/?mno=291696 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.5 AMA (American Medical Association) Style El-barasi OM, Azab AM, Feki M, Shakman E, El-tabakh MAM. Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Vet. J.. 2026; 16(2): 817-823. doi:10.5455/OVJ.2026.v16.i2.5 Vancouver/ICMJE Style El-barasi OM, Azab AM, Feki M, Shakman E, El-tabakh MAM. Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 817-823. doi:10.5455/OVJ.2026.v16.i2.5 Harvard Style El-barasi, O. M., Azab, . A. M., Feki, . M., Shakman, . E. & El-tabakh, . M. A. M. (2026) Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Vet. J., 16 (2), 817-823. doi:10.5455/OVJ.2026.v16.i2.5 Turabian Style El-barasi, Osama M., Ahmad M. Azab, Mariem Feki, Esmail Shakman, and Mohamed A. M. El-tabakh. 2026. Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Veterinary Journal, 16 (2), 817-823. doi:10.5455/OVJ.2026.v16.i2.5 Chicago Style El-barasi, Osama M., Ahmad M. Azab, Mariem Feki, Esmail Shakman, and Mohamed A. M. El-tabakh. "Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops." Open Veterinary Journal 16 (2026), 817-823. doi:10.5455/OVJ.2026.v16.i2.5 MLA (The Modern Language Association) Style El-barasi, Osama M., Ahmad M. Azab, Mariem Feki, Esmail Shakman, and Mohamed A. M. El-tabakh. "Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops." Open Veterinary Journal 16.2 (2026), 817-823. Print. doi:10.5455/OVJ.2026.v16.i2.5 APA (American Psychological Association) Style El-barasi, O. M., Azab, . A. M., Feki, . M., Shakman, . E. & El-tabakh, . M. A. M. (2026) Genetic taxonomical study of the families squalidae and squatinidae with the first mediterranean record of Squalus mitsukurii and the first Libyan record of Squalus megalops. Open Veterinary Journal, 16 (2), 817-823. doi:10.5455/OVJ.2026.v16.i2.5 |