| Research Article | ||
Open Vet. J.. 2026; 16(2): 953-961
Open Veterinary Journal, (2026), Vol. 16(2): 953-961 Research Article Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shopsHajer Shams Al-Din Abdullah1, Ashraf Saddik Alias2 and Omar Hashim Sheet3*1Department of Environmental Sciences, College of Environmental Sciences, University of Mosul, Mosul, Iraq 2Department of Physiology, Biochemistry and Pharmacology, College of Veterinary Medicine,University of Mosul, Mosul, Iraq 3Department of Veterinary Public Health, College of Veterinary Medicine, University of Mosul, Mosul, Iraq *Corresponding Author: Omar Hashim Sheet. Department of Veterinary Public Health, College of Veterinary Medicine, University of Mosul, Mosul, Iraq. Email: omar.sheet [at] uomosul.edu.iq Submitted: 23/10/2025 Revised: 05/01/2026 Accepted: 19/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
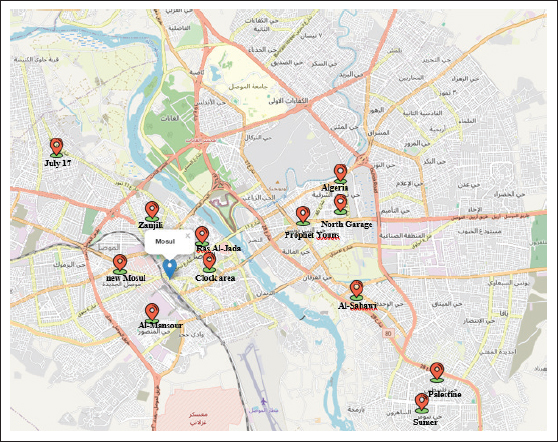
ABSTRACTBackground: The bacterium Escherichia (E.) coli has long been acknowledged as a major causative agent of food-borne illness, with the ability to cause several illnesses and fatalities nationwide. It has the ability to produce Stx1 and Stx2 toxins that cause food poisoning. Aim: The primary goals of the current study were to isolate and identify E. coli classical methods and confirmed by detecting the uidA gene using a polymerase chain reaction (PCR) assay to identify the Stx1 and Stx2 genes in E. coli isolated. Methods: Three hundred samples of veal meat sold at retail and various parts of butchers’ shops were collected from each side of Mosul city. Results: The prevalence of E. coli in the current investigation was 48%. The high prevalence of E. coli isolated from the floor was 61.7%. The prevalence of E. coli isolated from veal meat, tables, walls, and worker’s hand was 60%, 56.7%, 31.7%, and 30%, respectively. A statistically significant difference in contamination rates was observed among sample types (χ²=23.96, p < 0.001). The highest prevalence was recorded in the floor samples (61.7%). No statistically significant difference was observed between the two areas (χ²=0.00, p > 0.05), indicating similar hygienic and contamination conditions in butcher shops on both sides of the city. Additionally, PCR analysis confirmed the presence of the uidA gene in all E. coli isolates (100%), the presence of the Stx1 gene in 73.3%, and the presence of the Stx2 gene in 23.3% of E. coli isolates. Four different gene profiles were found in E. coli. Profile II (uidA + Stx1) accounted for 53.3% of all isolates. Conclusion: All butcher shops’ equipment spread E. coli, the retail meat was tainted by the bacteria, and the E. coli isolates had the Stx1 and Stx2 genes. Keywords: Butchers, E. coli, Retail meat, Shiga toxin genes, Shops. IntroductionMeat is one of the most valuable sources of nutrition for humans because of its rich composition of essential nutrients. Meat is a crucial source of high biological value protein that provides all of the essential amino acids required for human growth, maintenance, and metabolism. Meat is also a source of critical micronutrients, including iron, zinc, selenium, and B vitamins, particularly vitamin B12, which are important for development, tissue repair, and body function preservation. Numerous zoonotic foodborne microorganisms are capable of producing different kinds of toxins, which can cause food poisoning. The pathogenic bacteria can be transferred from animals to humans through consumers (Neamah et al., 2022). Meat can get contaminated by zoonotic foodborne germs while animals’ carcasses are being dressed, eviscerated, transported, and stored in the abattoir (Awosile et al., 2021). Furthermore, meat is exposed to contaminants, such as pollutants, rodents, and insects, and unsafe conditions during animal slaughter and meat preparation (Laury and Echeverry, 2009; Amniattalab and Ghalandarzadeh, 2019). Foodborne pathogenic bacteria isolated from meat and from numerous human food poisoning cases are of greatest importance (Kalin and Öngör, 2014; Yousif et al., 2021 ; Othman et al., 2023; Sheet et al., 2023). Escherichia coli has phenotypic characteristics and inhabits both human and animal digestive systems (Tenaillon et al., 2010). In addition, E. coli spreads through food and can cause a number of deadly infections in both humans and animals (Tarr et al., 2005). E. coli has several genes that encode virulence factors, such as uidA and Shiga toxin 1 and 2 genes (Krüger and Lucchesi, 2015). Furthermore, E. coli harboring the uidA gene produces the β-glucuronidase enzyme, which is essential for the hydrolysis of glucuronides into glucuronic acid (Majowicz et al., 2014). Two different types of toxins produced by E. coli are known as Shiga Toxins 1 and 2. Shiga toxin synthesis. This type of E. coli, frequently referred to as Escherichia coli (STEC), is responsible for global outbreaks and isolated cases of STEC illnesses, resulting in a variety of problems (Castro et al., 2019). Escherichia coli can be identified using various techniques, including molecular biology and conventional methods. Conventional methods rely on media (selection and enrichment media), Gram stain, which analyzes the shape and response of E. coli colonies for isolation, and biochemical tests, which use E. coli DNA to identify the bacteria (Lozinak et al., 2016). The molecular biological approaches depend on identifying the target sequence of the specific-species gene and applying them to validate the findings of traditional tests. Molecular biology methods produce simpler, less expensive, quicker, and more precise results, reducing the identification time from several days to just a few hours (Gao et al., 2011). The present study aims to: (1) isolate and identify E. coli isolated from Mosul City’s butchers’ shops; (2) confirm the E. coli isolates by detecting the uidA gene; and (3) detect the Stx1 and Stx2 genes in E. coli isolates. Materials and MethodsSamples collectionIn this study, 300 randomly collected samples were taken from veal meat sold at retail and various spots of butcher shops, including workers, hands, tables, walls, and floors, in various regions on the right (Ras Al-Jada, Zanjili, July 17, Clock area, Al-Mansour, and new Mosul) and left (Sumer, Palestine, North Garage, Al-Sabawi, Algeria, and Prophet Youns) sides of Mosul city, Iraq (Fig. 1). Furthermore, 60 samples were obtained from each worker’s hands, veal meat, tables, walls, and floors on both sides of the city. The study period began in February 2025 and ended in July 2025. Sterile containers were used to collect meat, and swabs were used to collect other samples. The samples were then taken to the Laboratory of Public Health at the College of Veterinary Medicine, University of Mosul, Iraq. All samples were pre-enriched by diving in peptone water and then incubated at 37°C for the entire night.
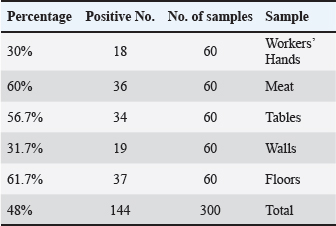
Fig. 1. Geographical map of Mosul illustrating the sampling points. Escherichia coli isolation and characterizationSwabs from butcher shops (workers, hands, tables, walls, and floors) and veal meat samples were used to isolate and identify pathogenic E. coli. After inoculation with the nutrient broth (LAB, UK), all samples and swabs were incubated for 24 hours at 37°C for 24 hours. One loop of nutrient broth was spread onto MacConkey agar (LAB, UK) and EMB for the classical culture method, which was then incubated for 24 h at 37°C. Additionally, the present study used Brilliance E. coli/coliform Agar (Oxoid, UK). The putative E. coli isolates were verified using the IMViC series of biochemical tests (Momtaz et al., 2013). Before being used further for workshop examination, all E. coli isolates were frozen in brain heart infusion broth (15% glycerol) at ?80°C. DNA isolationEscherichia coli was cultivated on selective media at 37°C for 24 h. E. coli’s deoxyribonucleic acid was isolated following the directions on the FlaPure Bacteria Genomic DNA Extraction Kit (Genesand Kit, China). A NanoDrop spectrophotometer (Jenway, UK) was used to determine the DNA concentration. The range of DNA concentrations was 30 ng/μl to 60 ng/μl. Reaction of the PCRThe detection of E. coli genes—including uidA, Stx1, and Stx2—was carried out using polymerase chain reaction (PCR) assays. The following amplicon sizes were expected for every target gene: uidA, 623 bp (Moyo et al., 2007), Stx1, 347 bp (Fujioka et al., 2013), and Stx2, 282 bp (Brian et al., 1992) (Table 1). These gene targets were selected for their relevance in confirming species identity and Shiga toxin production. The PCR reaction was performed in 200 μl PCR tubes (Biozym, Germany) with a total volume of 25 μl. After being amplified, the PCR products were identified by electrophoresis on a 2% agarose gel (Peqlab, Germany). A molecular size marker of 100 bp DNA was used to measure the fragment lengths of the ensuing amplicons. The PCR reaction mixture consisted of 12.5 μl of Master Mix (2×) (Addbio, Korea), 4 μl of E. coli genomic DNA, 1 μl of each primer (1 and 2), and 6.5 μl of DNeasy-free water (Promega Corporation, USA). Statistical analysisStatistical analysis was performed using the JMP® 16.1 software (SAS Institute Inc., 2021). The chi-square test revealed a significant variation in the percentage of E. coli isolates across various regions in Mosul City, with a p-value of 0.05. Ethical approvalAll animal procedures were approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Mosul’s College of Veterinary Medicine under approval number UM. Vet. 2025.072. The animal owners provided their informed consent for the sample to be utilized, and all procedures followed ethical guidelines. ResultsThe colony morphology showed that the E. coli isolates were positive for specific biochemical assays and IMViC tests. According to spectrophotometry, the quantity of genomic DNA extracted varied from 30 µg/l to 60 µg/l (Fig. 2). All E. coli isolates were confirmed to be positive for the species-specific uidA gene, supporting their molecular designation. PCR further supported the results from traditional microbiological techniques. Table 2 shows that the overall prevalence of E. coli isolated from veal meat sold at retail and butchers’ shops was 48% (144/300). The highest isolation rate was observed from floor samples, which was 61.7% (37/60) of the positive cases. In contrast, the lowest prevalence rates were detected in samples taken from workers’ hands (30%) (18/60). Furthermore, the percentages of E. coli isolated from veal meat, tables, and walls were 60% (36/60), 56.7% (34/60), and 31.7% (19/60), respectively. In addition, the chi-square test was used to evaluate differences in E. coli prevalence among sample types (hands, meat, tables, walls, and floors). The overall prevalence of E. coli among all examined samples was 48% (144/300). A statistically significant difference in contamination rates was observed among sample types (χ²=23.96, p < 0.001). The highest prevalence was recorded in floor samples (61.7%), meat samples (60%), and table surfaces (56.7%), whereas lower prevalence rates were observed on wall surfaces (31.7%) and workers’ hands (30%) (Table 2).
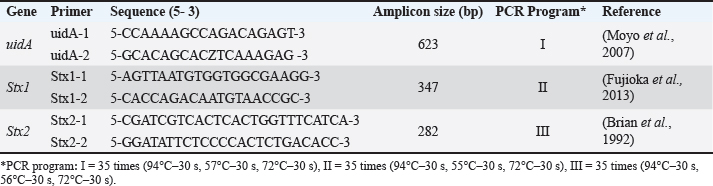
Fig. 2. Comparative DNA concentrations of different E. coli isolates. Table 1. PCR program and primers for uidA, Stx1, and Stx2 genes detection in E. coli.
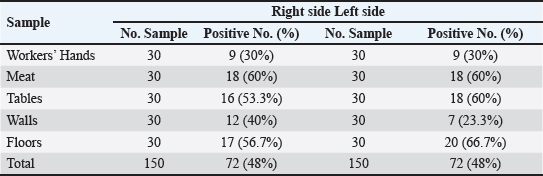
Furthermore, this investigation discovered that retail meat and butcher shops on Mosul’s left and right flanks contributed 48% (72/150) of the E. coli samples. Moreover, the highest occurrence of E. coli isolated from veal meat on the right side was 60% (18/30), whereas the lowest incidence of E. coli isolated from workers’ hands was 30% (9/30). The occurrence of E. coli isolated from walls, tables, and floors was 40% (12/30), 53.3% (16/30), and 56.7% (17/30), respectively. Additionally, the largest percentage of E. coli found on the flooring was found on the left side, at 66.7% (20/30). The lower incidence of E. coli found on the walls was 23.3% (7/30). Subsequently, it was discovered that 30% (9/30), 60% (18/30), and 60% (18/30) of the E. coli isolates came from the workers’ hands, veal meats, and tables, respectively. In addition, the results of the present study indicated no statistically significant differences in the isolation rates of E. coli among the various types of samples on the right side. Comparative analysis between the right and left sides of Mosul City showed identical overall prevalence rates (48%), with no statistically significant difference between the two areas (χ²=0.00, p > 0.05), indicating similar hygienic and contamination conditions in butcher shops on both sides of the city (Table 3). Table 2. E. coli prevalence in retail meat samples and butchers’ shops
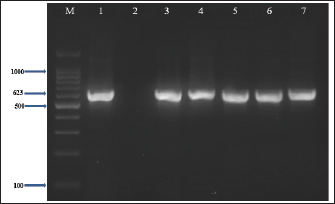
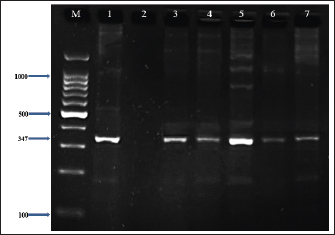
Fig. 3. Visualization of the uidA gene (623 bp) amplicon from E. coli isolates using 2% agarose gel electrophoresis. Lane M is DNA markers using a 100-bp ladder from Biozym Diagnostic. Lane 1 contains a positive control, Lane 2 shows a negative control, and Lanes 3–7 display-positive isolates. Table 3. Comparative prevalence of Escherichia coli isolates from meat and butcher shops in the right and left areas of Mosul City
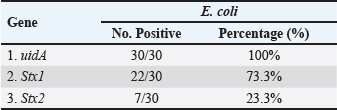
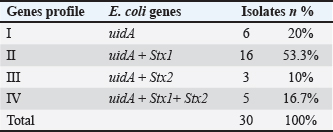
Table 4 PCR analysis confirmed the presence of the uidA gene in all E. coli isolates (100%) (Fig. 3). In addition, PCR analysis revealed that the Stx1 gene was present in 73.3% (22/30) of E. coli isolates, whereas the Stx2 gene was detected in 23.3% (7/30) (Figs. 4 and 5). Table 5 reveals that the E. coli isolates had four different gene profiles. Profile II (uidA + Stx1) was the most common profile, accounting for 53.3% (16/30) of all isolates. Profile IV (uidA + Stx1+ Stx2), which had the greatest gene-rich profile, was observed in 16.7% (5/30) of the isolates. Profile I (uidA) and Profile III (uidA + Stx2) were observed in 20% (6/30) and 10% (3/30) of the isolates, respectively.
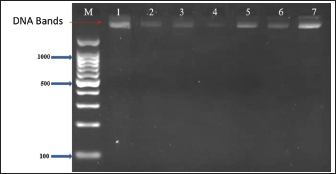
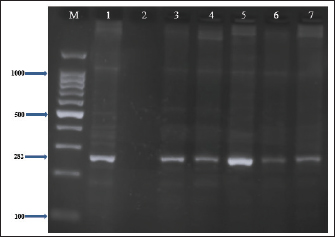
Fig. 4. Stx1 gene (347 bp) amplicon visualization from E. coli isolates using 2% agarose gel electrophoresis. Lane M is DNA markers using a 100-bp ladder from Biozym Diagnostic. Lane 1 contains a positive control, Lane 2 shows a negative control, and Lanes 3–7 display-positive isolates. Table 4. Frequency and prevalence of uidA, Stx1, and Stx2 in E. coli (n=30).
DiscussionMany types of foodborne bacteria cause food poisoning through the consumption of contaminated foods. The origin of E. coli from food-producing animals plays a crucial role in human infection, primarily through the ingestion of pathogen-carrying food products that serve as pathogen reservoirs (Busani et al., 2006). The data obtained confirmed the presence and distribution of E. coli in 48% of butchers’ shops. The obtained data were nearly of those reported in a study conducted in Iraq that showed the prevalence of E. coli in butchers and shops was 45% (Hameed et al., 2021). Moreover, the E. coli occurrence rate on each side of Mosul city was 72%. The results of our study exceeded the findings of other investigations, including the incidence of E. coli isolated from butchers and shops in Mansoura City, Egypt, which was 26.7% (Sallam et al., 2013); in Hawassa City, Ethiopia, where E. coli was isolated from butcher shops and meat samples at a rate of 2.3% (Worku et al., 2022); and in Buenos Aires, Argentina, where the isolation rate was 11.1% (Barril et al., 2019). In contrast, the current study’s findings were lower than those identified in a previous study conducted in Saudi Arabia, where the isolation rate of E. coli reached 60% (Iyer et al., 2013). The tolerance of bacteria to various physical and chemical surroundings, along with insufficient sanitation and disinfection practices designed to kill them, may be the cause of this variation in E. coli isolation rates. Furthermore, blood, feces, mucus fluids, insufficiently chlorinated water, and contaminated hides offer a perfect environment for E. coli to grow and survive for a long time.
Fig. 5. Stx2 gene (282 bp) amplicon visualization from E. coli isolates using 2% agarose gel electrophoresis. Lane M is DNA markers using a 100-bp ladder from Biozym Diagnostic. Lane 1 contains a positive control, Lane 2 shows a negative control, and Lanes 3–7 display-positive isolates. Table 5. Genotypic profiles of E. coli isolated from retail meat and butcher shops.
Additionally, 60% of E. coli were isolated from meat. The occurrence of E. coli from meat was greater in the United States (21.1%) (Tate et al., 2021), India (43.1%) (Sethulekshmi et al., 2018), and Egypt (54%) (Gwida et al., 2014) than in the other countries (60%) (Gwida et al., 2014). The incidence of E. coli isolated from meat was lower in South Africa (74.5%) (Vorster et al., 1994) and Burkina Faso (100%) (Kagambega et al., 2011). Differences in meat preparation procedures, national or district factors pertaining to animal feeding schemes, and the methods utilized for microbiological testing could all contribute to the mixture in E. coli infection rates in meat (Rigobelo et al., 2008). A number of variables, including animal skin, meat-cutting equipment, unsanitary conditions, and a drop in worker hygiene, contribute to the contamination of meat and related products with E. coli (Boukary et al., 2012). The majority of the population in impoverished republics choose to purchase low-cost meat from unregulated markets, which do not have hygienic requirements or meat safety protocols (James et al., 2014). Furthermore, this study found that the prevalence of E. coli in the hands of workers was 30%. Our results were in agreement with those of a study in Saudi Arabia, which recorded the prevalence of E. coli in workers’ hands as 30% (Elabbasy et al., 2021). In addition, our results showed that the prevalence of E. coli from workers’ hands was higher than that in Ghana (10%) (Adzitey et al., 2021) and Ethiopia (16.7%). (Sebsibe and Asfaw, 2020). The prevalence of E. coli in workers’ hands in Morocco was 80%, which was higher than the results of our study (Bahir et al., 2022). There are many reasons why butchers’ hands could be contaminated with E. coli. Being in direct contact with tainted meat or animal internal organs throughout slaughter and cutting is one of the main reasons, since fecal debris and intestinal contents. Poor handwashing practices, such as not using soap, disinfectants, or clean water after handling raw meat or utilizing the restroom, are another factor that leads to bacterial contamination. Additionally, the results of this study found that the prevalence of E. coli in the walls and floor of butchers’ hands was 31.7% and 61.7%, respectively. The floor and walls of butcher shops can become contaminated for a number of reasons. Throughout slaughter or evisceration, intestinal contents and fecal wastes may leak or splash, contaminating surrounding walls and floors, or aerosols created by high-pressure water hose cleaning transmit bacteria from contaminated floors to walls, tools, and meat surfaces. The persistence of E. coli on surfaces can be caused by insufficient cleaning, a lack of effective disinfectants, or a lack of time for sanitizers to come into contact with surfaces. Furthermore, 73.3% of E. coli had the Stx1 gene and 23.3% had the Stx2 gene. Our results were consistent with further research indicating that the Stx1 gene is more frequently found in E. coli isolates than the Stx2 gene (Ranjbar et al., 2017; Nehoya et al., 2020). Various studies have reported variations in the presence of the Stx1 and Stx2 genes in E. coli isolates; according to one study, 51% of isolates had the Stx2 gene and 29% of isolates had the Stx1 gene (Mora et al., 2007 ). Previous investigations found that 8.8% of isolates had both Stx1 and Stx2, 5.3% of isolates contained Stx1, and 86% of isolates had Stx2 (Llorente et al., 2014), whereas E. coli isolated from Vietnamese meat and fish did not possess the Stx1 or Stx2 genes (Van et al., 2008). The diversity in Stx1 and Stx2 gene occurrence across E. coli isolates can be attributed to host-related, ecological, and molecular characteristics. The observed heterogeneity is a result of differences in phage-integration effectiveness, gene stability, and horizontal gene transmission events. Bacteriophages carry both the Stx1 and Stx2 genes at the molecular level (Franz et al., 2007). Ecologically, gene prevalence depends on the source of isolation (cattle, goats, meat, humans, etc.). For example, some investigations showed that Stx1 was more prevalent in isolates from small ruminants, but Stx2 was more prevalent in strains related to humans or cows (Ndegwa et al., 2020). Lastly, sample type, region, seasonal variations, and animal husbandry techniques are additional factors that influence gene carriage rates. All of these factors work together to explain why certain E. coli populations have stx11ed, others have Stx2, and others have neither. ConclusionThis study assessed the sanitation practices of meat and butcher shops on both sides of the city of Mosul. The isolation of E. coli from meat samples that contamination may have occurred at various processing stages, including handling, transportation, storage, or slaughter, most likely under unhygienic conditions. Moreover, the presence of E. coli on the surfaces of butcher shops and workers’ hands underscores their significant contribution to the spread of the bacterium, facilitating contamination of meat and related products. Furthermore, the detection of E. coli on meat and within butcher shops indicates either repeated use of equipment without adequate sanitation or improper washing, cleaning, and sterilization procedures. Such lapses may lead to E. coli contamination of meat, posing a significant health risk to consumers. Unsanitary conditions can facilitate the proliferation of E. coli, enhancing the risk of Shiga toxin production and leading to foodborne diseases among consumers. The E. coli isolates obtained in this study exhibited considerable genetic diversity and carried both Stx1 and Stx2 genes, which was confirmed by the identification of several virulence-associated genes. AcknowledgmentOur heartfelt thanks go out to the College of Veterinary Medicine of the University of Mosul for their helpfulness and for providing the facilities and materials required for performing this study. FundingThe authors declare that no governmental or non-governmental organization provided financial support for this research. Authors’ contributionsThe authors were involved in the collection of samples and laboratory analysis. The authors performed data analysis and interpretation. The third author drafted and revised the manuscript, and approved the final manuscript. Conflict of interestThe author verifies that there were no conflicts of interest associated with the study’s publication, data analysis, or authorship. Data availabilityThe data supporting this study’s findings are available from the corresponding author on reasonable request. ReferencesAdzitey, F., Huda, N. and Shariff, A.H.M. 2021. Phenotypic antimicrobial susceptibility of Escherichia coli from raw meats, ready-to-eat meats, and their related samples in one health context. Microorganisms 9(2), 326; doi:10.3390/microorganisms9020326 Amniattalab, A. and Ghalandarzadeh, B. 2019. Prevalence and pathological evaluation of hepatic fatty change in cattle slaughtered at Urmia abattoir, northwest Iran. Iraqi J. Vet. Sci. 33(1), 45–50; doi:10.33899/ijvs.2019.125518.1040 Awosile, B., Eisnor, J., Saab, M.E., Heider, L. and McClure, J.T. 2021. Occurrence of extended-spectrum beta-lactamase and AmpC-producing Escherichia coli in retail meat products from the Maritime Provinces, Canada. Can. J. Microbiol. 67(7), 537–547; doi:10.1139/cjm-2020-0442 Bahir, M.A., Errachidi, I., Hemlali, M., Sarhane, B., Tantane, A., Mohammed, A., Belkadi, B. and Filali-Maltouf, A. 2022. Knowledge, attitude, and practices (KAP) regarding meat safety and sanitation among carcass handlers operating and assessment of bacteriological quality of meat contact surfaces at the Marrakech slaughterhouse. Food. Sci. 1(1), 4881494. Barril, P.A., Soto, S.A., Jaureguiberry, M.V., Gottardi, G., Bascur, I., Leotta, G.A. and Oteiza, J.M. 2019. Microbiological risk characterization in butcher shops from the province of Neuquen, Patagonia Argentina. LWT 107, 35–40; doi:10.1089/fpd.2020.2925 Boukary, A.R., Thys, E., Rigouts, L., Matthys, F., Berkvens, D., Mahamadou, I., Yenikoye, A. and Saegerman, C. 2012. Risk factors associated with bovine tuberculosis and molecular characterization of Mycobacterium bovis strains in Niger’s urban settings. Transbound. Emerg. Dis. 59(6), 490–502; doi:10.1111/j.1865-1682.2011.01302.x Brian, M.J., Frosolono, M., Murray, B.E., Miranda, A., Lopez, E.L., Gomez, H.F. and Cleary, T.G. 1992. Polymerase chain reaction for diagnosis of enterohemorrhagic Escherichia coli infection and hemolytic-uremic syndrome. J. Clin. Microbiol. 30(7), 1801–1806; doi:10.1128/jcm.30.7.1801-1806.1992 Busani, L., Scavia, G., Luzzi, I. and Caprioli, A. 2006. Laboratory surveillance for prevention and control of foodborne zoonoses. Super. Sanità. 42(4), 401–404; https://pubmed.ncbi.nlm.nih.gov/17361061/ Castro, V.S., Figueiredo, E.E.D.S., Stanford, K., Mcallister, T. and Conte-Junior, C.A. 2019. Shiga-toxin producing Escherichia coli in Brazil: a systematic review. Microorganisms 7(5), 137; doi:10.3390/microorganisms7050137 Elabbasy, M.T., Hussein, M.A., Algahtani, F.D., Abd El-rahman, G.I., Morshdy, A.E., Elkafrawy, I.A. and Adeboye, A.A. 2021. MALDI-TOF MS-based typing for rapid screening of multiple antibiotic-resistant E. coli and virulent non-O157 shiga toxin-producing E. coli isolated from slaughterhouse settings and beef carcasses. Foods 10(4), 820; doi:10.3390/foods10040820 Franz, E., Klerks, M.M., De Vos, O.J., Termorshuizen, A.J. and Van Bruggen, A.H.C. 2007. Prevalence of Shiga toxin-producing Escherichia coli stx 1, stx 2, eaeA, and rfbE genes and survival of E. coli O157: h7 in manure from organic and low-input conventional dairy farms. Appl. Environ. Microbiol. 73(7), 2180–2190; doi:10.1016/j.applenviron.2015.09.010 Fujioka, M., Otomo, Y. and Ahsan, C.R. 2013. A novel single-step multiplex polymerase chain reaction assay for detecting diarrheagenic Escherichia coli. J. Microbial Methods 92(3), 289–292; doi:10.1016/j.mimet.2012.12.010 Gao, J., Ferreri, M., Liu, X.Q., Chen, L.B., Su, J.L. and Han, B. 2011. Development of multiplex polymerase chain reaction assay for rapid detection of Staphylococcus aureus and selected antibiotic resistance genes in bovine mastitic milk samples. Diagn. Invest. 23(5), 894–901; doi:10.1177/1040638711416964 Gwida, M., Hotzel, H., Geue, L. and Tomaso, H. 2014. Occurrence of Enterobacteriaceae in raw meat and in human samples from Egyptian retail sellers. Int. Sch. Res. Notices 1(1), 565671; doi:10.1155/2014/565671 Hameed, M.A.K., Muhammed, H. and Al-Rashedi, A. 2021. The bacterial load and contamination rate of raw meat in butcher shops in different areas of Karbala City. Pak. J. Med. 4(4), 960; doi:10.1016/S014 Iyer, A., Kumosani, T., Yaghmoor, S., Barbour, E., Azhar, E. and Harakeh, S. 2013. Escherichia coli and Salmonella spp. in meat in Jeddah, Saudi Arabia. J. Infect. Dev. Ctries. 7, 812–818; doi:10.3855/jidc.3453 James, W.O., Cheryl, M.E.M., Makitabc, K. and Grace, D. 2014. Investigation of the food value chain of ready-to-eat chicken and the associated risk for staphylococcal food poisoning in Tshwane Metropole, South Africa. Food Control. 45, 87–94; doi: 10.1016/j.foodcont.2014.04.026 Kagambega, A., Haukka, K., Siitonen, A., Traore, A.S. and Barro, N. 2011. Prevalence of Salmonella enterica and the hygienic indicator Escherichia coli in raw meat at markets in Ouagadougou, Burkina Faso. Food Prot. 74, 1547–1551; doi:10.4315/0362-028X.JFP-11-124 Kalin, R. and Öngör, H. 2014. Examination of Escherichia coli O157: h7 and some Virulence Genes in Marketed Minced Meat Samples. Fak. Derg. 20(6), 957–960; doi:10.9775/kvfd.2014.11310 Krüger, A. and Lucchesi, P.M.A. 2015. Shiga toxins and stx phages: highly diverse entities. Microbiol 161(3), 451–462; doi:10.1099/mic.0.000003 Laury, A., Echeverry, A. and Brashears, M., 2009. Fate of Escherichia coli O157: H7 in Meat. In safety of meat and processed meat. New York, NY: Springer New York, pp: 31–53 Llorente, P., Barnech, L., Irino, K., Rumi, M.V. and Bentancor, A. 2014. Characterization of Shiga Toxin‐Producing Escherichia coli isolated from ground beef in different socioeconomic strata markets in Buenos Aires, Argentina. Biomed. Res. Int. 1, 795104; doi:10.1155/2014/795104 Lozinak, K.A., Jani, N., Gangiredla, J., Patel, I., Elkins, C.A., Hu, Z., Kassim, P.A., Myers, R.A. and Laksanalamai, P. 2016. Investigation of potential shiga toxin-producing Escherichia coli (STEC) associated with a local foodborne outbreak using a multidisciplinary approach. Food Sci. Hum. Well. 5(3), 163–168; doi:10.1016/j.fshw.2016.07.002 Majowicz, S.E., Scallan, E., Jones-Bitton, A., Sargeant, J.M., Stapleton, J., Angulo, F.J., Yeung, D.H. and Kirk, M.D. 2014. Global incidence of human Shiga toxin–producing Escherichia coli infections and deaths: a systematic review and knowledge synthesis. Foodborne. Pathogens. Dis. 11(6), 447–455; doi:10.1089/fpd.2013.1704 Momtaz, H., Dehkordi, F.S., Rahimi, E. and Asgarifar, A. 2013. Detection of Escherichia coli, Salmonella species, and Vibrio cholerae in tap water and bottled drinking water in Isfahan, Iran. BMC. Public. Health. 13, 556; doi:10.1186/1471-2458-13-556 Mora, A., Blanco, M., Blanco, J.E., Dahbi, G., López, C., Justel, P., Alonso, M.P., Echeita, A., Bernárdez, M.I., González, E.A. and Blanco, J. 2007. Serotypes, virulence genes and intimin types of Shiga toxin (verocytotoxin)-producing Escherichia coli isolates from minced beef in Lugo (Spain) from 1995 through 2003. BMC Microbiol. 7(1), 1–9. Mora, A., Blanco, M., Blanco, J.E., Dahbi, G., López, C., Justel, P., Alonso, M.P., Echeita, A., Bernárdez, M.I., González, E.A. and Blanco, J. 2013. Serotypes, virulence genes, and intimin types of Shiga toxin (verocytotoxin)-producing Escherichia coli isolates from minced beef in Lugo, Spain, from 1995 to 2003. BMC. Microbiol. 7(13), 13; doi:10.1186/1471-2180-7-13 Moyo, S.J., Maselle, S.Y., Matee, M.I., Langeland, N. and Mylvaganam, H. 2007. Identification of diarrheagenic Escherichia coli isolated from infants and children in Dar es Salaam, Tanzania. BMC Infect. Dis. 7(1), 92; doi:10.1186/1471-2334-7-92 Ndegwa, E., Alahmde, A., Kim, C., Kaseloo, P. and O’Brien, D. 2020. Age-related differences in phylogenetic diversity, prevalence of Shiga toxins, Intimin, Hemolysin genes and select serogroups of Escherichia coli from pastured meat goats detected in a longitudinal cohort study. BMC. Vet. Res. 16, 266; doi:10.1186/s12917-020-02479-0 Neamah, A.A., Fahed, K.H., Sadeq, J.N. and Alfatlawi, M.A. 2022. Molecular characterization and phylogenetic analysis of Escherichia coli isolated from the milk of cattle affected by mastitis. Iraqi J. Vet. Sci. 36(1), 251–254; doi:10.33899/ijvs.2021.129934.1702 Nehoya, K.N., Hamatui, N., Shilangale, R.P., Onywera, H., Kennedy, J. and Mwapagha, L.M. 2020. Characterization of Shiga toxin-producing E. coli in raw beef from informal and commercial abattoirs. PLos One. 15(12), 243828; doi:10.1371/journal.pone.0243828 Oguttu, J.W., McCrindle, C.M., Makita, K. and Grace, D. 2014. Investigation of the food value chain of ready-to-eat chicken and the associated risk of staphylococcal food poisoning in Tshwane, South Africa. Food Control 45, 87–94; doi:10.1016/j.foodcont.2014.04.026 Othman, S.M., Sheet, O.H. and Al-Sanjary, R. 2023. Phenotypic and genotypic characterization of Escherichia coli isolated from veal meats and butchers’ shops in the city of Mosul, Iraq. Iraq J. Vet. Sci. 37(1), 225–260; doi:10.33899/ijvs.2022.133819.2306 Ranjbar, R., Masoudimanesh, M., Dehkordi, F.S., Jonaidi-Jafari, N. and Rahimi, E. 2017. Shiga (Vero)-toxin producing Escherichia coli isolated from the hospital foods; virulence factors, o-serogroups and antimicrobial resistance properties. Antimicrob. Resist. 58(1), 4; doi:10.1186/s13756-016-0163-y Rigobelo, E.C., Santo, E. and Marin, J.M. 2008. Beef carcass contamination by Shiga toxin–producing Escherichia coli strains in an abattoir in Brazil: characterization and resistance to antimicrobial drugs. Foodborne. Pathog. Dis. 5(6), 811–817; doi:10.1089/fpd.2008.0138 Sallam, K.I., Mohammed, M.A., Ahdy, A.M. and Tamura, T. 2013. Prevalence, genetic characterization and virulence genes of sorbitol-fermenting Escherichia coli O157: h-and E. coli O157: H7 isolated from retail beef. Int. J. Food Microbiol. 165(3), 295–301; doi:10.1016/j.ijfoodmicro.2013.05.024 SAS Institute Inc. 2021. JMP®, version 16. SAS Institute Inc., Cary, North Carolina, USA. Sebsibe, M. A., and Asfaw, E. T. 2020. Occurrence of multi-drug resistant Escherichia coli and Escherichia coli O157: H7 in meat and swab samples of various contact surfaces at abattoir and butcher shops in Jimma town, southwest district of Ethiopia. Infect. 58, e013–e018. Sethulekshmi, C., Latha, C. and Anu, C.J. 2018. Occurrence and quantification of Shiga toxin-producing Escherichia coli in food matrices. Vet. World. 11(2), 104–111; doi:10.14202/vetworld.2018.104-111 Sharif, Y.M. and Tayeb, B.A. 2021. Estimation of the detection limit of Salmonella typhimurium in artificially contaminated chicken meat using cultured-based and polymerase chain reaction techniques. Iraq. J. Vet. Sci. 35(4), 621–625; doi:10.33899/ijvs.2020.127328.1496 Sheet, O.H., Al-Mahmood, O.A., Othamn, S.M., Al-Sanjary, R.A., Alsabawi, A.H. and Abdulhak, A.A. 2023. Detection of mecA-positive Staphylococcus aureus isolated from meat and butchers’ shops in Mosul city using PCR. Iraq J. Vet. Sci. 37(4), 865–870; doi:10.33899/ijvs.2023.136964 Tarr, P.I., Gordon, C.A. and Chandler, W.L. 2005. Shiga-toxin-producing Escherichia coli and hemolytic uremic syndrome. Lancet 365(9464), 1073–1086; doi:10.1016/s0140-6736(05)71144-2 Tate, H., Li, C., Nyirabahizi, E., Tyson, G.H., Zhao, S., Rice-Trujillo, C., Jones, S.B., Ayers, S., M’ikanatha, N.M., Hanna, S. and Ruesch, L. 2021. A National Antimicrobial Resistance Monitoring System Survey of Antimicrobial-Resistant Foodborne Bacteria Isolated from Retail Veal in the United States. Food. Prot. 84(10), 1749–1759; doi:10.4315/JFP-21-005 Tenaillon, O., Skurnik, D., Picard, B. and Denamur, E. 2010. The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 8(3), 207–217; doi:10.1038/nrmicro2298 Van, T.T.H., Chin, J., Chapman, T., Tran, L.T. and Coloe, P.J. 2008. Safety of raw meat and shellfish in Vietnam: an analysis of Escherichia coli isolation for antibiotic resistance and virulence genes. Int. J. Food Microbiol. 124(3), 217–223; doi:10.1016/j.ijfoodmicro.2008.03.029 Vorster, S.M., Greebe, R.P. and Nortje, G.L. 1994. Incidence of Staphylococcus aureus and Escherichia coli in ground beef, broilers and processed meats in Pretoria, South Africa. Food. Prot. 57(4), 305–310; doi:10.4315/0362-028X-57.4.305 Worku, W., Desta, M. and Menjetta, T. 2022. High prevalence and antimicrobial susceptibility pattern of Salmonella species and extended-spectrum β-lactamase producing Escherichia coli from raw cattle meat at butcher houses in Hawassa city, Sidama regional state, Ethiopia. PLos One 17, 262308; doi:10.1371/journal.pone.0262308 Yousif, M.S. and Tayeb, B.A. 2021. Estimation of limit of detection of Salmonella typhimurium in artificially contaminated chicken meat by cultured-based and polymerase chain reaction techniques. Iraqi J. Vet. Sci. 35(4), 621–5; doi: 10.33899/ijvs.2020.127328.1496 | ||
| How to Cite this Article |
| Pubmed Style Abdullah HSA, Alias AS, Sheet OH. Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Vet. J.. 2026; 16(2): 953-961. doi:10.5455/OVJ.2026.v16.i2.17 Web Style Abdullah HSA, Alias AS, Sheet OH. Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. https://www.openveterinaryjournal.com/?mno=292282 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.17 AMA (American Medical Association) Style Abdullah HSA, Alias AS, Sheet OH. Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Vet. J.. 2026; 16(2): 953-961. doi:10.5455/OVJ.2026.v16.i2.17 Vancouver/ICMJE Style Abdullah HSA, Alias AS, Sheet OH. Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 953-961. doi:10.5455/OVJ.2026.v16.i2.17 Harvard Style Abdullah, H. S. A., Alias, . A. S. & Sheet, . O. H. (2026) Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Vet. J., 16 (2), 953-961. doi:10.5455/OVJ.2026.v16.i2.17 Turabian Style Abdullah, Hajer Shams Al-din, Ashraf Saddik Alias, and Omar Hashim Sheet. 2026. Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Veterinary Journal, 16 (2), 953-961. doi:10.5455/OVJ.2026.v16.i2.17 Chicago Style Abdullah, Hajer Shams Al-din, Ashraf Saddik Alias, and Omar Hashim Sheet. "Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops." Open Veterinary Journal 16 (2026), 953-961. doi:10.5455/OVJ.2026.v16.i2.17 MLA (The Modern Language Association) Style Abdullah, Hajer Shams Al-din, Ashraf Saddik Alias, and Omar Hashim Sheet. "Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops." Open Veterinary Journal 16.2 (2026), 953-961. Print. doi:10.5455/OVJ.2026.v16.i2.17 APA (American Psychological Association) Style Abdullah, H. S. A., Alias, . A. S. & Sheet, . O. H. (2026) Molecular detection of the Stx1 and Stx2 genes in Escherichia coli isolated from veal meat sold at retail and butchers’ shops. Open Veterinary Journal, 16 (2), 953-961. doi:10.5455/OVJ.2026.v16.i2.17 |