| Research Article | ||
Open Vet. J.. 2026; 16(3): 1496-1510 Open Veterinary Journal, (2026), Vol. 16(3): 1496-1510 Research Article Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learningAxel Villalobos-Cortés1*, Ana Flores De León2 and Marcelino Jaén31Conservation and Animal Improvement. LABMA, Panama City, Panama 2Bachelor’s Degree from the Veterinary Medicine, Faculty of Veterinary Medicine, University of Panama, Panama City, Panama 3Tropical Veterinary Diseases, Animal Health Validity Laboratory, IDIAP, Institute of Agricultural Innovation of Panama, Panama City, Panama *Corresponding Author: Axel Villalobos-Cortés. Conservation and Animal Improvement. LABMA, IDIAP. City of Knowledge, Panama City, Panama. Email: villalobos.axel [at] gmail.com Submitted: 06/11/2025 Revised: 15/02/2026 Accepted: 26/02/2026 Published: 31/03/2026 © 2026 Open Veterinary Journal
ABSTRACTBackground: Guaymi and Guabala Creole cattle are unique genetic resources adapted to tropical systems in Panama. Aim: This study aimed to compare the morphometry and morphology using classical zoometric analysis and automated classification via the Mooebius web app and random forest models. Methods: Morphological and morphometric data from 225 adult cattle were analyzed using nested analysis of variance and chi-square tests. A supervised Random Forest (RF) classifier and web-based index calculator (Mooebius) were developed for aptitude prediction. Results: There were significant differences (p < 0.05) in body proportions and pelvic conformation. The Guaymi cattle showed lighter morphologies with dairy aptitude, whereas the Guabala were more robust and meat-oriented. The RF models achieved accuracies ranging from 86.4% (raw measurements) to 90.9% (index-based models), identifying body length and thoracic perimeter as the most influential predictors. Conclusion: The integration of morphological and morphometric data with artificial intelligence techniques provides an effective decision support tool for the conservation and selection of native breeds in tropical conditions. Keywords: Creole cattle, Machine learning, Morphometry, Tropical livestock, Zoometric indices. IntroductionThe introduction of cattle ranching in Central America dates back to at least 1,521, when the Spanish Crown granted Pedrarias Davila, governor of Castilla del Oro, a royal charter, allowing him to import cattle from the island of Santiago (now Jamaica) to the region of Darien on the Isthmus of Panama (Archivo General de Indias, 1521). Although there were initially adaptation difficulties in Darien, cattle were successfully established in areas such as Panama, Nata, and Remedios. This facilitated the consolidation of cattle ranching in these regions as a key economic resource that supported colonial expansion into Central and South America (Villalobos-Cortés et al., 2009). Techniques such as microsatellite and single-nucleotide marker analysis have been used for the genetic characterization of Creole breeds, such as Guaymi and Guabala. These breeds stand out for their genetic uniqueness and contribution to global biodiversity (Martínez et al., 2012; Ginja et al., 2013; Villalobos-Cortés et al., 2023). In 2019, studies using uniparental markers confirmed the Iberian heritage, both from Spain and Portugal, and the influence of African breeds, such as Gabou and Bafatá in Guinea-Bissau, Muturú in Nigeria, and the Egyptian areas of Baladi and Menoufis. These findings provide a more complete context for understanding livestock population migration and gene introgression in the New World (Ginja et al., 2019). Morphometric studies have also demonstrated the adaptability and productive characteristics of these breeds. In particular, the Guaymi breed exhibited morphological homogeneity, which is indicative of high consanguinity. However, this breed has a remarkable ability to adapt to tropical environments with limited forage and great potential to produce meat and dairy in dual-purpose production systems (Villalobos-Cortés et al., 2021). In many countries where advanced techniques, such as molecular or quantitative genetic testing, are not available, phenotypic analyses are the main methodology used to differentiate breeds and evaluate their productive capacity (Sierra, 2001). Morphometry is key to the differentiation and selection of animal biotypes with statistical validity (Holgado et al., 2015). This technique is especially useful for farmers because it allows the evaluation of each population’s response to selection standards and its relationship with productivity (Ramónez Cárdenas et al., 2017). Morphometric indices, such as height and hip width, are relatively easy to measure and provide valuable skeletal development indicators. Because skeletal development is progressive and slow, these measurements allow accurate growth curves to be drawn (Cerqueira et al., 2013). The use of descriptive statistics in the morphometric evaluation of domestic animals has facilitated the over time characterization of indigenous and Creole populations. Some examples include the Pyrenean breed in Spain, where differences were identified according to geographical origin, the morphometric variability of the Creole biotypes of Loja in Ecuador, and the analysis of the body conformation of the Argentine Creole (Martínez et al., 1998; Pastor et al., 2000; Aguirre-Riofrio et al., 2019). Morphological analysis, which refers to the external appearance of an organism, makes it possible to determine the degree of homogeneity or heterogeneity between individuals of a population or race (Griffin, 1962). Morphological variables are essential for this analysis (Canales, 2014). Phaneroptic traits, which are determined by qualitative variables, are also useful for the characterization of populations because they cover aspects such as the skin, wool color, horns, nails, and hooves, which can be analyzed using statistical techniques (Cevallos, 2012). In some studies, numerical taxonomy has been applied for the classification and characterization of Portuguese bovine breeds, highlighting this method as a key tool for studying morphological and phaneroptic traits (Sobral et al., 2002). The morphometric characterization of bovines with zoometric measurements and indices has been a tool traditionally used to evaluate the animals’ productive aptitude (meat or dairy). However, the application of morphometric characterization in the field may be limited by the need for expert interpretation, the variability in the criteria used, and the lack of accessible tools that systematize the calculations. Morphometric methods are based on standardized body measurements and derived indices that quantify the structural proportions of animals, providing objective size, conformation, and functional morphology descriptors. In bovines, these methods have been widely applied to breed characterization, phenotypic differentiation, and productive aptitude prediction, particularly for meat and dairy systems. Zoometric indices are especially valuable in tropical and low-input production systems, where access to molecular or advanced performance data is limited. Zoometric indices can be used to support selection, management, and conservation decisions based on easily obtainable field measurements (Sierra, 2001; Contreras et al., 2012; Aguirre-Riofrio et al., 2019). In this context, the development of interactive technological tools capable of automatically calculating and interpreting the main zoometric indices represents a significant advance. Incorporating a supervised learning model trained on previously classified morphometric data allows the objective and reproducible validation of the animals’ productive aptitude. This combination of web technologies and artificial intelligence provides an innovative solution for genetic selection and improvement, especially in tropical production systems where land breeds with high productive potential but little scientific characterization is used. This study aimed to compare the morphometry and morphology of the Guaymi and Guabala Creole cattle of Panama using zoometric indices and an automated approach based on the Mooebius application and a supervised learning model to classify productive aptitude and support conservation and genetic improvement programs. Based on scientific and conservation criteria, the analysis was intentionally limited to the Guaymi and Guabala breeds. These populations represent the two main Panamanian Creole cattle breeds under active conservation programs and exhibit well-defined and contrasting morpho-functional profiles, making them suitable for validating zoometric indices and supervised classification models. Materials and MethodsStudy areaThe data for this study were obtained from several herds of Creole cattle in nine regions of Panama. Samples of the Guabala breed were collected at the Ollas Arriba Research Subcentre in Capira, Panama Oeste (8° 48'13 "N, 79° 53'54" W); the Arenas de Mariato Experimental Station in Veraguas (7° 21,933'N, 80° 50,669'W); the Finca El Corotú, in Las Palmas, Veraguas (8° 08'14 "N, 81° 27'26" W); the Ganadera Guabala in Tole, Chiriqui (8° 12'51 "N, 81° 43'52" W); and the Finca Criollo Guabala in El Nancito, Remedios, Chiriqui (8° 13'00 "N, 81° 44'20" W). Samples of the Guaymi breed were collected at the Pacifico Marciaga Experimental Station in Coclé (8° 27,389'N, 80° 21,423'W); the Río Hato Sur Experimental Station in Antón, Coclé (8° 21,149'N, 80° 9,736'W); Calabacito Experimental Station in San Francisco, Veraguas (8° 14,709'N, 81° 4,765'W); Arenas de Mariato Experimental Station in Mariato, Veraguas (7° 21,933'N, 80° 50,669'W); and Carlos M. Ortega Experimental Station, in Gualaca, Chiriquí (8° 30.255'N, 82° 17.722'W). SamplingA comparative analysis of the Guabala and Guaymi populations in Panama was conducted. Both breeds correspond to Bos taurus (Creole cattle), a fact explicitly stated here to clarify their taxonomic classification. In the Guabala breed, the measurements included 100 individuals (80 females and 20 males) older than 2 years and in optimal body condition, without considering female parity. In the Guaymi population, 125 cattle were evaluated (98 females and 27 males) under the same criteria: age 3 years and good body condition, without considering the female reproductive history. The sample size was defined according to the FAO (2012) and ISAG guidelines for the phenotypic characterization of genetic resources of local and endangered animals, which recommend sampling at least 100 adult individuals per population whenever possible. The final number of animals included in this study reflects the actual availability of Guaymi and Guabala cattle within active conservation nuclei, while adult animals in good body condition were prioritized to ensure representative and reliable morphometric and morphological assessments. Age estimationThe age of the sample cattle was estimated from the records of each animal in the conservation centers and farms of collaborating producers. Morphological variablesThe morphological variables considered in this study cover various aspects of the bovine body and were analyzed to show qualitative differences between the Guaymi and Guabala breeds. The comparison of these variables allows the evaluation of morphological patterns associated with breed characteristics and their possible relationships with adaptation to local conditions. The head characteristics include the head profile (straight, concave, or convex); the presence, position, and direction of the horns (procero, ortocero, and opistocero); and the shape (bizco, brocho, cornipaso, cornivuelto, lira, playero, and rueda) and color of the horns (white, black and white, caramel, gray, and black). The ears’ direction (drooping, sloping, or horizontal) was also recorded. For the body, the variables included the dorsallumbar line (straight, saddled, or very saddled), buttock shape (straight, concave, or convex), and hump and dewlap presence or absence. Regarding the coat, the number of colors present (one color, two colors (overo), or more than two colors), their classification (intermixed or simple), and the predominant color (black, red, white, red, or bay) were considered, in addition to the coat type (long or short). Morphometric variables and indexesMorphometric data were gathered using a standard measuring tape and a zoometric rod, each with a precision of ±1 mm. The recorded measurements included live weight (LV), head width (HW), head length (HL), withers height (WH), crop height (CH), body length (BL), crop width (CW), crop length (CL), dorso-sternal diameter (DSD), bicostal diameter (BD), thoracic perimeter (TP), Cannon bone perimeter (CBP), and neck length (NL), following the methodology reported by various authors (Martinez et al., 2007; Martínez, 2008; Parés-Casanova, 2009; Herrera and Luque, 2009). Based on these measurements, four ethnological and five productive indices were calculated. The ethnological indices included: the Cephalic Index (CI), calculated as HW/HL) × 100; thoracic index (TI), calculated as BD/DSD) × 100; pelvic index (PI), calculated as CW/CL) × 100; and body index (BI), calculated as BL/TP) × 100. The productive indices included: the Dactyl-Costal Index (DCI), calculated as (CBP/BD) × 100; the dactyl-thoracic index (DTI), calculated as (CBP/TP) × 100; the transverse pelvic index (TPI), calculated as (CW/WH) × 100; the Longitudinal Pelvic Index (LPI), calculated as (CL/WH) × 100; the relative thoracic depth index (RTDI), calculated as (DSD/WH) × 100; and the lateral body proportionality index (LBPI), calculated as (WH/BL) × 100. This study also considered differences between genotypes (GR), with GR 1 representing Guaymi cattle and GR 2 representing Guabala cattle. The effect of sex (SX) was evaluated within each genotype group, with SX 1 referring to females and SX 2 to males. The aim of the analysis was to identify significant differences between the groups and between the sexes within each group. Statistical analysisNested analysis of variance (nested ANOVA) was chosen due to the hierarchical structure of the dataset, specifically to evaluate morphometric differences between sexes nested within each breed (Guaymi and Guabala). This modeling approach was selected to assess within-breed sexual dimorphism while accounting for breed-specific structure rather than testing crossed genotype × sex interactions. The inclusion of interaction terms could lead to unstable estimates and limited interpretability, given the unbalanced sex distribution across breeds. This approach enabled the precise detection of sex-specific differences within breeds, thereby enhancing the reliability of comparative assessments. Morphometric data underwent rigorous preprocessing before analysis, during which records with missing values or inconsistencies were systematically excluded. This step ensured accuracy, minimized biases, and maintained high-quality standards for statistical analyses. The applied statistical model was defined as follows: Y ijk=μ + α i + β j (i) + ϵ ijk where,
Statistical analysis was performed using the statsmodels package in Python, which allowed us to adjust the model and generate ANOVA tables for each variable. Statistics and 𝑝 values were calculated as follows:
Variables with p values <0.05 were considered statistically significant. For the analysis of the morphological variables, preprocessing was performed, within which the records with frequency values equal to zero in any of the comparison columns (Guaymi and Guabala) were eliminated to avoid biases in the later analyses. The statistical analysis focused on comparing the frequency distributions between the variables of the Guaymi and Guabala breeds. For this purpose, the chi-square test was used, implemented through the scipy.stats library’s chi2_contingency function. The categories of each characteristic were independently evaluated, and contingency tables were constructed for each of them. The analysis results included the chi-square statistic, the degrees of freedom, and the associated p values. Statistically significant differences were those characteristics for which the p-value was 0.05. The maximum values in each group (Guaymi and Guabala) were identified for the characteristics with significant differences. In this step, the specific category with the highest frequency in each group was determined along with the frequency value. These results were tabulated, and the categories and corresponding values for each significant characteristic were presented. Supervised machine learning applicationObjectiveFor the supervised classification of bovine productive aptitude (meat or dairy), 10 zoometric indices were selected for their proven discriminating power between functional biotypes, as documented in specialized zootechnical studies (Vargas-Villamil and Tedeschi, 2014; Ríos-Utrera et al., 2018). Animals were assigned to meat or dairy categories according to established cut-off points derived from these indices, validated in tropical cattle morphometric research (Contreras et al., 2012; Rojas-Espinoza et al., 2023). Predictor sets and preprocessingThe main predictive model used 12 raw morphometric measurements. Three auxiliary ablation models were designed for redundancy and robustness assessment: Data preprocessing included the following
The variables were left unscaled for the main random forest model; standardization was applied only in supplementary analyses. Model development and validationWe tuned the random forest classifiers (n estimators=300; class weight=balanced; random state=42) for max_depth, min_samples_leaf, and min_samples_split via nested grid search within stratified cross-validation. Three evaluation strategies were implemented: Performance assessmentThe evaluation metrics included accuracy, macro-precision, macro-recall, and macro-F1, reported as mean ± SD for CV and LOLO, and as point estimates for the hold-out set. Additional outputs included:
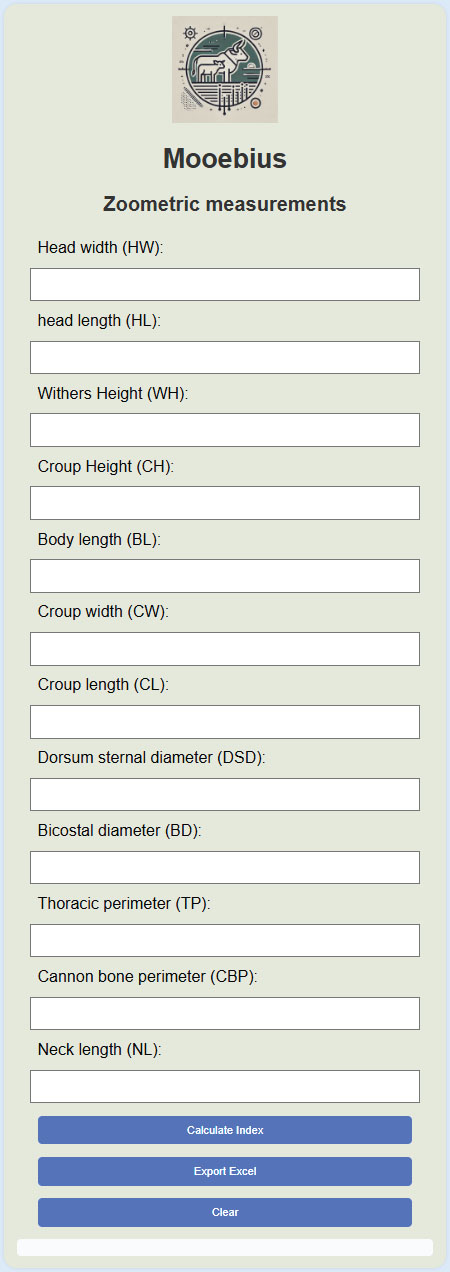
Circularity controlBecause the target label was derived from indices calculated from the same morphometric measurements that could serve as predictors, excluding indices from the main predictive model mitigated circularity. Indices were used only in the ablation models for interpretative purposes. Mooebius web applicationDefinition of the target variableThe categorical target variable, productive aptitude (meat/milk), was predetermined using a fixed zootechnical rule integrating five classical indices: CI, TI, PI, BI, and DCI. Each index was assessed against its literature-defined threshold for FBD. A weighted scoring scheme was applied (CI, BI: weight=2; TI, PI, DCI: weight=1) to generate composite scores for meat and milk. The assigned class corresponded to the higher score; ties or indeterminate cases were classified as undefined. This reference label was fixed and independent of any model output to ensure consistency. Predictor sets and preprocessingTo facilitate and standardize the interpretation of zoometric indices in the field, we developed a web application called Mooebius (Fig. 1), an interactive digital tool developed for bovine morphometric evaluation that focuses on the prediction of productive aptitude (dairy or meat) from basic body data. This system is implemented in a web environment using standard technologies such as Hyper Text Markup Language , Cascading Style Sheets, and JavaScript, allowing it to function on multiple devices such as cell phones or tablets. The program is structured in three main components: (1) a form for capturing 12 morphometric measurements of the animal; (2) an automatic calculation module of 10 fundamental zoometric indices, such as the CI, TI, PI, BI, and DCI, among others, all of which are evaluated according to functional ranges defined in the zootechnical literature; and (3) a scoring system weighted on five key indices, the sum of which allows a final classification of the animal as a “meat” or “dairy” producing animal.
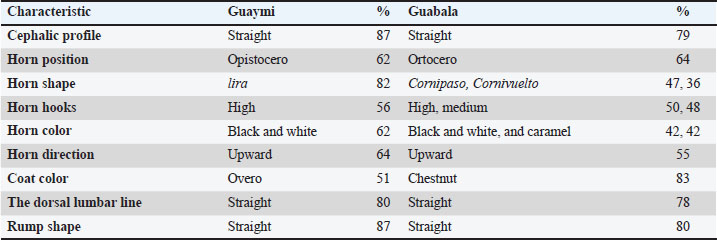
Fig. 1. Image sample from the Mooebius program. The results are presented in a color-coded pivot table, which facilitates visual interpretation. In addition, Mooebius exports the generated data into Excel format (.xlsx), which favors its application in field environments, laboratories, and technical documentation processes. Mooebius was used as a support tool to evaluate the morphometric data and validate the predictions generated by the random forest supervised classification model, confirming its usefulness as a complementary instrument in the processes of productive diagnosis and phenotypic selection. The combined use of the Mooebius app and the predictive model represents a methodological innovation that strengthens decision-making in the selection and genetic improvement of Creole breeds, especially in contexts of limited resources where traditional tools may not be available. Ethical approvalNot needed for this study. However, measurements were made to assess animal welfare and health in Panama. ResultsMorphological traitsPredominant traits have been reported to facilitate comparative interpretation; however, they do not preclude the presence of less frequent phenotypes within each population. Comparative analysis of the morphological characteristics of the Guaymi and Guabala cattle breeds revealed significant differences (p < 0.05) in several qualitative variables (Table 1). These differences highlight the clear phenotypic differentiation between the two breeds and reflect the underlying genetic and environmental influences. Table 1. Morphological characteristics of the Guaymi and Guabala breeds.
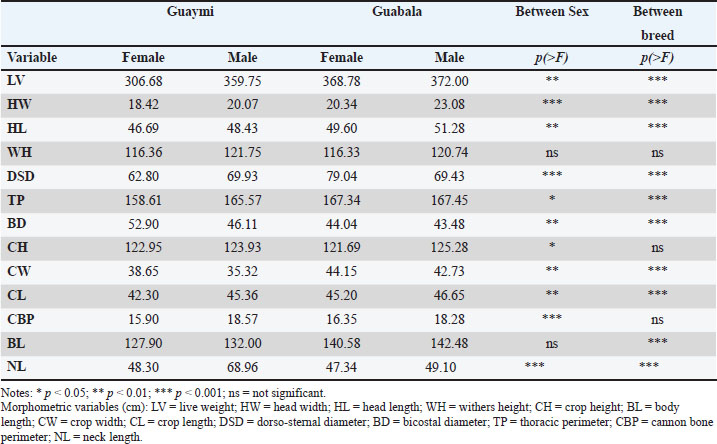
Regarding the cephalic profile, a predominance of the straight profile was observed in both breeds, although with a greater frequency in Guaymi (70%) than in Guabala (50%). Genetics and artificial selection factors may influence this trait, as has been documented in studies of morphological diversity in local bovine breeds (Casanova et al., 2012). Similar findings have been reported in bovine breeds native to Indonesia, where the head conformation varies significantly depending on the breed and adaptation to the environment (Setiaji, 2023). The horn position showed notable differences. In the Guaymi breed, the opistocero position was predominant (55%), whereas the ormocers position was more common (42%). Regarding the shape of the horns, Guaymi presented greater homogeneity with a predominance of the lira shape (68%), whereas greater diversity was observed in Guabala, with a balanced distribution between the cornipaso (47%) and cornivuelto forms (36%). This diversity in horn morphology could be related to local adaptations or selection preferences by breeders, as noted in studies on morphological variability and bovine functionality (Çakar et al., 2024). Recent research on bovine breeds from the Tarai region of India has shown that these differences may influence breed conservation and functionality within local production systems (Mohan et al., 2025). Another relevant aspect is the variation in the horn hooks. La Guaymi presented a greater proportion of high hooks (47%), whereas similar frequencies were found between high and medium hooks in Guabala (50% and 48%, respectively). Regarding the color of the horns, Guaymi showed greater uniformity, with a predominance of black and white (54%), whereas in Guabala, an equitable distribution was observed between black and white (42% each). Regarding the color of the coat, a predominance of the two-color pattern (overo) was observed in Guaymi (51%) and Guabala (83%). These differences may be related to inheritance patterns and phenotypic selection (Felius et al., 2011). A recent study in Türkiye cattle proved that the variation in cranial morphology between native and nonnative breeds is directly related to selective pressures and domestication patterns (Gündemir et al., 2025). For dorsal lumbar line and rump shape, both breeds showed a predominance of straight conformation (Table 1). Nevertheless, concave and ensaddled dorsal profiles, as well as alternative rump shapes, were also present, indicating that neither population is morphologically uniform. Overall, beyond the identification of predominant morphological traits summarized in Table 1, substantial intrabreed variability was detected across all evaluated qualitative characteristics. Several traits exhibited several expression levels, with frequencies distributed among multiple phenotypic categories. This intrabreed variability reflects the underlying phenotypic diversity of Guaymi and Guabala cattle and is consistent with their classification as Creole under conservation programs. Morphometric measurementsThe comparative analysis of the morphometric variables between the Guaymi and Guabala Creole breeds revealed significant differences in several measures, revealing both the genetic influence and the effect of sex on the variation in these characteristics (Table 2). Table 2. Morphometric variables by breed and sex between the Guaymi and Guabala breeds.
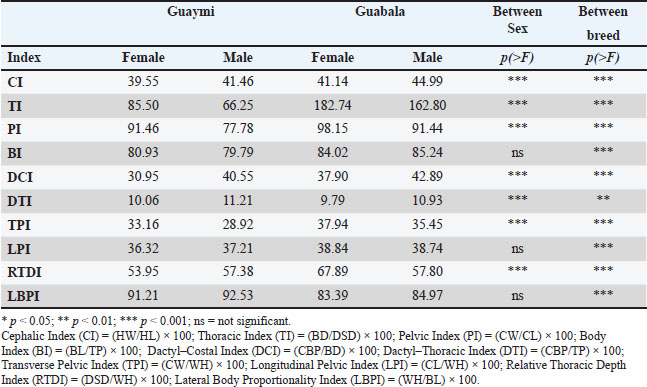
The most notable differences between breeds were observed in: HW (p < 0.0001), HL (p < 0.0001), CW (p < 0.0001), and DSD (p < 0.0001). Similarly, significant differences between sexes were observed within breeds in these same variables (p < 0.0001), consistent with the influences of hormonal or functional factors influence the expression of certain morphometric traits. In contrast, WH and CH did not significantly differ between breeds (p > 0.05), although the effect of sex within breeds was relevant (p <0.05). BL differed between breeds (p <0.0001) but not between sexes within breeds (p=0.212). On the other hand, CBP did not significantly differ between groups (p=0.168) but did significantly differ between sexes (p < 0.0001), highlighting the influence of sexual dimorphism in certain morphometric measurements. Zoometric indicesThe ANOVA of the zoometric indices of the Guaymi and Guabala Creole cattle in Panama allowed the identification of significant differences between the genetic groups and, to a lesser extent, between the sexes within each group (Table 3). These findings provide quantifiable evidence of the morphometric variability existing in these populations and contribute to the phenotypic characterization of Creole breeds in the region. Table 3. Comparison of zoometric indices by sex and between the Guaymi and Guabala breeds.
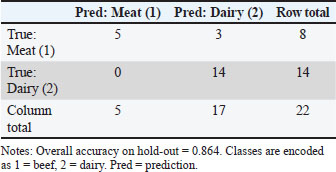
The CI, TI, PI, and BI indices reflect the general anatomical proportions of the animals and are essential for differentiating biotypes within a bovine population (Martínez et al., 2012). In this study, CI was significantly different between the genetic groups (p < 0.0001) and between the sexes within each group (p < 0.0001), suggesting that the shape and proportions of the head vary between Guaymi and Guabala, as well as between males and females within each breed. CI is a key marker for differentiating Creole breeds in Latin America, where variations related to ecological adaptation and selection pressure have been found (Ginja et al., 2013; Holgado et al., 2015). One of the most significant differences (p < 0.0001) was observed for TI, indicating that the depth and width of the thorax vary considerably between genotypes. This finding is consistent with that reported by Aguirre-Riofrio et al. (2019) in Creole cattle from Ecuador, where thoracic capacity was associated with adaptation to extensive grazing conditions and variable forage availability. However, PI was significantly different between the genetic groups (p < 0.0001) and between the sexes (p < 0.0001). This result differs from those of previous studies that have indicated that the variability in pelvic morphology in Creole breeds is usually less pronounced due to the natural selection stability in these populations. However, such variability was observed in this case (Contreras et al., 2011). Finally, BI was significantly different between the genetic groups (p < 0.001) but not between the sexes within each group (p=0.4846), indicating that the relationship between body length and thoracic perimeter is a determining feature in the differentiation of these breeds but not between the sexes. This index has been used to evaluate the productive aptitude and conformation of the breed biotype in Creole cattle of the Iberian Peninsula (Pastor et al., 2000). Productive indices, including DCI and the DTI, TPI, LPI, relative depth of the thorax RTDI, and LBPI indices, provide key information on the functional performance of cattle in terms of meat and dairy production. In this study, DCI and DTI showed statistically significant differences (p < 0.0001 and p < 0.001, respectively) between groups and between sexes (p < 0.0001 and p < 0.0001, respectively), suggesting that the proportions of the extremities in relation to the rib cage can be useful for selecting animals with better aptitudes for traction and resistance under extensive conditions (Cerqueira et al., 2013). Highly significant differences (p < 0.0001) were seen for TPI, which supports the hypothesis that these breeds have a differentiated morphometry that influences their reproductive ability and environmental adaptation. These results are consistent with those of Villalobos-Cortés et al. (2021), who reported that the pelvic and thoracic configurations of Panamanian Creole cattle resemble those of breeds adapted to tropical environments with limited forage availability. On the other hand, LPI was significantly different (p < 0.0001), indicating that the relationship between the length of the croup and the height of the withers varies between the breeds. This variation could be due to differences in the structural selection pressure exerted on each breed (Ramónez-Cárdenas and Zhunio-Samaniego, 2017). Thoracic index and BI showed high discriminative capacity between breeds and were consistently associated with differences in body conformation related to productivity. DiscussionBreed differentiation and productive aptitudeThese results indicate that the Guaymi breed exhibits greater morphological uniformity, whereas the Guabala breed shows greater phenotypic variability, especially in the shape and hooks of the horns. This contrast could be relevant in selection and genetic improvement programs because uniformity is valuable for commercial production, whereas diversity can represent adaptive advantages in different production environments (Gautier et al., 2010). In addition, recent studies have shown that genetic differences in bovine populations can impact both the breeds’ morphology and adaptive capacity (Lozada-Soto et al., 2022; Petrov and Kamaldinov, 2024). The ethnological and productive zoometric indices of the Guaymi and Guabala cultivars were analyzed to classify the predominant aptitudes of each population, revealing important functional and productive differences between the two populations. With respect to CI, both genotypes were classified as dolichocephalic, which is associated with meat-producing aptitude. This pattern was more pronounced in the Guaymi breed (39.98), and although the value in Guabala (41.91) was higher, both remained within the typical ranges of animals with long and narrow heads, typical of meat phenotypes (Contreras et al., 2012). In both breeds, TI reflected an elliptical thoracic conformation, with values lower than the cut-off point (Guaymi: 81.2; Guabala: 58.4). This conformation is associated with dairy-producing animals, suggesting that respiratory capacity and adaptability, which are key factors for dairy production under extensive conditions, are specializations (Rojas-Espinoza et al., 2023). PI was comparable to that of dolichopelvic animals (Guaymi: 88.39; Guabala: 96.81) and was more pronounced in the Guabala breed. This index is associated with pelvises that are longer than those that are wide, which is typical of dairy cows with greater calving efficiency, which supports their dairy-producing aptitude. In contrast, BI, which reflects the relationship between body length and thoracic perimeter, was greater in the Guabala breed (84.26) than in the Guaymi breed (80.68), indicating a more robust morphology in Guabala, although both breeds fall within the brevilinear classification, which is characteristic of meat-producing animals. Recent studies in Maharashtra cattle have revealed similar variations in morphometric parameters, consistent with a strong influence of natural selection and environmental adaptation, which highly influence these traits (Thalkar et al., 2016). Similarly, a study in Karnataka identified physical and morphometric characteristics in local breeds, highlighting the importance of phenotypic diversity in maintaining genetic biodiversity (Mundinamani et al., 2024). These differences may be relevant for breed characterization and selection in genetic improvement programs. The morphometric diversity between breeds can be influenced by genetic and functional factors, as reported in studies on cranial morphometry in bovines (Çakar et al., 2024). These results are important for the conservation and sustainable use of these land breeds. Their adaptation to local conditions and genetic variability makes them valuable resources for agricultural biodiversity (Felius et al., 2011). Therefore, implementing management and conservation strategies is essential to avoid the substitution of these breeds for exotic livestock with a lower degree of adaptation (Gautier et al., 2010). Differences in the productive indices were detected as follows: The DCI and DTI values were low in both breeds, with this decrease being more significant in Guaymi, indicating lower bone strength and reaffirming its dairy-producing orientation. Guaymi had a lower TPI value than the cut-off point (32.21), indicating less muscular development, whereas Guabala had a higher TPI value than the cut-off point (37.44), indicating better meat-producing ability. The LPI of Guaymi was within the dairy range (<37), while that of Guabala exceeded this value (38.82), indicating greater robustness and body development, which are favorable aspects for meat production. The RTDI, which is related to thoracic depth, was greater in Guabala (65.87), indicating greater lung capacity, which is related to better performance in meat production. Finally, the LBPI, which shows body proportionality, was higher in Guaymi (91.51), reflecting a more efficient conformation for dairy production, while Guabala had a lower value (83.71), which is typical of meat-producing animals (Mejía and Sebastián, 2015). These results indicate that the Guaymi breed has a morphology more adapted to dairy production, with lighter and more proportionate structures, whereas the Guabala breed shows greater robustness and muscular development, which are characteristic attributes of animals oriented toward meat production. This phenotypic differentiation coincides with earlier studies on biotypes adapted for different productive purposes in tropical production systems (Contreras et al., 2012; Rojas-Espinoza et al., 2023). Performance of the machine learning modelPerformance of the stratified hold-out set. The main random forest model, trained using all 12 raw morphometric measurements, achieved an overall accuracy of 86.4 % on the stratified 10 % hold-out set (n=22), despite a lower mean accuracy during 5-fold cross-validation (0.701 ± 0.101). As shown in Table 4, the classification performance was asymmetric between classes. Predictions for the dairy category were highly reliable, with a recall of 1.00 (14/14 correctly classified) and a precision of 0.824. Conversely, the meat category exhibited a recall of 0.625 (5/8 correctly classified) but a perfect precision of 1.00, indicating that the model was conservative in predicting meat, avoiding false positives (no dairy samples were misclassified as meat). Table 4. Confusion matrix on the stratified 10% hold-out (main model: 12 raw measurements).
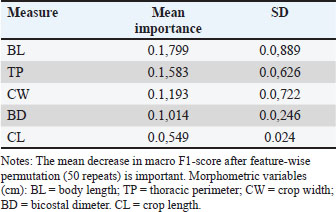
Table 5. Permutation importance (Top 5) on the main model’s hold-out set (12 raw measurements).
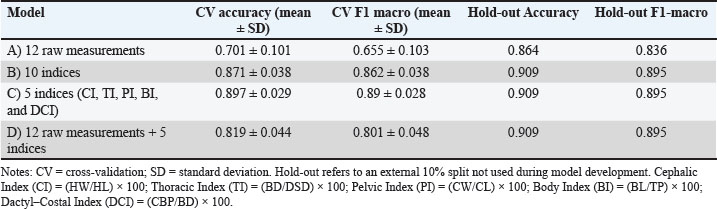
All misclassifications involved meat samples being labeled as dairy (37.5% of meat cases), indicating a tendency to favor dairy predictions. This pattern is consistent with the macro-precision (0.912) exceeding the macro-recall (0.812), reflecting a bias toward correct positive predictions at the expense of meat sensitivity. The absence of false positives for Meat underscores the specificity of the model for this class, albeit with reduced sensitivity. The superior performance on the hold-out set relative to cross-validation may be attributable to sampling variability, given the small evaluation size, as well as potential alignment between the hold-out distribution and patterns learned during training. Notably, the use of raw measurements—rather than derived indices—likely reduced information redundancy and circularity, making the classification task more challenging during cross-validation while still yielding robust generalization on unseen data. In addition to the main model based on raw morphometric data, auxiliary models trained with derived zoometric indices achieved higher accuracies of up to 90.9%, as shown in Table 6. Table 6. Ablation study comparing feature sets (Random Forest; 5-fold stratified cross-validation and 10% stratified hold-out).
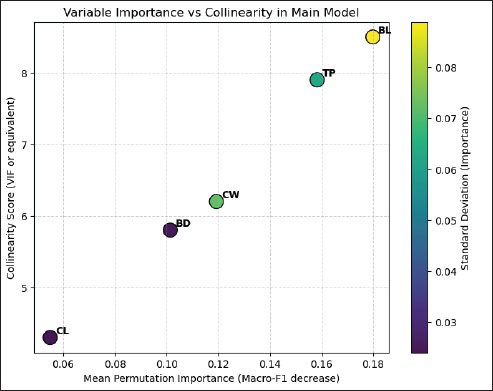
However, these models presented circularity because the predictors were partially derived from the same indices that define the target variable. Therefore, despite a slightly lower numerical performance, Model A (12 raw measurements) was considered the reference model due to its biological validity and independence from circular features. This trade-off highlights that morphometric information alone can capture functional aptitude patterns consistent with traditional zoometric evaluation when processed through supervised machine learning. The model was trained exclusively on the training portion and tuned via cross-validation, while the hold-out set was reserved for a single, unbiased performance assessment. This approach is widely adopted in machine learning research to prevent overfitting and simulate the behavior of the model on unseen data (Sun et al., 2025; Zhang et al., 2025). The stratified hold-out is particularly recommended in classification tasks with imbalanced classes because it reduces the risk of misleading performance metrics due to unequal class representation (Almuzadid and Subhiyakto, 2025; Chen et al., 2025). Training with raw measurements, rather than with defining indices, made the task more demanding, reducing circularity but lowering cross-validation performance. Nonetheless, the performance on the unseen set remained high. Given the limited sample size in the hold-out set, statistical confidence intervals for these metrics are relatively wide; therefore, performance differences between cross-validation and hold-out evaluation should be interpreted with caution until validated on larger, independent datasets. Variable importance and collinearityPermutation-based importance analysis on the hold-out set revealed that a limited subset of thoracic and pelvic measurements accounted for most of the model’s predictive performance in terms of macro-F1 (Table 5). BL was identified as the most influential trait (mean importance=0.1799, SD=0.0889), followed by thoracic perimeter (TP; 0.1583 ± 0.0626). Chest width, BD, and CL showed lower contributions but still provided measurable gains in predictive accuracy. The relatively high variability observed in BL importance is consistent with sensitivity to data partitioning, a behavior previously reported in morphometric trait-based classification models (Immaculate et al., 2020). Collinearity assessment using correlation matrices and variance inflation factors confirmed the presence of well-defined clusters of highly correlated traits, particularly among thoracic and pelvic measurements. This structural redundancy supports the use of ablation tests to evaluate whether a reduced set of variables can be used to preserve predictive performance, thereby improving model parsimony. Similar approaches have shown that removing redundant morphometric traits can enhance model generalizability without substantial loss of accuracy (Gizaw et al., 2007; Yakubu et al., 2011). Figure 2 integrates permutation-based feature importance and collinearity metrics, allowing simultaneous visualization of each trait’s predictive contribution and redundancy. Traits located in the upper-right quadrant, such as BL and TP, combine high predictive relevance with high collinearity, illustrating the trade-off between retaining informative variables and minimizing redundancy. This pattern supports targeted variable selection strategies, particularly for field-based applications where reducing the number of measurements can lower operational costs while maintaining predictive performance.
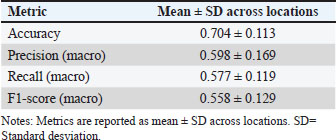
Fig. 2. Variable importance versus collinearity in the main model. In addition to individual morphometric traits, the TI and BI have emerged as key zoometric parameters for the differentiation of Guaymi and Guabala cattle and the assessment of their productive potential. These indices integrate multiple structural dimensions related to body capacity and skeletal development, providing a functionally meaningful summary of animal conformation. These results offer a phenotypic basis for supporting conservation strategies and genetic improvement programs in Panama. Future studies combining zoometric indices with genomic information and performance data under production conditions would further strengthen the understanding of the functional value of these Creole populations within sustainable production systems. Table 6 summarizes the ablation results obtained using a consistent partitioning scheme. Models B and C outperformed A due to their direct use of the index feature space that defines the target variable—simplifying the task but introducing circularity. Model D provided no hold-out gains and reduced CV performance, denoting redundancy from the combination of raw measurements with derived indices. Model A was selected as the baseline for its optimal trade-off between the biological validity and generalization performance. Geographic validation (Leave-one-location-out)For Model A, the mean across locations was accuracy 0.704 ± 0.113, macro-precision 0.598 ± 0.169, macro-recall 0.577 ± 0.119, and macro-F1 0.558 ± 0.129. The site-level performance ranged from 0.556 to 0.889 and was influenced by the sample size and local biotype distribution. These results indicate moderate spatial generalization and highlight the need for broader sampling in locations with small n or class imbalance and for future models incorporating site effects (Table 7). Table 7. Geographic validation (Leave-One-Location-Out, LOLO) of the main model (12 raw measurements).
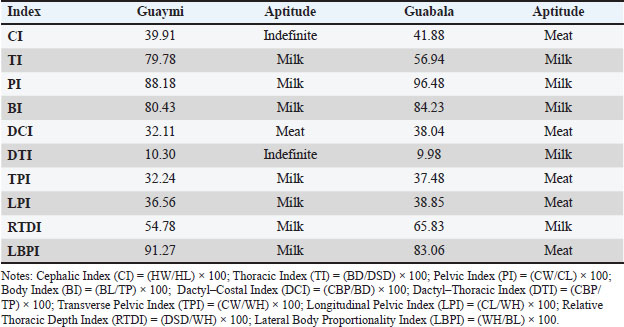
Applications and implications for conservationThe Random forest (RF) supervised classification model, trained with morphometric variables and weighted zoometric indices, showed outstanding performance in predicting cattle productive aptitude (Table 8). The target classes were “meat” and “dairy,” defined through a weighted scoring system applied to five key zoometric indices (CI, TI, PI, BI, and DCI). These indices were selected based on their functional relevance in productive characterization and their documented discriminative capacity between bovine biotypes in morphostructural studies of creole cattle populations (López-Aguirre et al., 2023; Ccalta et al., 2025). Table 8. Classification of the Guaymi and Guabala breeds based on suitability (meat or dairy) using Mooebius and supervised learning.
Numerous studies have endorsed ensemble algorithms, particularly random forests, in zootechnical and genetic contexts because of their ability to manage multivariate data, resistance to overfitting, and superior performance compared with traditional classifiers (Mahmud et al., 2021; Chafai et al., 2023). RF applications have been successful in predicting dairy production, carcass quality, and breed and phenotypic characterization (Fuentes, 2022; Gebreyesus et al., 2023). To confirm and automate the classification of productive aptitude (dairy or meat) in Creole cattle from the 10 zoometric indices generated, a supervised learning model was trained using the random forest classifier algorithm (Table 8). This approach has previously been used to classify animals according to performance and morphometry, with good precision (Gebreyesus et al., 2023; Kasarda et al., 2023). This method offers a practical tool to support phenotypic selection in Creole and Mestizo cattle production systems. In particular, the implementation of this approach within the Mooebius system facilitates the integration of AI in tropical livestock, providing concrete benefits in terms of the objectivity, speed, and reproducibility of the selection and genetic improvement processes (Bovo et al., 2021; Kasarda et al., 2023). Future research should explore the development of machine learning-based body weight prediction models stratified by breed and sex using expanded and more balanced morphometric datasets. Such approaches would allow a more refined assessment of sexual dimorphism and breed-specific growth patterns, thereby enhancing the applicability of morphometric tools for precision management, conservation, and genetic improvement of Creole cattle populations. ConclusionThis study presents the first comprehensive morphometric and morphological comparison between the Guaymi and Guabala Creole cattle breeds of Panama, integrating classical zoometric analysis with automated classification through the Mooebius web application and a supervised random forest model. The results revealed marked phenotypic differentiation: Guaymi cattle displayed lighter, more proportionate morphologies associated with dairy aptitude, whereas Guabala cattle exhibited greater robustness and muscular development, traits typical of meat-oriented biotypes. Significant differences in key morphometric variables and zoometric indices underscore the value of these breeds as well-adapted to tropical production systems. The strong predictive performance of the automated model demonstrates its applicability as a practical decision-support tool for phenotypic selection, conservation, and genetic improvement, particularly in regions with limited access to advanced genomic technologies. Overall, these findings highlight the need to preserve the genetic diversity of Creole breeds, not only for their historical and cultural heritage but also for their proven adaptability to local environmental challenges. Future research should combine genomic analysis with on-farm performance evaluations to reinforce breeding programs in tropical livestock systems that balance conservation priorities with productivity goals. AcknowledgmentsWe thank the Faculty of Veterinary Medicine of the University of Panama for the support provided in this research, as well as the IDIAP field technicians who supported the collection of samples from the farms and producers of the Association of Creole Breeders of Panama (ACCRIPA), to the Conbiand Network and the Ibero-American Network on Zoogenomic Resources and their Resilience (REZGEN-IBA, 123RT0139), for their support of this research. Conflicts of interestThe authors have no conflicts of interest to declare. FundingThis work was financed by the Institute of Agricultural Innovation of Panama (IDIAP), the National Secretariat of Science, Technology and Innovation (SENACYT), and the National System of Researchers (SNI). Authors’ contributionsAxel Villalobos-Cortes: Conceptualization, methodology, validation, investigation, formal analysis, and preparation of the original draft. Ana Flores De Leòn: investigation writing—review and editing. Marcelino Jaen: validation and supervision. Data availabilityAll data supporting this study’s findings are available from the corresponding author upon reasonable request. AI transparency statementAI-assisted tools were used solely for language editing and formatting, while the authors performed all analyses, interpretations, and conclusions. ReferencesAguirre-Riofrio, E.L., Abad-Guamán, R.M. and Uchuari-Pauta, M. 2019. Morphometric evaluation of phenotypic groups of southern Ecuadorean creole cattle. Diversity 11(12), 221; doi:10.3390/d11120221 Almuzadid, G. and Subhiyakto, E.R. 2025. Stroke risk classification using the XGBoost and random forest ensemble learning method. J. Appl. Inf. Comput. 9(1), 20–31. Archivo General de Indias. 1521. Despacho para la ciudad de Panamá. Portal de Archivos Españoles. Arrieta-González, A., Hernández-Beltrán, A., Barrientos-Morales, M., Martínez-Herrera, D.I., Cervantes-Acosta, P., Rodríguez-Andrade, A. and Dominguez-Mancera, B. 2022. Caracterización y tipificación tecnológica del sistema de bovinos doble propósito de la Huasteca Veracruzana México. Rev. MVZ. Córdoba. 27(2), e2444; doi: 10.21897/rmvz.2444 Bovo, M., Agrusti, M., Benni, S., Torreggiani, D. and Tassinari, P. 2021. Random forest modeling of dairy cow milk yield under heat stress conditions. Animals 11(5), 1305; doi:10.3390/ani11051305 Çakar, B., Tandir, F., Güzel, B.C., Bakıcı, C., Ünal, B., Duro, S., Szara, T., Spataru, C., Spataru, M.C. and Gündemir, O. 2024. Comparison of skull morphometric characteristics of Simmental and Holstein cattle. Animals 14, 2085; doi:10.3390/ani14142085 Canales, A. 2014. Caracterización genética y morfológica de vacas de la raza Criollo-Lechero Tropical. MSc thesis, Universidad Veracruzana. Casanova, P.M.P.I., Sinfreu, I. and Villalba, D. 2012. Principal component analysis of cephalic morphology for the classification of some Pyrenean cattle. Anim. Genet. Resour. 50, 59–64; doi:10.1017/S2078633611000385 Ccalta, R., Huayta Arizaca, R., Salcedo Quispe, E., Valverde, A., Cucho Dolmos, H., Canaza-Cayo, A., Acuña Leiva, A. and Estrada, R. 2025. Morphometric characterization and zoometric indices of High-Andean creole cows from Southern Peru. Vet. Sci. 12(8), 782; doi:10.3390/vetsci12080782 Cerqueira, J., Araújo, P., Vaz, P.S., Cantalapiedra, J., Blanco, P. and Niza, R. 2013. Relación entre medidas zoométricas en vacas Holstein-Friesian y dimensiones de cubículos en granjas lecheras. Int. J. Morphol. 31(1), 55–63; doi:10.4067/S0717-95022013000100008 Cevallos, F.O. 2012. Caracterización morfoestructural y faneróptica del bovino criollo en la provincia de Manabí. MSc thesis, Universidad de Córdoba. Chafai, H., Hayah, E., Houaga, I. and Badaoui, B. 2023. Machine learning models applied to genomic prediction in animal breeding. Front. Genet. 14, 1150596; doi:10.3389/fgene.2023.1150596 Chen, Z., Dazard, J.E., Salerno, P.R.V.D.O., Sirasapalli, S.K., Makhlouf, M.H., Rajagopalan, S. and Al-Kindi, S. 2025. Composite socio-environmental risk score for cardiovascular assessment: an explainable machine learning approach. Am. J. Prev. Cardiol. 15, 100650; doi:10.1016/j.ajpc.2025.100650 Contreras, G., Chirinos, Z., Molero, E. and Paéz, A. 2012. Medidas corporales e índices zoométricos de toros Criollo Limonero de Venezuela. Zootec. Trop. 30(2), 175–181. Contreras, G., Chirinos, Z., Zambrano, S., Molero, E. and Páez, E. 2011. Caracterización morfológica e índices zoométricos de vacas Criollo Limonero de Venezuela. Rev. Cient. FCV-LUZ 21(1), 91–113. Cuevas-Reyes, V. and Rosales-Nieto, C. 2018. Caracterización del sistema bovino doble propósito en el noroeste de México: productores, recursos y problemática. Rev. MVZ. Córdoba. 23(1), 6448–6460; doi:10.21897/rmvz.1240 FAO. 2012. Phenotypic characterization of animal genetic resources. Rome, Italy: FAO . Felius, M., Koolmees, P.A., Theunissen, B., European Cattle Genetic Diversity Consortium. and Lenstra, J.A. 2011. On the breeds of cattle—historic and current classifications. Diversity 3, 660–692; doi:10.3390/d3040660 Fuentes., S. and et al. 2022. Animal biometric assessment using non-invasive computer vision and machine learning as predictors of age and welfare of dairy cows: the future of automated veterinary support systems. J. Agric. Food Res. 10, 100388; doi:10.1016/j.jafr.2022.100388 Gautier, M., Laloë, D. and Moazami-Goudarzi, K. 2010. Insights into the genetic history of French cattle from dense SNP data on 47 worldwide breeds. PLos One. 5(9), e13038; doi:10.1371/journal.pone.0013038 Gebreyesus, G., Milkevych, V., Lassen, J. and Sahana, G. 2023. Supervised learning techniques for predicting the body weight of dairy cattle from 3D digital images. Front. Genet. 13, 947176; doi:10.3389/fgene.2022.947176 Ginja, C., Gama, L.T., Cortes, O., Delgado, J.V., Dunner, S., García, D., Landi, V., Martín-Burriel, I., Martínez, A., Penedo, C.T., Rodellar, C., Zaragoza, P., Cañon, J. and Consortium, B. 2013. Conservation priorities of Iberoamerican cattle based on autosomal microsatellite markers. Genet. Sel. Evol. 45(1), 35; doi:10.1186/1297-9686-45-35 Ginja, C., Gama, L.T., Cortés, O., Martín-Burriel, I., Vega-Pla, J.L., Penedo, C., Sponenberg, P., Cañón, J., Sanz, A., Do Egito, A.A., Alvarez, L.A., Giovambattista, G., Agha, S., Rogberg-Muñoz, A., Lara, M., Delgado, B.C. and Martínez, A. 2019. Genetic ancestry of American creole cattle inferred from uniparental and autosomal genetic markers. Sci. Rep. 9, 11486; doi:10.1038/s41598-019-47636-0 Gizaw, S., Van Arendonk, J.A.M., Komen, H., Windig, J.J. and Hanotte, O. 2007. Population structure, genetic variation, and morphological diversity in Ethiopian indigenous sheep. Anim. Genet. 38(6), 621–628; doi:10.1111/j.1365-2052.2007.01659.x Griffin, D.R. 1962. Estructura y función animal. Compañía Editorial Continental, México City, Mexico. Gündemir, O., Manuta, N., Güzel, B.C. and Bakıcı, C. 2025. Skull morphology in native and non-native cattle breeds in Türkiye. J. Anim. Sci. Adv. 15(2), 179–188; doi:10.1111/joa.14234 Herrera, M. and Luque, M. 2009. Valoración morfológica de los animales domésticos. Universidad de Córdoba, Córdoba, Spain. Holgado, F.D., Ortega, M.F. and Fernández, J. 2015. Evolución con la edad de diferentes medidas corporales en hembras bovinas de la raza Criollo Argentino. Actas. Iberoam. Conserv. Anim. 6, 178–183. Immaculate, J., Ebenanjar, E., Sivaranjani, K. and Terence, S. 2020. Application of machine learning algorithms in agriculture. Test Eng. Manag. 82, 9312–9320. Kasarda, R., Moravčíková, N., Mészáros, G., Simčič, M. and Zaborski, D. 2023. Classification of cattle breeds using the random forest approach. Livest. Sci. 267, 105143; doi:10.1016/j.livsci.2022.105143 López-Aguirre, R., Montiel-Palacios, F., Carrasco-García, A.A., Ahuja-Aguirre, C., López-de-Buen, L. and Severino-Lendechy, V.H. 2023. Morphometric characterization and zoometric indices of the Criollo Mixteco cattle from Oaxaca, Mexico. Rev. Bio. Cienc. 10, 1387. Lozada-Soto, E., Vargas-Bello-Pérez, E., Ríos-Utrera, Á. and Martínez-Velázquez, G. 2022. Animals 12(9), 1154; doi: 10.3168/jds.2022-22116 Mahmud, M.S., Zahid, A., Das, A.K., Muzammil, M. and Khan, M.U. 2021. A systematic literature review on deep learning applications for precision cattle farming. Comput. Electron. Agric. 187, 106313; doi:10.1016/j.compag.2021.106313 Martínez, A.M., Gama, L.T., Cañón, J., Ginja, C., Delgado, J.V., Dunner, S., Landi, V., Martín-Burriel, I., Penedo, M.C.T., Rodellar, C., Vega-Pla, J.L., Acosta, A., Álvarez, L.A., Camacho, E., Cortés, O., Marques, J.R., Martínez, R., Martínez, R.D., Melucci, L., Martínez-Velázquez, G., Muñoz, J.E., Postiglioni, A., Quiroz, J., Sponenberg, P., Uffo, O., Villalobos, A., Zambrano, D. and Zaragoza, P. 2012. Genetic footprints of Iberian cattle in America 500 years after the arrival of Columbus. PLos One 7, e49066; doi:10.1371/journal.pone.0049066 Martínez, R.D. 2008. Caracterización genética y morfológica de bovinos criollos. Universidad Nacional del Nordeste, Corrientes, Argentina. Martínez, R.D., Delgado Bermejo, J.V., Martínez, A. and Baselga, M. 1998. Morphological characterization of Argentine Creole cattle. Arch. de Zootec. 47, 11–20. Martínez, M., Rubén., Fernández T., Eduardo., Abbiati C., Nora. and Broccoli R., Ana. 2007. Caracterización zoométrica de bovinos criollos: patagónicos vs noroeste Argentino. Revista MVZ Córdoba, 12(2), 1042–1049. Available via https://www.redalyc.org/articulo.oa?id=69312210 Mejía, A. and Sebastián, R. 2015. Caracterización morfométrica e índices zoométricos en bovinos criollos. Undergraduate thesis, Universidad de las Américas, Quito, Ecuador. Mohan, K., Kumar, P. and Kundu, A. 2025. Physical and phenotypic traits of native cattle (Bos indicus) in the Tarai region of North Bihar for conservation. Vet. World 18(1), 95–101; doi:10.14202/vetworld.2025.95-101 Mundinamani, V.I., Gouri, M.D., Prasad, C. and Patil, V.M. 2024. Physical and morphometric characteristics of unidentified cattle breed of northern Karnataka region of India. Indian J. Dairy Sci. 77(4), 354–361; doi: 10.33785/IJDS.2024.v77i04.008 Parés-Casanova, P.M. 2009. Zoometric indices in livestock species. University of Lleida, Lleida, Spain. Pastor, F., Picot, A., Quintin, F., Ruiz, M., Sevilla, E. and Vijil, E. 2000. Características zoométricas de la raza bovina Pirenaica en función de su origen geográfico. Arch. Zootec. 49, 223–227. Petrov, A. and Kamaldinov, E. 2024. Genetic structure of Siberian branch cattle by microsatellite loci. J. Livestock Biodiversity 14(1), 45–52; doi:10.31677/2072-6724-2024-72-3-230-239 Ramónez-Cárdenas, M.A. and Zhunio-Samaniego, L.E. 2017. Caracterización morfométrica e índices zoométricos de los grupos raciales bovinos existentes en los cantones occidentales de la provincia del Azuay. BSc thesis, Universidad de Cuenca. Ríos-Utrera, A., Montaño-Bermúdez, M., Vega-Murillo, V.E., Martínez-Velásquez, G. and Báez-Rodríguez, J.J. 2018. Genetic parameters for scrotal circumference, frame score and yearling weight of Mexican Charolais and Charbray young bulls. Rev. Colomb. Cienc. Pecu. 31(3), 204–212. Rojas-Espinoza, R., Macedo, R., Suaña, A., Delgado, A., Manrique, Y.P., Rodríguez, H., Quispe, Y.M., Perez-Guerra, U.H., Pérez-Durand, M.G. and García-Herreros, M. 2023. Phenotypic characterization of creole cattle in the Andean highlands using biomorphometric measures and zoometric indices. Animals 13, 11843; doi:10.3390/ani13111843 Setiaji, A. 2023. Differences in phenotypic characteristics and morphology of Jabres and Pasundan cattle. Biodiversitas 25, 3239–3246; doi:10.13057/biodiv/d250745 Sierra, I. 2001. El concepto de raza: evolución y realidad. Arch. Zootec. 50, 547–564. Sobral, M.F., Cravador, A., Navas, D., Roberto, C., Reis, C. and Lima, M.B. 2002. Classification and morphological characterization of native Portuguese cattle using numerical taxonomy. Rev. Port. Zootec. 2, 123–137. Sun, J., Li, Y., Yu, Z., Towns, J.M., Soe, N.N., Latt, P.M., Zhang, L., Ge, Z., Fairley, C.K., Ong, J.J. and Zhang, L. 2025. Exploring artificial intelligence for differentiating early syphilis from other skin lesions: a pilot study. BMC Infect. Dis. 25(1), 113; doi:10.1186/s12879-024-10438-5 Thalkar, M.G., Bhagat, D.J., Kumar, S., Burte, R.G., Desai, B.G., Mayekar, A.J. and Prasade, N.N. 2016. Morphological and morphometric features of non-descript cattle in Raigad district of Maharashtra. J. Livestock Biodiversity 6(2), 67–70. Vargas-Villamil, L.M. and Tedeschi, L.O. 2014. Potential integration of multi-fitting, inverse problem and mechanistic modelling approaches to applied research in animal science: a review. Anim. Prod. Sci. 55(5), 557–568; doi:10.1071/AN14568 Villalobos, A., Martínez, A. and Delgado, J. 2009. Historia de los bovinos en Panamá y su relación con las poblaciones bovinas de Iberoamérica. Arch. Zootec. 58, 121–129. Villalobos-Cortés, A., Carbono, M., Rodríguez, A., Arosemena, E. and Jaén, M. 2021. Phenotypic characterization of the Guaymi breed in Panama’s conservation centers. Afr. J. Agric. Res. 17(6), 907–915; doi:10.5897/AJAR2021.15495 Villalobos-Cortés, A., Rodríguez-Espino, G., Murillo-Alcedo, M., Castillo-Mayorga, H. and Franco-Schafer, S. 2023. Polimorfismos de nucleótido simple asociados a variables ambientales en el genoma de bovinos criollos panameños. Cienc. Agropecu. 36, 37–52. Yakubu, A., Salako, A.E. and Imumorin, I.G. 2011. Multivariate analysis of the morphostructural characteristics of West African dwarf goats. J. Anim. Sci. 89(9), 2697–2704; doi:10.4081/ijas.2011.e17 Zhang, H., Zhang, L., Li, N., Zhang, Y., Zhang, X. and Wang, D. 2025. Machine learning survival models for non-alcoholic fatty liver disease based on a health checkup cohort. BMC. Gastroenterol. 25, 4120; doi:10.1186/s12876-025-04120-6 | ||
| How to Cite this Article |
| Pubmed Style Villalobos-cortés A, León AFD, Jaén M. Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Vet. J.. 2026; 16(3): 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 Web Style Villalobos-cortés A, León AFD, Jaén M. Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. https://www.openveterinaryjournal.com/?mno=295008 [Access: March 31, 2026]. doi:10.5455/OVJ.2026.v16.i3.9 AMA (American Medical Association) Style Villalobos-cortés A, León AFD, Jaén M. Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Vet. J.. 2026; 16(3): 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 Vancouver/ICMJE Style Villalobos-cortés A, León AFD, Jaén M. Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Vet. J.. (2026), [cited March 31, 2026]; 16(3): 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 Harvard Style Villalobos-cortés, A., León, . A. F. D. & Jaén, . M. (2026) Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Vet. J., 16 (3), 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 Turabian Style Villalobos-cortés, Axel, Ana Flores De León, and Marcelino Jaén. 2026. Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Veterinary Journal, 16 (3), 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 Chicago Style Villalobos-cortés, Axel, Ana Flores De León, and Marcelino Jaén. "Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning." Open Veterinary Journal 16 (2026), 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 MLA (The Modern Language Association) Style Villalobos-cortés, Axel, Ana Flores De León, and Marcelino Jaén. "Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning." Open Veterinary Journal 16.3 (2026), 1496-1510. Print. doi:10.5455/OVJ.2026.v16.i3.9 APA (American Psychological Association) Style Villalobos-cortés, A., León, . A. F. D. & Jaén, . M. (2026) Morphological and morphometric comparison and digital classification of Guaymi and Guabala cattle breeds using machine learning. Open Veterinary Journal, 16 (3), 1496-1510. doi:10.5455/OVJ.2026.v16.i3.9 |