| Research Article | ||
Open Vet. J.. 2025; 15(2): 724-730 Open Veterinary Journal, (2025), Vol. 15(2): 724-730 Research Article Neospora caninum in goats from Iraq: A serological and molecular studyRoqaya B. Mohammed* and Mansour J. AliDepartment of Microbiology, College of Veterinary Medicine, University of Al-Qadisyiah, AL-Diwaniyah, Iraq *Corresponding Author: Roqaya B. Mohammed. Department of Microbiology, College of Veterinary Medicine, University of Al-Qadisyiah, AL-Diwaniyah, Iraq. Email: Vet.post22.22 [at] qu.edu.iq Submitted: 22/10/2024 Accepted: 12/01/2025 Published: 28/02/2025 © 2025 Open Veterinary Journal
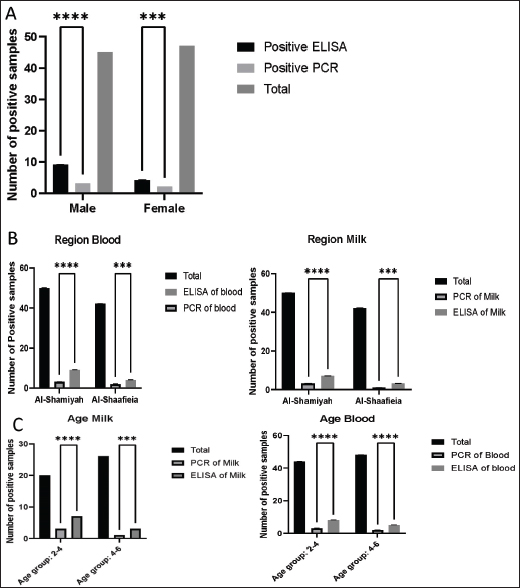
AbstractBackground: Neospora caninum is an Apicomplexa parasite that is an important abortive infectious agent in cattle, goats, and other domestic and wild animals. Aim: This study aimed to identify N. caninum using Enzyme-linked immunosorbent assay (ELISA) and polymerase chain reaction (PCR) in suspected goats in Al-Diwaniyah Province, Iraq. Methods: This study was conducted to identify N. caninum in blood and milk samples of 138 goats (92 blood and 46 milk samples) from suspected infected goats in Al-Diwaniyah Province, Iraq. Two techniques were used; ELISA and PCR (NC5-gene-based and partial sequencing), and the recorded data were categorized into age, sex, and region. Results: ELISA results revealed that the seroprevalence rates were 14.13% (13/92) in blood and 21.73% (10/46) in milk samples. The PCR results demonstrated that for the blood samples, 3/45 (6.66%) and 2/47 (4.25%) were positive for males and females, respectively. The PCR milk findings showed that the highest rates were 3/20 (15%) and 1/26 (3.84%) in the 2–4 and 4–6 age groups, respectively. Sequencing demonstrated close identities of the examined isolates to those from New Zealand and Australia for the blood and milk samples. Conclusions: The findings of this study reveal that N. caninum is widely distributed in goats, which could be an important source of zoonosis in the tested areas. The phylogenetic analysis shows a shared ancestor between the study isolates and those from New Zealand and Australia. Keywords: Serology, Molecular biology, Neosporosis, Goat, Iraq. IntroductionNeospora caninum is an apicomplexan protozoan parasite belonging to the Sarcocystidae group that is globally distributed (Tsakmakidis et al., 2024). Domestic dogs and other canids, the Australian dingo, coyote, and gray wolf, are the only absolute hosts that can spread oocysts throughout the world via their waste in a varying time frame. They are not only intermediate hosts, but they are also essential to the cycle of transmission for N. caninum and the transmission of infection to other infected animals (Lefkaditis et al., 2020). Neospora caninum is an Apicomplex parasite that is an important abortive infectious agent in cattle. Vertical transmission is the main transmission route in N. caninum, which causes reproductive failure in mammals, especially in cattle and goats. Congenitally infected cattle are important sources of horizontal transmission, and infected dogs are important hosts in the life cycle (Gharekhani and Yakhchali, 2020; Baldini et al., 2022). The diagnosis of neosporosis using traditional methods based on clinical symptoms, histopathological findings, and immunohistochemistry is time-consuming and labor-intensive. These methods are not widely used in clinical diagnosis, and they cannot identify the route of transmission and prevalence of infection. The prevalence of N. caninum in horizontal transmission was monitored using polymerase chain reaction (PCR) in dog plasma and serological methods in cattle milk. However, the detection of N. caninum in blood samples is not often possible. The antibodies attached to N. caninum or the characteristic stage-specific gene expression affect the circulation of N. caninum in the blood (Nazari et al., 2023; Zanet et al., 2023). The Sarcocystidae group exhibits two major modes of transmission. Toxoplasma gondii and Hammondia hammondi infect cats for a short time, but oocysts form that pollute the environment. Neospora caninum and Sarcocystis spp., by contrast. Cause chronic urinary and cardiac infections in firm hosts, such as dogs, and intermittent oocyst shed. The water buffalo can be transmitted via birth and horizontal contact with infected hosts, urine, infected food, and water. In cattle, nonphysiological pathways produce environmental contamination and infectious diseases in dairy farms. N. caninum’s host diversity, international distribution, and zoonotic potential and its absence of any effective controls make it a problem for the health and productivity of livestock (Demir et al., 2020; Gharekhani et al., 2020, 2021; Nayeri et al., 2022). The Nc5 gene was initially identified as a surface protein with high polymorphism among N. caninum isolates. Arguably, gene variability triggers large diversification of the isolate. The gene has the potential for differentiating between parasite and animal isolates, neosporotic parasites, and hidden infections. It is currently unclear whether the specific genetic subtypes of N. caninum are the same in goats and other mammals at the same site (Page, 2020; Yi et al., 2024). The NC5 gene in N. caninum is highly polymorphic, and hence, it is used to distinguish strains and identify genetic diversity. This is a polymorphism because of genetic variation in the gene and so the genotypic histories of isolates from different hosts, and from different regions of the world, are different. This diversity is needed to determine which strains of N. caninum there are because they can be identified molecularly and phylogenetically very precisely. The NC5 gene makes a highly immunodominant surface protein (NcSRS2) that allows the parasite to circumvent host defenses. Those variations are important to study host-parasite interactions, virulence genes, and patterns of transmission. NC5 is a high-sensitivity and highly specific polymerase chain reaction marker in diagnosis. Its polymorphism also offers data for evolution and epidemiology, and this is one of the most important features of the N caninum research (Al-Qassab et al., 2010). Neospora caninum, obligatory heteroxene cyst-forming protozoan, family Sarcocystidae, closely related to T. gondii, Hammondia spp., and other Cystoisospora spp. morphological and molecular traits. Neospora caninum is the only species in the genus that infects and re-infects itself in the intermediate host. Moreover, N. caninum is believed to have evolved from an ancestor of T. gondii by losing the ability to encyst in the cat intestine and maintaining only the capacity to infect warm-blooded animals in identical manners (Lindsay and Dubey, 2020; Winzer et al., 2020). In the current study area, using PCR, the microorganism was detected in sheep in 7(3.5%) of the isolates (Al-Shabany and Alkhaled, 2024). No more data can be detected in the literature based on the current study area. The Enzyme-linked immunosorbent assay (ELISA) technique is still a reliable method for the detection of the parasite. This could be strengthened using PCR (Page, 2020; Yi et al ., 2024). This study aimed to identify N. caninum using ELISA and PCR in suspected goats in Al-Diwaniyah Province, Iraq. Materials and MethodsEthical consentThe sample collection methods were approved on August 22, 2023 (approval issue number: 1890) by the Ethical Approval Committee of the College of Veterinary Medicine, University of Al-Qadisiyah. SamplesThe criteria for the sample size depended on the fact that the study region had reported on the disease in different animals but not in goats, which allowed us to survey more than decide the exact number of samples to be collected. The current study contained 138 samples (92 blood by sterile syringes and 46 milk by sterile containers) from suspected infected goats. No definite symptoms, but abortion appeared in some animals. The study was conducted in Al-Diwaniyah Province (31.9740° N, 44.8957° E), Iraq. The samples were collected from different farms and then transferred to the Parasitology Lab at the College of Veterinary Medicine, University of Al-Qadisiyah, Al-Diwaniyah City, Iraq. The blood was taken aseptically from the jugular vein using sterile syringes and placed into clotting anticoagulant tubes. Samples of milk were collected in sterile containers and hygienically prepared to not contaminate it. All samples were immediately frozen and transported to the Parasitology Laboratory of the College of Veterinary Medicine, University of Al-Qadisiyah. The blood samples were centrifuged at 3,000 rpm for 10 minutes to extract serum then stored at 20°C for ELISA. ELISANeospora caninum-specific antibodies were detected in goat blood and milk by ELISA. Neospora caninum antigen was coated on microplates and incubated overnight at 4°C. Following washes of the unbound antigen, blocking was performed with a solution of bovine serum albumin (BSA) to prevent nonspecific binding. The goat serum and milk samples were diluted in buffer, placed into the wells, and incubated for t2 hat 37°C. The samples were washed well and then an enzyme-conjugated secondary antibody was introduced to the goat immunoglobulin. Another second incubation was added an, and Tetramethylbenzidine (TMB) substrate was left to react with the enzyme, causing a hue variation proportional to the number of N. caninum antibodies. The reaction was stopped with sulfuric acid, and the optical densities were read at 450 nm using an ELISA plate reader. Results were read according to the manufacturer’s instructions, and positive samples were interpreted with reference to positive and negative controls. IgG-ELISA was used to detect N. caninum-specific antibodies in goat blood and milk samples because IgG shows recent and past infection (an important part of the parasite’s epidemiology). The test serum samples were diluted 1:100 in the diluent buffer according to the instructions on the ELISA kit for the highest sensitivity and specificity. Molecular techniquesDNA was extracted from blood and milk samples using the kit from (AddBio, Korea). DNA was measured and quantified using a NanoDrop. The primers were F: CAGTCAACCTACGTCTTC and R: GTGCGTCCAATCCTGTAA (328 bp), and they were used by (Al-Ubaidy and Alsultan, 2023). In a volume of 20µl, Master mix, each primer, PCR water, and DNA were 10 µl, 1.5 (0.5 pmol/20), 5 µl, and 2 µl. The conditions of the thermal cycler were as follows: one-cycle initial denaturation, 38 cycles of the triple steps of denaturation, annealing, and extension, and one-cycle final extension at 95°C for 5 minutes, 95°C for 30 seconds, 53°C for 35 seconds, 72°C for 40 seconds, and 72°C for 5 minutes. The PCR products were analyzed via 1.5% agarose gel electrophoresis at 100 V and 80 Amp for 60 seconds. The gel was visualized using a UV-equipped imager. The PCR amplicons were sent to the Macrogen Sequencing facility (Korea). The phylogenetic analysis started with the NCBI website and ended with the recruitment of MEGA 11 software. Statistical analysisThe data were analyzed using Chi-square with the SPSS program as previously described. The results were considered significant when p ≤ 0.05. ResultsELISA and PCR results of blood examinationThe current study found that the total infection rates of the parasite were 13/92 (14.13%) using ELISA compared with 5/92 (5.43%) using PCR, and 9/45 (20%) and 2/47 (4.25%) of samples were serologically positive for males and females, respectively Figure 2. The PCR analyses demonstrated that 6.66% and 4.25% of samples were positive for males and females, respectively (Fig. 1). It was found that, for the blood samples, 9/50 (18%) and 4/42 (9.52%) of samples were positive serologically for the Al-Shamiyah and Al-Shaafieia Districts, respectively, as well as the total PCR blood findings revealed that 3/50(6%) and 2/42 (4.76%) were positive for the Al-Shamiyah and Al-Shaafieia Districts, respectively (Fig. 1). For the blood samples, 8/44 (18.18%) and 5/48 (10.41%) of samples were positive for the 2–4- and 4–6-years age groups, respectively, by ELISA. In addition, the total PCR findings showed that 3/44(6.8%) and 2/48(4.16%) were positive for the 2–4 and 4–6 age groups, respectively (Fig. 1). Fig. 1. Prevalence of neosporosis (using ELISA and polymerase chain reaction) According to (A) sex; (B) region; (C) age. The samples were tested, and the results were recorded, grouped, and analyzed using SPSS software. The statistical analysis was performed using the chi-square test, and the results were considered significant when ***p < 0.001, ****p < 0.0001.
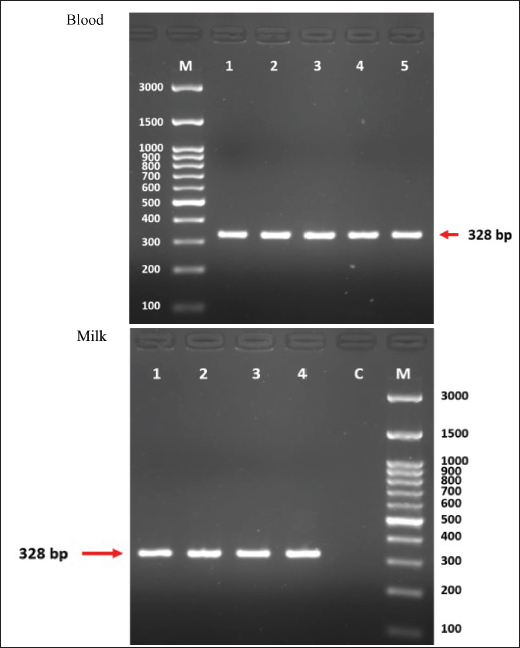
ELISA and PCR results of milk examinationFor the milk samples, 7/30 (23.33%) and 3/16 (18.75%) of samples were positive by ELISA for the Al-Shamiyah and Al-Shaafieia Districts, respectively, while the PCR milk findings showed that 3/30 (10%) and 1/16 (6.25%) were positive for the Al-Shamiyah and Al-Shaafieia Districts, respectively (Fig. 3). In the milk samples, 7/20 (35%), 3/26 (11.53%), and 2/9 (21.73%) of samples were positive serologically for 2–4 and 4–6 years of age groups, respectively. The PCR milk findings showed that 3/20 (15%) and 1/26 (3.84%) were positive for 2–4 and 4–6 age groups, respectively (Fig. 3). For sequencing, the blood and milk samples were similar to those from New Zealand and Australia, respectively (Fig. 3). DiscussionThis is the first study reporting the presence of N. caninum DNA in goat blood, demonstrating the possibility of this species becoming a potential carrier and, if adequate conditions are met, a potential source of environmental contamination. Owning a goat dairy farm is not a risk factor for human toxoplasmosis. However, the presence of N. caninum in goat milk poses a potential infection risk, particularly for immune-compromised consumers (Qian et al., 2020). This study identified the presence of antibodies against microorganisms using ELISA. This result aligns with those of Tsakmakidis et al., (2024). The authors reported that antibodies against N. caninum were detected in 38 of the 172 samples. The total herd prevalence in all farms was 33.33%, and the prevalence of N. caninum antibodies in samples of buffalo milk was 22.10%, which suggests a high population exposure. Fig. 2. Agarose gel electrophoresis of NC5 gene for Neospora caninum from milk and blood samples of goats.
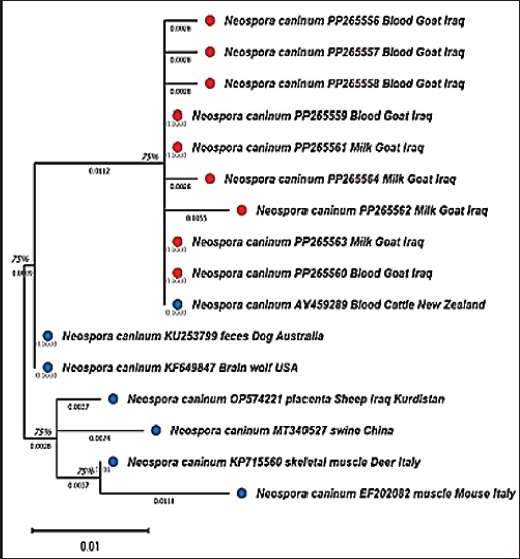
Fig. 3. Phylogenetic analysis of Neospora caninum isolates based on the NC5 gene from blood and milk samples of goats. This tree was developed for the evolution of N. caninum isolates from Iraqi goat blood and milk. The NC5 gene sequences were aligned multiple times using ClustalW in MEGA 11, and the trees were built using the maximum likelihood algorithm. The fit used a nucleotide substitution model, and bootstrap testing with 1,000 replicates was performed to verify the branching patterns. Its tree exhibited similar clustering of Iraqi isolates and New Zealand and Australia isolates.
After the molecular detection of protozoa DNA, part of the positive PCRs was confirmed by sequencing. The NC5 gene is a multi-copy gene encoding the NcSRS2 protein, which is a highly immunodominant protein on tachyzoite that is recognized by the majority of sera raised against the parasite in both experimental and natural infections with any parasite stage. In this study, the PCR-sequencing method worked exceptionally well on naturally infected animal blood and milk samples, revealing valuable insights into the presence of the parasite (Wu et al., 2022; Al-Shabany and Alkhaled, 2024). Neospora caninum strains used here were measured by ELISA and PCR in goat blood and milk samples (different advantages and disadvantages of each). ELISA showed higher infection rates (14.13% in blood and 21.73% in milk) than PCR (5.43% in blood and 4.34% in milk), which illustrates how sensitive it is to antibodies for evidence of infection prior or ongoing. PCR that had gone after the NC5 gene instead revealed N. caninum DNA, direct evidence of the parasite and infection. This disparity between the two methods is what makes ELISA good at screening populations while PCR is better at diagnosing infections at the molecular level. At the statistical scale, these inconsistencies require coordinated diagnostic methods: ELISA works for prevalence studies, and PCR gives the specificity needed for phylogenetic and transmission dynamics. These technologies in total improve the diagnosis of N. caninum infections and add to the species’ epidemiology. The epidemiological data of the current Iraq study echo and supplement other comparable studies and provide important patterns and differences in N. caninum epidemiology. Al-Saadi et al., (2023) found 24.8% N. caninum seroprevalence in cattle in Central India, far higher than the level found in goats in Al-Diwaniyah Province. This could be because of the species or the environment or even the veterinarians. Just like our work, Hebbar et al. (2021) an age risk factor, and recommended long-term parasite exposure. They also included risk factors such as dogs, access to pond water, and abortions, reinforcing the significance of environmental contamination and reproductive malfunction in N. caninum transmission dynamics. Fereig et al. (2024) reported 13.2% N. caninum seroprevalence in cattle, sheep, and goats, which is very similar to our result in Iraq. They found no difference in prevalence between cattle and small ruminants, which is consistent with our finding that the parasite is common in both. These studies show that ELISA is useful for large-scale serological surveys and emphasizes integrated management to control transmission. Yi et al. (2024), in an outbreak in Hunan Province, China, et al. described a lower seroprevalence of N. caninum in goats (2.0%) than ours (which indicates that the risk of exposure varies with region). They also used the NC5 gene to determine if N. caninum was present (this was also done by molecular analysis), just as we did, PCR-based identification and phylogenetic analysis using the NC5 gene. Additionally, Yi et al. stated that age and management with the existence of specific hosts such as dogs and semi-intensive farming systems are risk factors that demand biosecurity. In all these reports, the synergistic application of ELISA and PCR was highlighted as an effective method for diagnosing N. caninum. Serological techniques are ideal for prevalence research, but molecular techniques provide specificity and reassurance of parasite genetics. These findings, taken together, add to our knowledge of the epidemiology of N. caninum and highlight the need for regional control measures that do not compromise animal health and yield. Cross-reactivity between antibodies against T. gondii, Neospora spp., Hammondia spp., Sarcocystis spp., and Besnoitia besnoiti is a prime issue for serological diagnosis using ELISA. These cyst-forming protozoan parasites have antigenic similarity with each other, which creates false positives, making them difficult to diagnose and epidemiologist. Gondim et al. (2017) revealed how cross-reactive T. gondii and H. hammondi (and T. gondii and N. caninum) are. So, infections with N. caninum and N. hughesi, whose serology almost guarantees their same thing, Neospora spp., B. besnoiti, and Sarcocystis spp., there is also serologic cross-reactivity in cows. Hebbar et al. (2022) also took this into consideration when examining cross-reactivity in goats and found that serological tests have to have cut-off dilutions (1:200 for an ELISA) to reduce cross-reactivity. The results show that even at dilutions of serum to the highest level, a high percentage of sera from animals infected with T. gondii were responsive to N. caninum antigens and vice versa, indicating a common antigenic architecture. That’s why serological tests need to be more specific for antigens to make better diagnostic decisions. Gondim et al. (2017) identified such markers as crucial for increasing the specificity of ELISA and other serological tests. Neospora caninum NC5 gene-derived protein, for example, provides a consistent molecular marker that can be used to isolate it from other parasites and may be exploited along with serology. These aspects are essential to the correct diagnosis and epidemiology of close parasites. ConclusionThe findings of the present study reveal that N. caninum is widely distributed in goats, and it could be an important source of zoonosis in the tested areas. The phylogenetic analysis shows a shared ancestor between the study isolates and those from New Zealand and Australia. AcknowledgmentThe College of Veterinary Medicine, University of Al-Qadisiyah, Al-Diwaniyah, Iraq, provided funding for the study. Conflict of interestThe authors declare that they have no conflicts of interest related to the publication of this work. Highlights1. The study investigated a clinical examination and identified specific antibodies in milk and Blood to diagnose caprine neoporosis. 2. The clinical examination was compared to the ELISA results. 3. NC5 gene-based PCR and partial sequencing produced highly accurate results that can be used for the confirmation of neosporosis. ReferencesAl-Ubaidy, Yathrib, and Amjed Alsultan. 2023. Using of Integrons as Biomarker to Assess Dissemination and Diversity of Antimicrobial Resistance Genes in Farm Animal Manure. Journal of Pure and Applied Microbiology 17, no. 3, 1708–1714. Al-Qassab, S.E., Reichel, M.P. and Ellis, J.T. 2010. On the biological and genetic diversity in Neospora caninum. Diversity 2(3), 411–438. Al-Saadi, M., Al-Sallami, D. and Alsultan, A., 2023. Molecular identification of Anaplasma platys in cattle by nested PCR. Iranian Journal of Microbiology, 15(3), p.433. Al-Shabany, A.C. and Alkhaled, M.J. 2024. Nc-5 gene-dependent molecular identification and phylogenetic investigation of Neospora caninum in sheep. Iraqi J. Vet. Sci. 38(2), 443–447. Baldini, M.H.M., Sandoval, E.D.P. and Duarte, J.M.B. 2022. Assessment of transplacental transmission of Neospora caninum and Toxoplasma gondii in Neotropical deer. An estimative based on serology. Vet. Parasitol. 303, 109677. Demir, P.A., Eşki, F. and Ütük, A.E. 2020. Estimating the total economic costs of Neospora caninum infections in dairy cows in Turkey. Trop. Anim. Health Prod. 52(6), 3251–3258. Fereig, R.M., Salama, D.B., Salem, F.K., Rouby, S.R., Shaapan, R.M., Draz, S., Elsawy, B.S.M., Elgioushy, M.M., Altwaim, S.A., Aboelhadid, S.M. and Frey, C.F. 2024. Frequency of Besnoitia besnoiti and Neospora caninum antibodies in cattle and small ruminants from Greater Cairo and Beni Suef governorates, Egypt. Vet. Parasitol. Reg. Stud. Rep. 53, 101078. Gharekhani, J. and Yakhchali, M. 2020. Vertical transmission of Neospora caninum in Iranian dairy cattle. Ann. Parasitol. 66(4), 495–500. Gharekhani, J., Yakhchali, M. and Berahmat, R. 2020. Neospora caninum infection in Iran: a review. J. Parasit. Dis. 44(4), 671–686. Gharekhani, J., Yakhchali, M., Keramat, F. and Berahmat, R. 2021. A literature review of Neospora caninum infection in humans. Avicenna J. Clin. Microbiol. Infect. 8(3), 113–116. Gondim, L.F.P., Mineo, J.R. and Schares, G. 2017. Importance of serological cross-reactivity among Toxoplasma gondii, Hammondia spp., Neospora spp., Sarcocystis spp., and Besnoitia besnoiti. Parasitology 144(7), 851–868. Hebbar, B.K., Mitra, P., Khan, W., Chaudhari, S., Shinde, S. and Deshmukh, A.S. 2022. Seroprevalence and associated risk factors of Toxoplasma gondii and Neospora caninum infections in cattle in Central India. Parasitol. Int. 87, 102514. Lefkaditis, M., Evagelopoulou, G., Sossidou, A. and Spanoudis K. 2020. Neosporosis and toxoplasmosis are two prevalent and important protozoses in dairy cows in small farms from Thessaly, Central Greece. Hellenic Vet. Med. Soc. 71(3), 2357–2362. Lindsay, D.S. and Dubey, J.P. 2020. Neosporosis, toxoplasmosis, and sarcocystosis in ruminants: an update. Vet. Clin. Food Anim. Pract. 36(1), 205–222. Nayeri, T., Moosazadeh, M., Sarvi, S. and Daryani, A. 2022. Neospora caninum infection in aborting bovines and lost fetuses: a systematic review and meta-analysis. PLoS One 17(5), e0268903. Nazari, N., Khodayari, M.T., Hamzavi, Y., Raeghi, S., Karamati, S.A., Falahi, S., Bozorgomid, A. and Sajedi, M.T. 2023. Systematic review and meta-analysis of the role of felids as intermediate hosts in the life cycle of Neospora caninum based on serological data. Acta Parasitol. 68(1), 266–276. Page, R.J. 2020. Associations among beef cattle genotypes, Neospora caninum infections, and reproductive performance. University of Arkansas ProQuest Dissertations & Theses. Qian, W., Yan, W., Lv, C., Bai, R., Wang, T., Wei, Z. and Zhang, M. 2020. Molecular detection and genotyping of Toxoplasma gondii and Neospora caninum in slaughtered goats in Central China. Foodborne Pathog. Dis. 17(5), 348–356. Tsakmakidis, I., Lefkaditis, M., Sofia, M., Zaralis, K. and Giantsis, I.A. 2024. Prevalence of Neospora caninum and associated risk factors in semi-intensive female water buffaloes reared in Greece, using an ELISA kit on milk samples. Vet. Parasitol. Reg. Stud. Rep. 54, 101081. Winzer, P., Müller, J., Imhof, D., Ritler, D., Uldry, A.C., Braga-Lagache, S., Heller, M., Ojo, K.K., Van Voorhis, W.C., Ortega-Mora, L.M. and Hemphill, A. 2020. Neospora caninum: differential proteome of multinucleated complexes induced by the bumped kinase inhibitor BKI-1294. Microorganisms 8(6), 801. Wu, K., Song, X., Wu, Y., Yang, X., Liu, J. and Liu, Q. 2022. Deleting ku80 improves the efficiency of targeted gene editing in Neospora caninum. Mol. Biochem. Parasitol. 251, 111508. Yi, X.L., Yang, W.H., Zheng, H.L., Cao, M.L., Xiong, J., Chen, W.C., Zhou, Y.J., Li, F., Zhu, X.Q. and Liu, G.H. 2024. Seroprevalence and molecular detection of Toxoplasma gondii and Neospora caninum in beef cattle and goats in Hunan Province, China. Parasites Vectors 17(1), 195. Zanet, S., Poncina, M. and Ferroglio, E. 2023. Congenital transmission of Neospora caninum in wild ungulates and foxes. Front. Vet. Sci. 10, 1109986. | ||
| How to Cite this Article |
| Pubmed Style Mohammed RB, Ali MJ. Neospora caninum in goats from Iraq: A serological and molecular study. Open Vet. J.. 2025; 15(2): 724-730. doi:10.5455/OVJ.2025.v15.i2.21 Web Style Mohammed RB, Ali MJ. Neospora caninum in goats from Iraq: A serological and molecular study. https://www.openveterinaryjournal.com/?mno=225585 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i2.21 AMA (American Medical Association) Style Mohammed RB, Ali MJ. Neospora caninum in goats from Iraq: A serological and molecular study. Open Vet. J.. 2025; 15(2): 724-730. doi:10.5455/OVJ.2025.v15.i2.21 Vancouver/ICMJE Style Mohammed RB, Ali MJ. Neospora caninum in goats from Iraq: A serological and molecular study. Open Vet. J.. (2025), [cited January 25, 2026]; 15(2): 724-730. doi:10.5455/OVJ.2025.v15.i2.21 Harvard Style Mohammed, R. B. & Ali, . M. J. (2025) Neospora caninum in goats from Iraq: A serological and molecular study. Open Vet. J., 15 (2), 724-730. doi:10.5455/OVJ.2025.v15.i2.21 Turabian Style Mohammed, Roqaya B., and Mansour J. Ali. 2025. Neospora caninum in goats from Iraq: A serological and molecular study. Open Veterinary Journal, 15 (2), 724-730. doi:10.5455/OVJ.2025.v15.i2.21 Chicago Style Mohammed, Roqaya B., and Mansour J. Ali. "Neospora caninum in goats from Iraq: A serological and molecular study." Open Veterinary Journal 15 (2025), 724-730. doi:10.5455/OVJ.2025.v15.i2.21 MLA (The Modern Language Association) Style Mohammed, Roqaya B., and Mansour J. Ali. "Neospora caninum in goats from Iraq: A serological and molecular study." Open Veterinary Journal 15.2 (2025), 724-730. Print. doi:10.5455/OVJ.2025.v15.i2.21 APA (American Psychological Association) Style Mohammed, R. B. & Ali, . M. J. (2025) Neospora caninum in goats from Iraq: A serological and molecular study. Open Veterinary Journal, 15 (2), 724-730. doi:10.5455/OVJ.2025.v15.i2.21 |