| Research Article | ||
Open Vet. J.. 2025; 15(10): 5335-5345 Open Veterinary Journal, (2025), Vol. 15(10): 5335-5345 Research Article Silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus)Mohammad Sukmanadi1*, Sri Agus Sudjarwo1, Mustofa Helmi Effendi2, Aswin Rafif Khairullah3, Sri Pantja Madyawati4, Imam Mustofa4, Pudji Srianto4, Mirni Lamid5, Hani Plumeriastuti6, Riza Zainuddin Ahmad3, Adeyinka Oye Akintunde7, Mo Awwanah8, Bima Putra Pratama9 and Agung Prasetyo101Division of Basic Veterinary Medicine, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 2Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 3Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor, Indonesia 4Division of Veterinary Reproduction, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 5Division of Animal Husbandry, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 6Division of Veterinary Pathology, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 7Department of Agriculture and Industrial Technology, Babcock University, Ilishan Remo, Nigeria 8Research Center for Applied Botany, National Research and Innovation Agency (BRIN), Bogor, Indonesia 9Research Center for Agroindustry, National Research and Innovation Agency (BRIN), South Tangerang, Indonesia 10Research Center for Estate Crops, National Research and Innovation Agency (BRIN), Bogor, Indonesia *Corresponding Author: Mohammad Sukmanadi. Division of Basic Veterinary Medicine, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia. Email: moh-s [at] fkh.unair.ac.id Submitted: 27/04/2025 Revised: 14/08/2025 Accepted: 05/09/2025 Published: 31/10/2025 © 2025 Open Veterinary Journal
AbstractBackground: Mycotoxins are secondary metabolites produced by fungi that cause serious diseases and death. Among the mycotoxins, aflatoxins stand out for their significant toxic impact, especially Aflatoxin B1 (AFB1) produced by Aspergillus species. Capsaicin, known for its pungent taste, is also highlighted in this study for its potential health benefits, which have been shown to have beneficial effects on the cardiovascular system, metabolism, analgesia, and management of disorders. Aim: This study aimed to determine the in silico inhibition effect of capsaicin on the expression of AKT1 and MAPK1 target proteins in mice due to AFB1 induction. Methods: Molecular docking is a bioinformatics technique that combines biological knowledge with databases (big data) to ascertain the pattern, affinity, and possible activity of chemical compounds interacting with target proteins. When the Gibbs free energy (ΔG) changes, protein-ligand interaction occurs. Results: The findings demonstrated that molecular docking, protein-ligand interactions, homology modeling, target protein prediction, protein interaction analysis, and molecular visualization in silico. The target proteins AKT1 and MAPK1 were highly expressed in the AFB1 binding prediction results in mice (Mus musculus). The tethering of negative Gibbs free energy molecules was deemed satisfactory, and 3D virtual visualization demonstrated excellent adhesion to the receptors of the molecules. Conclusion: Capsaicin can synergistically inhibit regulatory cells through the AKT 1 and MAPK 1 signaling pathways, potentiating the hepatotoxicity action protective effect of capsaicin against AFB-1 intoxication. Therefore, capsaicin is a potential candidate for herbal medicine as a hepatoprotective agent. Keywords: Aflatoxin B1, Capsaicin, Hepatoprotection, Herbal medicine, Good health and wellbeing IntroductionMycotoxins are secondary metabolites produced by fungi that can cause serious illness, including death, in humans and animals (Awuchi et al., 2021). However, some mycotoxins or their derivatives also have pharmacological activity and have been used as antibiotics, growth promoters, and other drugs (Bilal et al., 2023). Aftoxin is one of the most well-known mycotoxins is aflatoxin, which is produced by fungi species of the genus Aspergillus, such as Aspergillus flavus and Aspergillus parasiticus (Shabeer et al., 2022). Aflatoxins are often found in hot and humid climates, especially at temperatures of 27°C–40°C and relative humidity of 85% (Mahato et al., 2019). Aflatoxin B1 (AFB1) and Aflatoxin B2 (AFB2) are the two main forms of aflatoxin formed, while Aflatoxin M1 and M2 are metabolites resulting from the hydroxylation of AFB1 and AFB2 by the cytochrome P450 1A2 enzyme in animals and humans who consume contaminated food (Popescu et al., 2022). After entering the body, AFB1 can activate a series of key proteins that play a role in hepatotoxicity, including various protein kinases such as Glycogen Synthase Kinase 3 Beta (Gsk3b), Mechanistic Target of Rapamycin (Mtor), Epidermal Growth Factor Receptor (Egfr), AKT Serine/Threonine Kinase 1 (Akt1), Mitogen-Activated Protein Kinase 1 (Mapk1), Mitogen-Activated Protein Kinase 3 (Mapk3), and (RAF Proto-Oncogene Serine/Threonine-Protein Kinase) Raf1 (Antonius et al., 2022). The direct interaction of AFB1 with these proteins may trigger the activation of certain genes that potentially increase the risk of liver damage (Marchese et al., 2018). Of the AFB1 target proteins, seven protein kinases have been identified to play important roles in cell pathology (Tran et al., 2024). Alpha kinase threonine 1 (AKT1) is one of the three types of Akt kinases involved in the regulation of metabolism, cell proliferation, growth, and angiogenesis (Benkerroum, 2020). Meanwhile, Mapk1 and Mapk3 proteins, which are serine/threonine kinases, regulate MAP signal transduction, resulting in responses related to cell growth, adhesion, survival, and differentiation (Pearson, 2001). Capsaicin (8-methyl-N-vanillyl-6-nonenamide) is responsible for the spiciness of chili peppers (Wang et al., 2025). When consumed, capsaicin stimulates sensory neurons that transmit pain information to the central nervous system, creating a burning sensation of pain (Alalami et al., 2024). Chilies’ spiciness level depends on the capsaicin and other capsaicinoid compounds, which account for 90% of the total capsaicinoids in chilies (Orellana-Escobedo et al., 2012). Capsaicin has a variety of positive health effects, including analgesic effects, cardiovascular support (McCarty et al., 2015), diabetes management (Szallasi, 2022), gastric protection (Xiang et al., 2022), treatment of urogenital disorders (Shao et al., 2023), and weight loss (Elmas and Gezer, 2021). However, the therapeutic potential of capsaicin in treating respiratory system diseases remains controversial, and the therapeutic effects of capsaicin agonists and antagonists and the role of TRPV1 require further investigation. Among the existing mycotoxins, aflatoxins stand out for their significant toxic impact, especially AFB1 produced by Aspergillus species (Benkerroum, 2020). This research not only focuses on the adverse effects of aflatoxins but also investigates the possibility of developing new therapeutic approaches based on the pharmacological activity profile of mycotoxins. Capsaicin, which is known for its spicy taste, was also highlighted in this study for its potential health benefits (Fattori et al., 2016). Capsaicin improves metabolism, the cardiovascular system, analgesia, and the treatment of illnesses (Thongin et al., 2022). Although capsaicin is often considered a positive compound, this new approach shows that it is important to assess the dual effects of capsaicin, which can be either beneficial or detrimental. This study explores new dimensions in the understanding of mycotoxins and capsaicin and their potential roles in the context of human and animal health. Through the in silico approach, the potential bioactive interactions of these compounds can be explored before conducting in vivo studies, thereby minimizing risks and maximizing the understanding of the mechanisms involved. This study aimed to determine the inhibitory effect of capsaicin on the expression of AKT1 and MAPK1 target proteins in mice following AFB1 induction in silico. This study not only contributes to basic science but also has practical implications in the development of safer and more effective alternative therapies for diseases associated with mycotoxins and inflammatory processes. Materials and MethodsIn silico studyA computerized simulation known as "in silico" is used to develop or evaluate medicinal items, technologies, or therapies using current technology and a database-linked understanding of biology (Musuamba et al., 2021). Several terms must be understood, and with their operational definitions. A technique for determining a chemical compound’s interaction pattern, affinity, and possible activity with a target protein is molecular docking (Agu et al., 2023). In a spontaneous chemical process where protein-ligand binding occurs when the Gibbs free energy changes (ΔG), the magnitude of binding energy serves as a measure of a molecule’s ability to interact with a target protein (Du et al., 2016). Tools required for the in silico analysisThe tools needed for in silico analysis are a computer with a RAM capacity of 2 GB, Java, Phyton, and Adobe Flash Player features to support analysis compatibility, an Intel Core i3-4005U 1.7 GHz CPU, and Windows 10 OS. AFB1 molecule samples with CID 186907 and Capasicin (CAP) with CID 1548943 were obtained from the chemical compound database PubChem (pubchem.ncbi.nlm.nih.gov). PubChem is a database that stores specific information related to organic and synthetic compounds from laboratories. This database displays physicochemical, pharmacological, and pharmacodynamic properties and is equipped with online compound drawing facilities (Kim et al., 2016). Samples were downloaded in structure data format (sdf) to be minimized using the Open Babel plug-in in PyRx 0.8 software to obtain the 3D structure of AFB1. Compound minimization during preparation before docking aims to change the target compound file format from.sdf to.pdb and increase the flexibility of the chemical compound structure by obtaining positive bond energy between its atoms (Adnan et al., 2023). In addition to the 3D structure, Canonical SMILE AFB1 is stored on a notepad to allow prediction of target proteins in the body of mice (Mus musculus). SMILE Canonical is a type of chemical formula format that allows researchers to perform advanced analysis in bioinformatics for predicting drug physicochemical properties, toxicity, target proteins, gene expression, and other parameters (Badwan et al., 2023). Prediction of target proteinTarget protein prediction is done with SMILE Canonical AFB1 stored in a notepad and inputted to the online target protein prediction server, SwissTargetPrediction, with the page www.swisstargetprediction.ch, and the target organism is mice (M. musculus). The names of the AFB1 target proteins are stored in a notepad after making the prediction. SwissTargetPrediction is a server that allows the prediction of macromolecular targets from a small molecule, such as a bioactive molecule. The prediction is found through a combination of 2D and 3D similarity in a database with known activities of 370,000 from 3,000 types of proteins in different species (Daina et al., 2019). Protein interaction analysis (PIA)PIA was conducted using the name of the AFB1 target protein obtained from the prediction analysis, which was then inputted into the biological pathway database, namely the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING), with the string-db.org page. This stage aims to determine the biological function of a protein that acts as a hepatocarcinoma oncogene in the body of mice (M. musculus). STRING is a database or server used for protein-PIA based on the name of the single target protein, multi-protein, and sequence. The interaction between proteins is shown in the line connecting the shell of each protein and shows the specific biological activity of each interaction (Susanto et al., 2018). Homology modelingHomology modeling is done by: oncogene proteins that are successfully identified in the biological pathway database, then their amino acid sequences are accessed in a specific database, namely Uniprot, with the page www.uniprot.org with FASTA format. The sequence is inputted into the homology modeling server, namely SWISS MODEL, with the page swissmodel.expasy.org to perform the process of modeling the 3D structure of HCA oncogene proteins. Homology modeling is a method of modeling query amino acid sequences formed from templates in a database to produce homologous protein structures. Protein structures with a similarity score above 20% are declared similar to the template (Guex et al., 2009; Waterhouse et al., 2018). Molecular dockingMolecular docking is the 3D structure of the target protein obtained from the modeling results downloaded in the protein databank (pdb) format, and the AFB1 ligand is input into the PyRx 0.8 software for the molecular docking process. This analysis aims to determine the level of affinity and atomic bond distance of AFB1 to each target protein to determine the interaction pattern. Molecular docking is used to determine the interaction pattern, affinity, and potential activity of a chemical compound when interacting with a target protein (Kharisma et al., 2018). LiganPada penelitian ini menggunakan suatu ligan yang diinteraksikan dengan lebih dari satu protein target untuk mengetahui besar kecilnya energi pengikatan ligan untuk aktivasi protein target. Protein-ligand interactions and molecular analysisThe results of the molecular docking analysis were analyzed for protein-ligand interactions using LigPlus software to determine the type of chemical bond interactions that occur in the atoms that make up AFB1 with the amino acid residues of the target protein. Representative molecular visualization of the 3D structure of the protein was performed using PyMol software, which displays the secondary protein structure and the number of constituent chains (El Khoury et al., 2023). Data analysisTarget protein prediction, PIA, homology modeling, molecular docking, protein-ligand interactions, and molecular visualization in the form of figures and tables are all included in the results from the in silico study. Ethical approvalNot needed for this study. ResultsThe animal model setting on the server was selected as the experimental animal mice (M. musculus). The predicted results of AFB1 binding in the body of the mice (M. musculus) (Table 1) showed interactions with 3 types of protein families, namely kinase, phosphatase, and an G protein-coupled receptor, with binding probabilities of 46.7%, 20.0%, and 6.7%, respectively. Table 1. Aflatoxin B1 target proteins in Mus musculus.
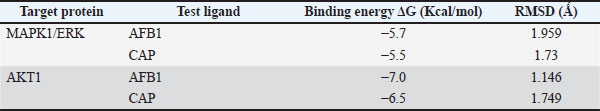
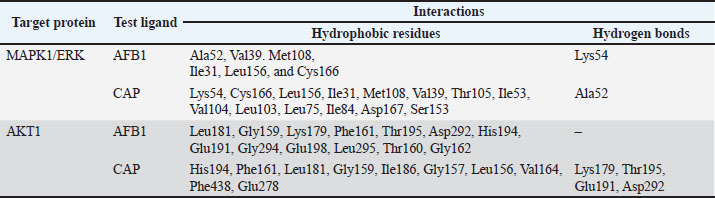
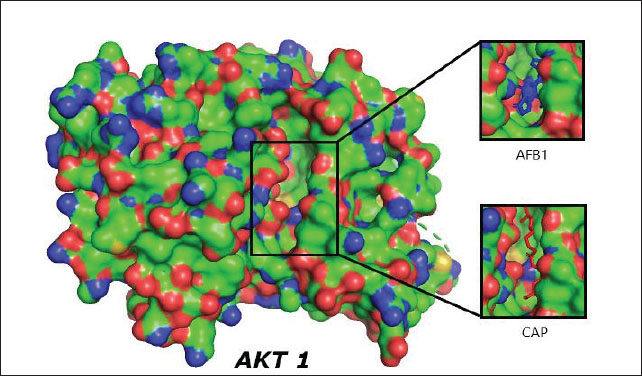
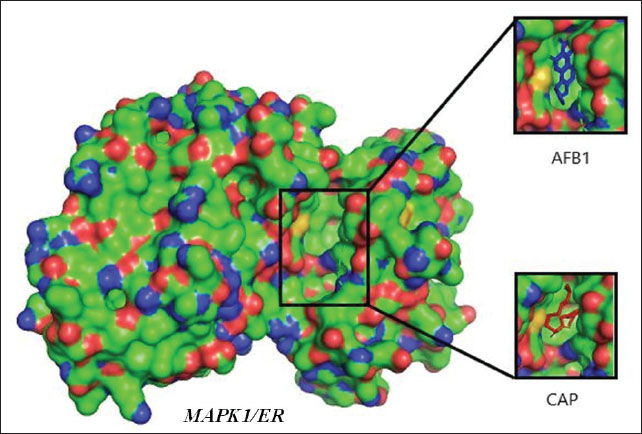
Proteins involved in the apoptotic pathway of cells caused by aflatoxin intoxication are MAPK1 in the MAPK1 signaling pathway and AKT1 in the PI3K/Akt signaling pathway. Aftoxin induces the formation of reactive oxygen species (ROS) and DNA damage in hepatocytes (Zhang et al., 2023). Molecular docking was performed on MAPK1 (ERK) and Akt1 proteins using the Autodock Vina 4.0 program and the CAP and Aflatoxin (AFB1) test ligands as intervening (control variables). The results of molecular docking are Gibbs free energy (ΔG) and virtual 3D visualization results. After performing molecular docking, the process of determining the active side of the protein and the amino acids that will bind to the test ligand (residue) is visualized using Ligplot+ and PyMOL. The quality of the molecular docking test results can be determined by analyzing the binding energy between the target protein and the test ligand using the Gibbs free energy (ΔG) and by analyzing the binding site to determine the interaction between the target protein and the test ligand. The value of ΔG comes from the scoring function with the following equation: S1 (ΔG)=g (c1 − cintra1)=g (cinter1) Where g is a nonlinear function and c is the conformation or interaction caused by intramolecular or intermolecular hydrophobic reactions or hydrogen bonds. If ΔG is negative, the reaction will occur spontaneously, and the affinity of the test ligand to the target protein will be higher (Trott and Olson, 2010). Validation of the molecular docking is that the root mean square deviation (RMSD) value must be less than 2 Ǻ because the smaller the RMSD value, the closer it is to the original binding position of the test ligand (binding pose) with the simulated target protein. If it exceeds 2 Ǻ, the binding pose gets worse, rendering the results invalid. After molecular docking, it was found that CAP binds to AKT1 with a ΔG of −6.5 Kcal/mol, with a ΔG value for Aflatoxin (AFB1) of −7.0 Kcal/mol, with visualization of the binding and binding site in Figure 1. In the test with the MAPK1/ERK target protein (Fig. 2), the ΔG for CAP was −5.5 Kcal/mol and for AFB1 was −5.7 Kcal/mol. All of these ΔGs are very negative, so the reaction will occur spontaneously and cause high affinity for both AFB1 and CAP on the MAPK1/ERK and Akt target proteins, and all RMSD values for the four tests are below <2 Ǻ (Table 2).
Fig. 1. Visualization of the interaction of the test ligands CAP (marked in red and Aflatoxin (AFB1) (intervening) marked in blue) with the target protein AKT1, both binding together at the same binding site/cavity.
Fig. 2. Visualization of the interaction of the test ligands CAP [marked in red and Aflatoxin (AFB1) (intervening) marked in blue] with the target protein MAPK1/ERK, both of which bind together at the same binding site/cavity. Table 2. Molecular docking test results between MAPK1 and AKT1 target proteins with CAP and Aflatoxin (AFB1).
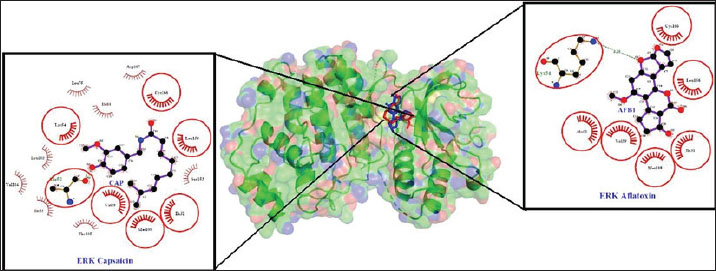
In Figures 1 and 2, the type of interaction between the target protein and the test ligand is known using amino acid analysis that binds to the test ligand, or known as residues. After molecular docking using the Ligplot+ application, residue analysis is used, and the results can be combined using PyMOL. Visualization of the interaction of the test ligand CAP is marked in red, and AFB1 (intervening) is marked in blue, with the target proteins AKT1 and MAPK1, both of which bind together at the same binding site/cavity. The type of interaction between the target protein and the test ligand can be determined using amino acid analysis that binds to the test ligand or is known as residue. After molecular docking using the Ligplot+ application, residue analysis is used, and the results can be combined using PyMOL (Kharisma et al., 2018). The results of the interaction analysis for the target proteins MAPK1/ERK and AKT1 and the test ligands Aflatoxin (AFB1) and CAP are shown in Table 3, Figures 3 and 4.
Fig. 3. Amino acid residues of the AKT1 target protein CAP test ligand in red and Aflatoxin (AFB1) as control in blue. Green lines indicate hydrogen bonds, red eyelash-shaped lines indicate hydrophobic reactions, and red circles indicate similar amino acid residues between the two test ligands.
Fig. 4. Amino Acid Residues of MAPK1/ERK Target Proteins CAP test ligands are colored in red, and Aflatoxin (AFB1) test ligands are colored in blue. Green lines indicate hydrogen bonds, red eyelash-shaped lines indicate hydrophobic reactions, and red circles indicate similar amino acid residues between the two test ligands. Table 3. Interaction analysis results by looking at the residues between the test ligand and the target protein.
DiscussionMolecular docking was performed on MAPK1 (ERK) and AKT1 proteins using the Autodock Vina 4.0 program and the CAP and Aflatoxin (AFB1) test ligands as intervening ligands. The virtual 3D visualization findings and Gibbs free energy (ΔG) are the outcomes of this molecular docking. Since it indicates that the drug being examined by molecular docking will bind to its receptor very well and occur spontaneously, a negative Gibbs free energy value is favorable (Du et al., 2016). After performing molecular docking, the process of determining the active site of the protein and the amino acids that will bind to the test ligand (residue) is visualized using Ligplot+ and PyMOL (Rauf et al., 2015). The binding energy between the target protein and the test ligand was analyzed using the Gibbs free energy (ΔG), and the binding site was examined to ascertain the interaction between the target protein and the test ligand. These two methods were used to assess the quality of the molecular docking test results (Ivanova and Karelson, 2022). The value of ΔG is derived from the scoring function. The Capsaicin Ligand Files and the AKT1 and MAPK1 protein files that had previously been created in Autodock 4.0 were put through molecular docking. Vina is a docking application that yields binding energy from ligands to receptors (Trott and Olson, 2010). The most promising ligand will then be validated using the Passonline webserver to ascertain the compound’s potential, and the ligand file with the highest potential will be visualized using PyMOL and Ligplot+ to ascertain the bond between the ligand and the amino acid residue at the binding site (Saliu et al., 2021). Gibbs free energy (ΔG) affects bond energy; spontaneous reactions will have a negative Gibbs free energy at constant temperature (Chen, 2019). Several components influence the bond energy, which is expressed by the following equation: ∆GBond=∆GGauss + ∆GRepulsi + ∆GHBond+ ∆GHydrophobic + ∆GTorsion. As the number of energy components increases, the ∆GBond value will decrease (become negative), strengthening the bond and creating a high affinity (Kharisma et al., 2018). Trott and Olson (2010) explained that the conformation and interactions that occur greatly affect the value of the ΔG binding energy, and its impact indicates the potential for binding from the test ligand to the target protein. Similar amino acid residues between two test ligands also indicate the binding site or place where the target protein binds to the test ligand. The more similar the residues, the more similar the binding site, resulting in a competition effect between the two test ligands and an inhibitory effect (Roy and Kapoor, 2018). A sample of AFB1 compound ID: 186907 and 3D structure, along with Smile Canonical: COC1=C2C3=C(C(=O)CC3)C(=O)OC2=C4C5C=COC5OC4=C1 with molecular weight of 312.277 g/mol and capsaicin compound (CAP) ID: 1548943 and 3D structure, along with Smile Canonical: CC(C)C=CCCCCC(=O)NCC1=CC(=C(C=C1)O)OC with molecular weight of 305.4 g/mol were obtained from PubChem database (pubchem.ncbi.nlm.nih.gov). Information related to the target proteins of AFB1 and CAP was obtained from the Swiss Target Prediction server (www.swisstargetprediction.ch), an online analysis page that predicts target proteins from a chemical compound based on Canonical SMILE. In this study, the animal model setting on the server was chosen as M. musculus. SwissTargetPrediction is a server that allows users to estimate target proteins that bind small molecules or can be called bioactive. Approximately 370,000 predictions have been made from 3,000 proteins found in three different organisms, namely Homo sapiens, M. musculus, and Rattus norvegicus (Gfeller et al., 2014). The IDs of the target proteins can be used to access further information without writing the full name of the target protein in the database. The protein group with ID in the AFB1 and CAP target prediction results are used to access further information through the UniProt database (Daina et al., 2019). In the AKT1 target protein, there are eight amino acid residues that are similar between the two test ligands, namely, Leu181, Gly159, Lys179, Phe161, Thr195, Asp292, His194, and Glu191 in both the CAP test ligand and the intervening AFB1. The striking difference is in the residues Lys179, Thr195, Glu191, and Asp292 in the CAP test ligand, which have interactions in the form of hydrogen bonds, whereas all interactions in AFB1 are hydrophobic. In the MAPK1/ERK target protein, there are seven similar residues, including Ala52 and Val39. Met108, Ile31, Leu156, Cys166, and Lys54, where each target protein has 1 hydrogen bond. In the CAP test ligand, there is 1 hydrogen bond on the Ala52 residue, whereas in the AFB1 test ligand, there is a hydrogen bond on the Lys54 residue. Based on the examination of amino acid residues, capsaicin against MAPK1 and AKT1 have almost identical binding sites and impact the receptor at a comparable location, specifically the receptor/protein. Therefore, further analysis is needed to determine the effect of capsaicin on MAPK1 and AKT1. The Passonline webserver must be used to analyze the potential of Capsicum sp. herbs based on their chemical structure. In the target proteins AKT1 and MAPK1/ERK with the test ligands CAP and AFB, both test ligands have similar binding sites. Thus, it can be predicted that both test ligands will cause a competitive inhibitory effect, where CAP can inhibit the expression of AKT1 and MAPK1/ERK proteins by inhibiting the binding between AFB1 and MAPK1/ERK by occupying the same binding site. Capsaicin has an excellent activity value in this test and may be a hepatoprotection because it inhibits AKT1 and MAPK1/ERK signaling and has an agonist apoptotic action, which stops cell proliferation (Sukmanadi et al., 2020). The results of AFB1 binding prediction in the body of mice (M. musculus) showed interactions with three types of protein families, namely kinase, phosphatase, and A G protein-coupled receptor, with respective binding probability percentages of 46.7%, 20.0%, and 6.7%, respectively. The results show that AFB1 has a dominant target protein, including the kinase group, because its binding probability score is higher than that of other proteins. The probability on the SwissTargetPrediction server shows the level of ability of a small molecule to bind to the target protein. The higher the score, the higher the binding potential (Daina et al., 2019). The server used in this study makes predictions based on the online ligand-based target prediction method, which was developed from the previous version and works quickly to predict the target protein of a small molecule (Cervello et al., 2017). However, the results of this prediction are still a hypothesis and can be further confirmed through laboratory analysis (Sulaimany et al., 2024). Kinase is a group of enzymes that catalyze phosphate groups to form ATP and ADP molecules for dephosphorylation (Seok, 2021). This mechanism impacts the activity, reactivity, and binding ability of molecules such as proteins, lipids, and carbohydrates, thus playing an important role in the physiological function of living things (Martínez-Alcantar et al., 2024). Protein kinase plays a role in the phosphorylation mechanism of amino acid residues serine, threonine, tyrosine, and histidine (Ardito et al., 2017). Phosphorylation can modify the function of a protein through many pathways. It can increase or decrease the activity of a protein, establish stabilization mechanisms, localize within cellular compartments, and initiate the degradation of other proteins or biological functions such as cell proliferation and differentiation (Zhong et al., 2020). AFB1 can interact with target proteins dominated by the kinase group when entering the body of mice (M. musculus). However, the predicted AFB1 target protein must be interacted with to determine the function of the biological pathway and determine its association with the structure and physiology of liver cells in the body of mice (M. musculus) (Marchese et al., 2018). Hydrogen bonds also significantly affect the affinity of the test ligand to the target protein. The higher the number of hydrogen bonds, the better the affinity of a test ligand to the target protein (Madushanka et al., 2023). Hydrophobic interactions direct molecules toward the binding site, and hydrogen bonds increase affinity and direct ligands to proteins (Snyder et al., 2011). In the AKT1 protein, the CAP test ligand has more hydrogen bonds compared to AFB1, proving that CAP has a tendency to bind to AKT compared to AFB1 to inhibit signaling from the AKT pathway, which can inhibit the occurrence of hepatocyte necrosis downstream. Although the in silico approach in this study provides important initial insights into potential molecular interactions between capsaicin and target proteins involved in AFB1-induced hepatotoxicity, the results have several inherent limitations that need to be recognized. Molecular docking simulations rely heavily on the accuracy of the force fields and scoring functions used to estimate the target protein’s ligand binding affinity (Pantsar and Poso, 2018). For example, force fields, such as those used in AutoDock software, are simplified mathematical representations of complex interactions between atoms (Mohanty and Mohanty, 2023). These force fields do not fully capture the dynamic contributions of the molecular environment, such as explicit solvent effects, thermal fluctuations, or interactions between protein domains (Trott and Olson, 2010). This can result in overestimation or underestimation of ligand affinity; therefore, docking results should be considered preliminary predictions rather than definitive evidence. Scoring functions used in docking generally only consider the binding free energy under static conditions and ignore the contributions of entropy, protein conformational flexibility, and induced reactions after binding (Tripathi and Bankaitis, 2017). Proteins in biological systems have complex structural dynamics, and their active pockets are not always as rigid as assumed in most docking simulations (Lexa and Carlson, 2012). The possibility of significant conformational changes (induced fit) or the influence of biological cofactors is not adequately accounted for in these studies. Another limitation involves the homology model quality used for the target protein. If an experimental crystallographic structure is not available and a 3D model is obtained through homology modeling, the structure accuracy will depend heavily on the sequence identity between the template and target and on the modeling quality and structure refinement (Forrest et al., 2006). Structural inaccuracies can directly impact the binding site position and docking results. This study did not include MD analysis, which can capture real-time protein-ligand conformational changes, evaluate complex stability, and calculate parameters such as RMSD, RMSF, and radius of gyration. Without MD data, the validity of binding interactions remains predictive and does not reflect dynamic biological conditions. ConclusionThis study examines the role of mycotoxins, especially aflatoxins, produced by fungi in diseases in humans and animals. AFB1 is involved in hepatotoxicity by activating protein kinases, such as Gsk3b and Akt1. Based on computational studies with in silico bioinformatics, it is concluded that the effect of capsaicin as a hepatoprotection in preventing hepatic injury by AFB1 is through the target proteins AKT1 and MAPK1 expressed in hepatocyte cells. AcknowledgmentsThe authors would like to thank Dr. Nove Hidajati, DVM., M.Health., for managerial support, Prof. Dr. Lilik Maslachah, DVM., M.Health., for expertise support, Dr. Rahmi Sugihartuti, DVM., M.Health. for technical support. Conflict of interestThe authors declare no conflict of interest. FundingThe authors funded this study. Author’s contributionsMS, SAS, IM, and MHE: conceived the idea, designed the mainframe of this manuscript, and acquired, analyzed, and interpreted data. BPP, RZA, AOA, AP, and ARK: drafted the manuscript. PS, SPM, ML, MA, and HP: critically read and revised the manuscript for intellectual content. All authors have read and approved the final version of the manuscript. Data availabilityAll data are available in the revised manuscript. ReferencesAdnan, M., Siddiqui, A.J., Ashraf, S.A., Bardakci, F., Alreshidi, M., Badraoui, R., Noumi, E., Tepe, B., Sachidanandan, M. and Patel, M. 2023. Network pharmacology, molecular docking, and molecular dynamics simulation to elucidate the molecular targets and potential mechanism of phoenix dactylifera (Ajwa dates) against candidiasis. Pathogens 12(11), 1369. Agu, P.C., Afiukwa, C.A., Orji, O.U., Ezeh, E.M., Ofoke, I.H., Ogbu, C.O., Ugwuja, E.I. and Aja, P.M. 2023. Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Sci. Rep. 13(1), 13398. Alalami, K., Goff, J., Grimson, H., Martin, O., McDonald, E., Mirza, T., Mistry, D., Ofodile, A., Raja, S., Shaker, T., Sleibi, D. and Forget, P. 2024. Does Topical Capsaicin Affect the Central Nervous System in Neuropathic Pain? A Narrative Review. Pharmaceuticals (Basel). 17(7), 842. Antonius, Y., Kharisma, V.D., Widyananda, M.H., Ansori, A.N.M., Trinugroho, J.P., Ullah, M.E., Naw, S.W., Jakhmola, V. and Wahjudi, M. 2022. Prediction of Aflatoxin-B1 (AFB1) Molecular Mechanism Network and Interaction to Oncoproteins Growth Factor in Hepatocellular Carcinoma. J. Pure Appl. Microbiol. 16(3), 1844–1854. Ardito, F., Giuliani, M., Perrone, D., Troiano, G. and Muzio, L.L. 2017. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). Int. J. Mol. Med. 40(2), 271–280. Awuchi, C.G., Ondari, E.N., Ogbonna, C.U., Upadhyay, A.K., Baran, K., Okpala, C.O.R., Korzeniowska, M. and Guiné, R.P.F. 2021. Mycotoxins Affecting Animals, Foods, Humans, and Plants: types, Occurrence, Toxicities, Action Mechanisms, Prevention, and Detoxification Strategies-A Revisit. Foods 10(6), 1279. Badwan, B.A., Liaropoulos, G., Kyrodimos, E., Skaltsas, D., Tsirigos, A. and Gorgoulis, V.G. 2023. Machine learning approaches to predict drug efficacy and toxicity in oncology. Cell Rep. Methods 3(2), 100413. Benkerroum, N. 2020. Chronic and Acute Toxicities of Aflatoxins: mechanisms of Action. Int. J. Environ. Res. Public Health 17(2), 423. Bilal, R.M., Tahir, M.A., Shahzad, A., Farag, M.R., Siddiq, A., El-Shall, N.A., Dhama, K., Elnesr, S.S. and Alagawany, M. 2023. Yeast and derived products: their uses in preventing mycotoxins in poultry feeds. World’s. Poult. Sci. J. 79(2), 351–378. Cervello, M., Augello, G., Cusimano, A., Emma, M.R., Balasus, D., Azzolina, A., Mccubrey, J.A. and Montalto, G. 2017. Pivotal roles of glycogen synthase-3 in hepatocellular carcinoma. Adv. Biol. Regulation 65(1), 59–76. Chen, L.Q. 2019. Chemical potential and Gibbs free energy. MRS. Bull. 44(1), 520–523. Daina, A., Michielin, O. and Zoete, V. 2019. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47(1), W357–W364. Du, X., Li, Y., Xia, Y.L., Ai, S.M., Liang, J., Sang, P., Ji, X.L. and Liu, S.Q. 2016. Insights into Protein-Ligand Interactions: mechanisms, Models, and Methods. Int. J. Mol. Sci. 17(2), 144. El Khoury, G., Azzam, W. and Rebehmed, J. 2023. PyProtif: a PyMol plugin to retrieve and visualize protein motifs for structural studies. Amino Acids 55(10), 1429–1436. Elmas, C. and Gezer, C. 2021. Capsaicin and Its Effects on Body Weight. J. Am. Nutr. Assoc. 41(8), 831–839. Fattori, V., Hohmann, M., Rossaneis, A., Pinho-Ribeiro, F. and Verri, W. 2016. Capsaicin: current Understanding of Its Mechanisms and Therapy of Pain and Other Pre-Clinical and Clinical Uses. Molecules 21(7), 844. Forrest, L.R., Tang, C.L. and Honig, B. 2006. On the accuracy of homology modeling and sequence alignment methods applied to membrane proteins. Biophys. J. 91(2), 508–517. Gfeller, D., Grosdidier, A., Wirth, M., Daina, A., Michielin, O. and Zoete, V. 2014. SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res. 42(Web Server issue), W32–W38. Guex, N., Peitsch, M.C. and Schwede, T. 2009. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30(Suppl 1), S162–S173. Ivanova, L. and Karelson, M. 2022. The Impact of Software Used and the Type of Target Protein on Molecular Docking Accuracy. Molecules 27(24), 9041. Kharisma, V.D., Septiadi, L. and Syafrudin, S. 2018. Prediction of Novel Bioactive Compound from Zingiber officinale as Non-nucleoside Reverse Transcriptase Inhibitors (NNRTIs) of HIV-1 through Computational Study. Bioinform. Biomed. Res. J. 1(2), 49–55. Kim, S., Thiessen, P.A., Bolton, E.E., Chen, J., Fu, G., Gindulyte, A., Han, L., He, J., He, S., Shoemaker, B.A., Wang, J., Yu, B., Zhang, J. and Bryant, S.H. 2016. PubChem Substance and Compound databases. Nucleic Acids Res. 44(D1), D1202–D1213. Lexa, K.W. and Carlson, H.A. 2012. Protein flexibility in docking and surface mapping. Q. Rev. Biophys. 45(3), 301–343. Madushanka, A., Moura, R.T., Verma, N. and Kraka, E. 2023. Quantum Mechanical Assessment of Protein-Ligand Hydrogen Bond Strength Patterns: insights from Semiempirical Tight-Binding and Local Vibrational Mode Theory. Int. J. Mol. Sci. 24(7), 6311. Mahato, D.K., Lee, K.E., Kamle, M., Devi, S., Dewangan, K.N., Kumar, P. and Kang, S.G. 2019. Aflatoxins in Food and Feed: an Overview on Prevalence, Detection and Control Strategies. Front. Microbiol. 10(1), 2266. Marchese, S., Polo, A., Ariano, A., Velotto, S., Costantini, S. and Severino, L. 2018. Aflatoxin B1 and M1: biological Properties and Their Involvement in Cancer Development. Toxins (Basel). 10(6), 214. Martínez-Alcantar, L., Hernández-Padilla, L., Díaz-Pérez, A.L., Villalón-Magallán, L.G., Durán-Maldonado, M.X., Díaz-Pérez, C., Campos-Morales, M.E., Figueroa-Guzmán, C. and Campos-García, J. 2024. Cyclic peptides as protein kinase modulators and their involvement in the treatment of diverse human diseases. Kinases Phosphatases 2(4), 346–378. Mccarty, M.F., Dinicolantonio, J.J. and O'Keefe, J.H. 2015. Capsaicin may have important potential for promoting vascular and metabolic health. Open Heart 2(1), 262. Mohanty, M. and Mohanty, P.S. 2023. Molecular docking in organic, inorganic, and hybrid systems: a tutorial review. Monatsh. Chem. 1(1), 1–25. Musuamba, F.T., Skottheim Rusten, I., Lesage, R., Russo, G., Bursi, R., Emili, L., Wangorsch, G., Manolis, E., Karlsson, K.E., Kulesza, A., Courcelles, E., Boissel, J.P., Rousseau, C.F., Voisin, E.M., Alessandrello, R., Curado, N., Dall’Ara, E., Rodriguez, B., Pappalardo, F. and Geris, L. 2021. Scientific and regulatory evaluation of mechanistic in silico drug and disease models in drug development: building model credibility. CPT. Pharmacometrics Syst. Pharmacol. 10(8), 804–825. Orellana-Escobedo, L., Garcia-Amezquita, L.E., Olivas, G.I., Ornelas-Paz, J.J. and Sepulveda, D.R. 2012. Capsaicinoids content and proximate composition of Mexican chili peppers (Capsicum spp.) cultivated in the State of Chihuahua. CYTA. J. Food 11(2), 179–184. Pantsar, T. and Poso, A. 2018. Binding Affinity via Docking: fact and Fiction. Molecules 23(8), 1899. Pearson, G., Robinson, F., Gibson, T.B., Xu, B.E., Karandikar, M., Berman, K. and Cobb, M.H. 2001. Mitogen-activated protein (MAP) kinase pathways: regulation and physiological functions. Endocr. Rev. 22(2), 153–183. Popescu, R.G., Rădulescu, A.L., Georgescu, S.E. and Dinischiotu, A. 2022. Aflatoxins in Feed: types, Metabolism, Health Consequences in Swine and Mitigation Strategies. Toxins (Basel). 14(12), 853. Rauf, M.A., Zubair, S. and Azhar, A. 2015. Ligand docking and binding site analysis with pymol and autodock/vina. Int. J. Basic Appl. Sci. 4(2), 168–177. Roy, S.S. and Kapoor, M. 2018. In silico identification and analysis of the binding site for aminocoumarin type inhibitors in the C-terminal domain of Hsp90. J. Mol. Graph. Model. 84(1), 215–235. Saliu, T.P., Umar, H.I., Ogunsile, O.J., Okpara, M.O., Yanaka, N. and Elekofehinti, O.O. 2021. Molecular docking and pharmacokinetic studies of phytocompounds from Nigerian Medicinal Plants as promising inhibitory agents against SARS-CoV-2 methyltransferase (nsp16). J. Genet. Eng. Biotechnol. 19(1), 172. Seok, S.H. 2021. Structural insights into protein regulation by phosphorylation and substrate recognition of protein kinases/phosphatases. Life (Basel). 11(9), 957. Shabeer, S., Asad, S., Jamal, A. and Ali, A. 2022. Aflatoxin Contamination, Its Impact and Management Strategies: an Updated Review. Toxins (Basel). 14(5), 307. Shao, Z., Chen, C.Y., Chen, X., Chen, H., Su, M., Sun, H., Li, Y., Tu, B., Wang, Z. and Liu, C.M. 2023. Capsaicin exerts anti-benign prostatic hyperplasia effects via inhibiting androgen receptor signaling pathway. Biocell 47(6), 1389–1396. Snyder, P.W., Mecinovic, J., Moustakas, D.T., Thomas, S.W., Harder, M., Mack, E.T., Lockett, M.R., Héroux, A., Sherman, W. and Whitesides, G.M. 2011. Mechanism of the hydrophobic effect in the biomolecular recognition of arylsulfonamides by carbonic anhydrase. Proc. Natl. Acad. Sci. U. S. A. 108(44), 17889–17894. Sukmanadi, M., Sudjarwo, S.A. and Effendi, M.H. 2020. Molecular mechanism of capsaicin from (Capsicum annuum L.) on expression of MAPK1 and AKT1 protein as candidate of anticancer drugs: in silico study. Pharmacognosy J. 12(4), 916–919. Sulaimany, S., Farahmandi, K. and Mafakheri, A. 2024. Computational prediction of new therapeutic effects of probiotics. Sci. Rep. 14(1), 11932. Susanto, H., Kharisma, V.D., Listyorini, D., Taufiq, A., Sunaryono, S. and Aulanni’am. 2018. Effectivity of black tea polyphenol in adipogenesis related IGF-1 and its receptor pathway through in silico based study. J. Phys. Conf. Ser. 1093(1), 012037. Szallasi, A. 2022. Dietary capsaicin: a spicy way to improve cardio-metabolic health?. Biomolecules 12(12), 1783. Thongin, S., Den-Udom, T., Uppakara, K., Sriwantana, T., Sibmooh, N., Laolob, T., Boonthip, C., Wichai, U., Muta, K. and Ketsawatsomkron, P. 2022. Beneficial effects of capsaicin and dihydrocapsaicin on endothelial inflammation, nitric oxide production and antioxidant activity. Biomed. Pharmacother. 154(1), 113521. Tran, Q.H., Pham, Q.T., Nguyen, T.T., Nguyen, D.V., Le, V.T., Tran, V.D., Phan, H., Nguyen, V.H. and Nguyen, T.D. 2024. Pathogen producing aflatoxin in contaminated sandwich: identification and preservation. Microb. Pathog. 195(1), 106890. Tripathi, A. and Bankaitis, V.A. 2017. Molecular docking: from lock and key to combination lock. J. Mol. Med. Clin. Appl. 2(1), 106; doi: 10.16966/2575-0305.106 Trott, O. and Olson, A.J. 2010. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31(2), 455–461. Wang, N., Zhou, X., Zhang, T., Jian, W., Sun, Z., Qi, P., Feng, Y., Liu, H., Liu, L. and Yang, S. 2025. Capsaicin from chili peppers and its analogues and their valued applications: an updated literature review. Food Res. Int. 208(1), 116034. Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., Heer, F.T., deBeer, T.A.P., Rempfer, C., Bordoli, L., Lepore, R. and Schwede, T. 2018. SWISS-MODEL: homology modeling of protein structures & complexes. Nucl. Acids Res. 46(W1), 296–303. Xiang, Y., Xu, X., Zhang, T., Wu, X., Fan, D., Hu, Y., Ding, J., Yang, X., Lou, J., Du, Q., Xu, J. and Xie, R. 2022. Beneficial effects of dietary capsaicin in gastrointestinal health and disease. Exp. Cell Res. 417(2), 113227. Zhang, L., Shi, S., Liu, Y., Cui, Y., Zhu, Y., Bao, Y., Chen, B. and Shi, W. 2023. Aflatoxin B1 triggers apoptosis in rabbit hepatocytes via mediating oxidative stress and switching on the mitochondrial apoptosis pathway. Ecotoxicol. Environ. Saf. 264(1), 115478. Zhong, Q., Xiao, X., Qiu, Y., Xu, Z., Chen, C., Chong, B., Zhao, X., Hai, S., Li, S., An, Z. and Dai, L. 2020. Protein posttranslational modifications in health and diseases: functions, regulatory mechanisms, and therapeutic implications. MedComm 4(3), 261. | ||
| How to Cite this Article |
| Pubmed Style Sukmanadi M, Sudjarwo SA, Effendi MH, Khairullah AR, Madyawati SP, Mustofa I, Srianto P, Lamid M, Plumeriastuti H, Ahmad RZ, Akintunde AO, Awwanah M, Pratama BP, Prasetyo A. In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Vet. J.. 2025; 15(10): 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 Web Style Sukmanadi M, Sudjarwo SA, Effendi MH, Khairullah AR, Madyawati SP, Mustofa I, Srianto P, Lamid M, Plumeriastuti H, Ahmad RZ, Akintunde AO, Awwanah M, Pratama BP, Prasetyo A. In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). https://www.openveterinaryjournal.com/?mno=254742 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i10.50 AMA (American Medical Association) Style Sukmanadi M, Sudjarwo SA, Effendi MH, Khairullah AR, Madyawati SP, Mustofa I, Srianto P, Lamid M, Plumeriastuti H, Ahmad RZ, Akintunde AO, Awwanah M, Pratama BP, Prasetyo A. In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Vet. J.. 2025; 15(10): 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 Vancouver/ICMJE Style Sukmanadi M, Sudjarwo SA, Effendi MH, Khairullah AR, Madyawati SP, Mustofa I, Srianto P, Lamid M, Plumeriastuti H, Ahmad RZ, Akintunde AO, Awwanah M, Pratama BP, Prasetyo A. In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Vet. J.. (2025), [cited January 25, 2026]; 15(10): 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 Harvard Style Sukmanadi, M., Sudjarwo, . S. A., Effendi, . M. H., Khairullah, . A. R., Madyawati, . S. P., Mustofa, . I., Srianto, . P., Lamid, . M., Plumeriastuti, . H., Ahmad, . R. Z., Akintunde, . A. O., Awwanah, . M., Pratama, . B. P. & Prasetyo, . A. (2025) In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Vet. J., 15 (10), 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 Turabian Style Sukmanadi, Mohammad, Sri Agus Sudjarwo, Mustofa Helmi Effendi, Aswin Rafif Khairullah, Sri Pantja Madyawati, Imam Mustofa, Pudji Srianto, Mirni Lamid, Hani Plumeriastuti, Riza Zainuddin Ahmad, Adeyinka Oye Akintunde, Mo Awwanah, Bima Putra Pratama, and Agung Prasetyo. 2025. In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Veterinary Journal, 15 (10), 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 Chicago Style Sukmanadi, Mohammad, Sri Agus Sudjarwo, Mustofa Helmi Effendi, Aswin Rafif Khairullah, Sri Pantja Madyawati, Imam Mustofa, Pudji Srianto, Mirni Lamid, Hani Plumeriastuti, Riza Zainuddin Ahmad, Adeyinka Oye Akintunde, Mo Awwanah, Bima Putra Pratama, and Agung Prasetyo. "In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus)." Open Veterinary Journal 15 (2025), 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 MLA (The Modern Language Association) Style Sukmanadi, Mohammad, Sri Agus Sudjarwo, Mustofa Helmi Effendi, Aswin Rafif Khairullah, Sri Pantja Madyawati, Imam Mustofa, Pudji Srianto, Mirni Lamid, Hani Plumeriastuti, Riza Zainuddin Ahmad, Adeyinka Oye Akintunde, Mo Awwanah, Bima Putra Pratama, and Agung Prasetyo. "In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus)." Open Veterinary Journal 15.10 (2025), 5335-5345. Print. doi:10.5455/OVJ.2025.v15.i10.50 APA (American Psychological Association) Style Sukmanadi, M., Sudjarwo, . S. A., Effendi, . M. H., Khairullah, . A. R., Madyawati, . S. P., Mustofa, . I., Srianto, . P., Lamid, . M., Plumeriastuti, . H., Ahmad, . R. Z., Akintunde, . A. O., Awwanah, . M., Pratama, . B. P. & Prasetyo, . A. (2025) In silico study: Hepatoprotective effects of capsaicin on the liver of aflatoxin B1-intoxicated mice (Mus musculus). Open Veterinary Journal, 15 (10), 5335-5345. doi:10.5455/OVJ.2025.v15.i10.50 |