| Research Article | ||
Open Vet. J.. 2025; 15(10): 4928-4935 Open Veterinary Journal, (2025), Vol. 15(10): 4928-4935 Research Article NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversityPanca Andes Hendrawan1, Sony Hartono Wijaya2, Cece Sumantri3, Jakaria Jakaria3*1Doctoral Graduate School of Animal Production and Technology, Faculty of Animal Science, IPB University, Bogor, Indonesia 2Department of Computer Science, Faculty of Mathematics and Natural Science, IPB University, Bogor, Indonesia 3Department of Animal Production and Technology, Faculty of Animal Science, IPB University, Bogor Indonesia *Corresponding Author: Jakaria Jakaria, Department of Animal Production and Technology, Faculty of Animal Science, IPB University, Bogor, Indonesia. Email: jakaria [at] apps.ipb.ac.id Submitted: 17/06/2025 Revised: 21/08/2025 Accepted: 01/09/2025 Published: 31/10/2025 © 2025 Open Veterinary Journal
AbstractBackground: Genetic diversity plays a critical role in the conservation and enhancement of genetic resources, including indigenous livestock populations. Aim: This study aimed to analyze the genetic diversity of the Jambi buffalo (Bubalus bubalis) population based on the mitochondrial NADH dehydrogenase subunit 1 (ND1) gene. Methods: This study analyzed 93 samples from five subpopulations in Jambi: Tebo (23), Batanghari (21), Merangin (9), Kerinci (13), and Tanjung Jabung Barat (9), as well as comparison samples from North Sumatra/Sumut (6), Banten (4), and Riau (8). Sequencing of the ND1 gene was performed, and the resulting data were analyzed using MEGA11, BioEdit, DnaSP6, and NETWORK software. Results: In this study, seven haplotypes were identified, with haplotype and nucleotide diversity values of 0.1184 and 0.0015, respectively. Analysis of haplotypes indicated that the Jambi buffalo populations (Tebo, Batanghari, and Kerinci) did not receive gene flow from other groups and were probably the original ancestors of other buffalo populations in Jambi Province. Phylogenetic tree analysis further revealed that Jambi buffaloes share common ancestry with populations from China and mainland Southeast Asia, including Thailand, Laos, Vietnam, and Myanmar. Conclusion: Based on the mitochondrial ND1 gene, the genetic diversity of Jambi buffalo is low. Keywords: Bubalus bubalis, Haplotype diversity, NADH dehydrogenase subunit 1. IntroductionIndonesia is characterized by a high diversity of AnGR. These genetic resources play a vital role in ensuring the sustainable fulfillment of food needs. Among the potential genetic resources of livestock, the buffalo holds significant importance. Indonesia has identified 14 distinct local buffalo populations, each of which is associated with specific geographical regions. These include the Pampangan, Simeulue, Kuntu, Toraya, Javanese, Gayo, Kalang South Kalimantan, Kalang East Kalimantan, Moa, Murrah, Anoa, North Sumatra, West Sumatra, and Sumbawa buffaloes (FAO, 2023). Swamp buffaloes (Bubalus bubalis) exhibit considerable adaptability to diverse environmental conditions and low-quality feed, facilitating their widespread distribution throughout Indonesia, including Jambi Province. Buffaloes are significant in various traditional ceremonies and cultural practices, such as wedding processions, customary legal fines (Kerlogue, 2011), thanksgiving events (Fadilah et al., 2019), ritual slaughter during the welcoming of Ramadan (bantai adat), and as sacrificial animals (Kurniadi and Putri, 2021). Consequently, buffalo livestock conservation is of considerable importance. However, over the past decade, the buffalo population in Jambi Province has shown a declining trend (Hendrawan et al., 2025). If this decline continues unchecked, it may result in a severe reduction in population size and potentially lead to long-term extinction. The analysis of genetic diversity is one strategy for conserving animal genetic resources (Pineda et al., 2021). Genetic diversity refers to the variance of genes both within and among a species’ populations. Diversity is a crucial element of biodiversity and significantly contributes to the long-term survival and adaptability of species (Kumar, 2023). Mitochondrial DNA markers can be used to analyze livestock genetic diversity. Mitochondrial DNA can be used in maternal genetic inheritance studies (Sun et al., 2020a), comparisons between breeds (Jakaria et al., 2019), evolutionary studies (Singh et al., 2020), domestication areas, and phylogenetic analyses (Kumar et al., 2007). Mitochondrial DNA consists of a displacement loop (D-loop) region and 37 genes [13 protein parts (polypeptides), 22 transfer RNA parts, and two small (12S) and large (16S) ribosomal RNA parts]. The study of the genetic diversity of buffalo using mitochondrial DNA markers usually focuses on the D-loop region (Suhardi et al., 2023), Cyt b (Winaya et al., 2019; Rusdin et al., 2020), and COI (Saputra et al., 2013). Unfortunately, there are no reports on the genetic diversity of buffalo based on NADH dehydrogenase subunit 1 (ND1) gene sequences. NADH is a coenzyme that transports electrons from metabolic reactions, such as glycolysis and the citric acid cycle, to the electron transport chain in the mitochondria. This process produces energy in the form of adenosine triphosphate, which is essential for cellular activity in livestock (Xie et al., 2020). The ND1 gene of mitochondrial DNA is known for its high diversity, making it a promising marker for the analysis of genetic diversity. ND1 gene diversity has been reported in beef cattle (Mulyati et al., 2025), Iraqi cattle (Muhaidi et al., 2017), and Sumba Ongole and Peranakan Ongole cattle (Mubarak et al., 2019). However, there have been no reports on the genetic diversity of Jambi buffaloes based on the ND1 gene. The use of the ND1 gene as a genetic marker is particularly significant because of its high variability and its essential role in mitochondrial function, providing new insights into maternal lineage and evolutionary history. The findings of this study are expected to fill a critical knowledge gap, inform conservation strategies, and contribute to the sustainable management and preservation of local buffalo genetic resources in Indonesia and the world. Therefore, this study aimed to analyze the genetic diversity of Jambi buffaloes based on the ND1 gene. Materials and MethodsEthical approvalThis study was approved by the Ethics Commission of the Faculty of Medicine, Universitas Andalas, Indonesia, with license number 588/UN.16.2/KEP-FK/2023. Sample collectionA total of 93 DNA samples were analyzed in this study, including five buffalo sub-populations from Jambi Province: Tebo (23), Batanghari (21), Merangin (9), Kerinci (13), and Tanjung Jabung Barat (9), as well as samples from the Genetic and Molecular Laboratory of the Faculty of Animal Science, IPB University, including North Sumatra/Sumut (6), Banten (4), and Riau (8). DNA extraction, amplification, and sequencingDNA was extracted from blood samples collected in EDTA tubes using a DNA Extraction Kit (Qiagen). DNA purity was evaluated using a NanoDrop one microvolume UV-Vis Spectrophotometer (Thermo Fisher Scientific, USA). ND1 gene amplification with a product length of 1,013 bp was amplified with a pair of primers: forward 5’CAG AGC CCG GTA ATT GCA TA’3 and reverse 5’GGG GAA TGC TTG ATA GGA GAA3’. Primers were designed using Primer 3 software (primer3.ut.ee) based on the complete sequence of Gene Bank Accession No. (KX758395.1). PCR was performed with a total volume of 25 μl, consisting of 50 ng/μl template DNA, 25 pmol primers, GoTaq® Red master mix, and distilled water. The amplification of DNA fragments was performed using an AB System PCR machine. DNA amplification consists of 35 repeated cycles of predenaturation at 95°C for 5 minute, denaturation at 95°C for 30 second, annealing at 62°C for 45 second, and extension at 72°C for 1 minute . The final extension was performed at 72°C for 5 minute. PCR products were visualized on a 1% agarose gel in 0.5× tris borate EDTA buffer. Further sequencing of PCR products of the ND1 gene was performed using the services of the 1st Base Malaysia. Data analysisThe ND1 gene sequence was aligned, and a phylogenetic tree was constructed using MEGA 12 and BioEdit version 7.7.1.0 (Hall et al., 2011; Kumar et al., 2024). The ClustalW method was used for alignment. The phylogenetic tree was constructed using the maximum likelihood method with Kimura-2 parameters with 1,000 bootstrap replications, a gamma distribution (+G) with five rate categories, and evolutionary invariance (+I). Reference sequences accessed from the NCBI GenBank were also used to construct the phylogenetic tree (Fig. 4). Haplotype diversity (Hd), nucleotide diversity (π), polymorphic sites (S), and Fu’s values were analyzed using DnaSP6 version 6.12.1 (Rozas et al., 2017). The median-joining network was constructed using the NETWORK software version 10.2.00 (Kong et al., 2016).
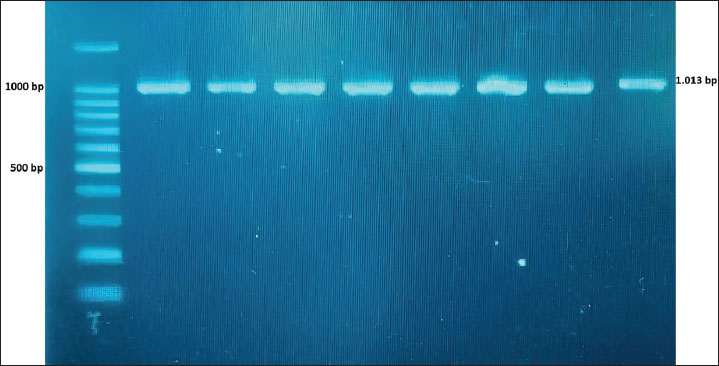
Fig. 1. PCR amplification of the mitochondrial ND1 gene in swamp buffalo (Bubalus bubalis).
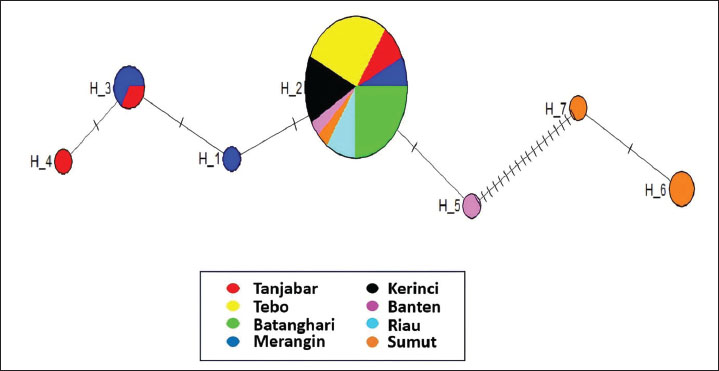
Fig. 2. Median-joining network of the ND1 gene haplotypes in buffalo populations from Jambi (Tanjabar, Tebo, Batanghari, Merangin, and Kerinci), Banten, Riau, and Sumut.
Fig. 3. Geographic distribution map illustrating the frequency of buffalo haplotypes across eight distinct populations. Hap 1 (red), Hap 2 (yellow), Hap 3 (green), Hap 4 (blue), Hap 5 (sky blue), Hap 6 (orange), and Hap 7 (black).
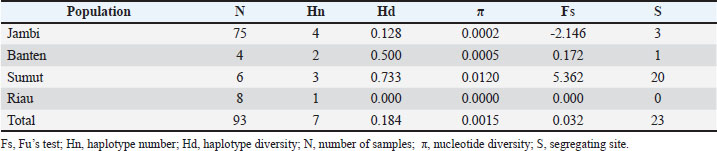
Fig. 4. Phylogenetic tree of buffaloes constructed based on ND1 gene sequences using maximum likelihood analysis ResultAnalysis of haplotype and genetic diversityThe ND1 gene was successfully amplified with a product length of 1,013 bp (Fig. 1). The ND1 sequence of buffalo has a length of 952 base pairs (bp) and has been submitted to the gene bank (NCBI: access number PX210530). The Jambi buffalo population had more haplotypes (Hn) than the North Sumatra/Sumut, Banten, and Riau populations, but the Hd and nucleotide diversity (π) were lower than those of the North Sumatra/Sumut and Banten populations (Table 1). This study reports the first ND1 gene in buffalo. The Fu’s test (Fs) value shows that only the Jambi buffalo population has a negative value, indicating population expansion or gene flow after experiencing a bottleneck due to a large population decline resulting in inbreeding. Table 1. Genetic diversity of the Jambi buffaloes (Tebo, Merangin, Tanjabar, Kerinci, and Batanghari), Banten, Sumut, and Riau.
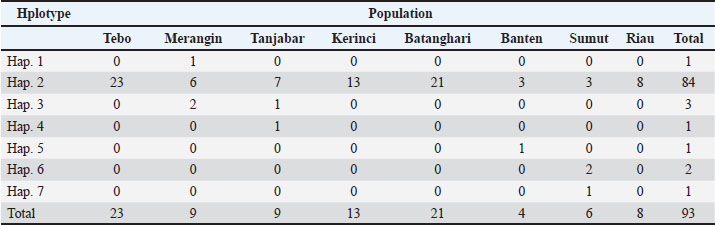
Haplotype distribution and network structureTable 2 shows the haplotype distribution of the 93 analyzed samples. The haplotype distribution showed that 93.33% of the observed buffaloes were classified as haplotype 2. The median-joining network (Fig. 2) showed that haplotype 2 is a common ancestor within the population, indicating that buffaloes in Jambi (Tanjung Jabung Barat and Merangin), Banten, North Sumatra/Sumut, and Riau share a common ancestor. We also found that three Jambi buffalo populations (Tebo, Batanghari, and Kerinci) did not introduce gene flow from other populations (Fig. 3). Table 2. Distribution of haplotypes in buffalo populations from Jambi, Banten, Sumut, and Riau.
Genetic distance and phylogenetic analysisThe pairwise distance between the Jambi, Banten, North Sumatra/Sumut, and Riau buffaloes shows that the North Sumatra/Sumut population has a large genetic distance from the Jambi, Banten, and Riau populations. In contrast, the Jambi, Banten, and Riau populations have relatively small genetic distances (Table 3). Pairwise distances for the five observed buffalo populations show that geographic factors do not affect the genetic distance between populations. The phylogenetic tree of buffaloes in this study shows that the Jambi, Banten, and Riau buffaloes are in the same cluster as Chinese buffalo populations and buffaloes from mainland Southeast Asia, such as Thailand, Myanmar, Laos, and Vietnam (Fig. 4). This finding could indicate the possibility of gene flow or shared ancestry from mainland Southeast and South/East Asia. Table 3. Pairwise genetic distances among Jambi buffalo populations in Banten, Sumut, and Riau Provinces
DiscussionAnalysis of haplotype and genetic diversityThe number of Hn and Hd in this study were relatively low compared with those in previous studies on local buffalo from various regions in Indonesia. Rusdin et al. (2020) used 78 samples in the Cyt b region, which included 10 Hn and Hd of 0.659. Saputra et al. (2013) used 28 samples in the COI region and found three Hn, whereas (Suhardi et al., 2023) used 120 Borneo buffalo samples in the D-loop region, with 47 Hn and Hd of 0.936. In this study, the Hd is low, which is also in line with the low value of nucleotide diversity (π). According to Nei and Kumar (2000), the Hd value has two categories: low (<0.50) and high (0.51–1.00). Meanwhile, the value of μ in research on experimental animals in each population falls within the low category. The μ value is classified into three levels: low (0.01–0.04), moderate (0.05–0.07), and high (0.08–1.00). The number of haplotypes and Hd were also consistent with the segregation site (S) in the observed population. Another possible hypothesis is that the low genetic diversity in this population is caused by a population bottleneck event due to a significant decline in population size over the past 10 years. Jager et al. (2025) reported low genetic diversity in Cape Buffalo, which was strongly suspected to be caused by a bottleneck event resulting from a previous drastic decline in population size. Given the low genetic diversity in the Jambi banteng population, establishing a conservation program that not only aims to increase genetic variation but also preserves the unique local genetic characteristics of the Jambi banteng is crucial. The introduction of genetic material from external sources must be done carefully to avoid the loss of the original genetic identity, which has been shaped through adaptation to the local environment and cultural practices in Jambi. The observed negative Fu’s Fs test values are consistent with the three Jambi buffalo populations, Batanghari, Tebo, and Kerinci, which belong to the same haplogroup. Similar findings of negative Fu’s Fs values have been reported in studies on Kalimantan buffalo populations (Suhardi et al., 2023) and Sragen Black Cattle (Kusumaningrum et al., 2020). Haplotype distribution and network structureThe median-joining network suggests that buffalo populations in Merangin, Tanjung Jabung Barat, North Sumatra/Sumut, and Banten likely received genetic introductions from larger populations, such as Batanghari, Tebo, and Kerinci. The diversity of haplotypes in these populations, including the dominant haplotype (haplotype 2), which is also found in Tebo, Kerinci, and Batanghari, supports this hypothesis. Batanghari and Tebo are the regions with the largest buffalo populations (Jambi Province Central Statistics Agency, 2024), and they serve as the main breeding stock source (Hendrawan et al., 2025). Populations identified as not receiving gene flow, such as those from Tebo, Batanghari, and Kerinci, should be prioritized as local germplasm sources. Breeding interventions or seedling exchanges must consider the importance of maintaining local genetic purity and uniqueness to preserve the adaptive diversity and cultural value of the Jambi buffalo. Genetic distance and phylogenetic analysisNorth Sumatra/Sumut buffaloes are in a distinct cluster with Indian buffaloes and Bubalus arnee, which are wild Asian buffaloes, indicating that they have a strong ancestry or relationship with Indian buffaloes. Evidence supporting the presence of the Indian river buffalo in the North Sumatra/Sumut region has been further substantiated by the government’s crossbreeding program initiated in 2015, which involves hybridization between swamp buffalo and river buffalo through AI. The semen used in these programs was sourced from the North Sumatra/Sumut Artificial Insemination Center and the Lembang facility in Bandung (Aritonang et al., 2018). The North Sumatra/Sumut buffalo is geographically closer to the Jambi and Riau populations, but has a large genetic distance from these two populations. In contrast, Banten buffaloes have a close genetic distance from the Jambi and Riau populations. According to Sukri et al. (2014), the genetic distance of livestock can be influenced by its biogeographic location or the presence of barriers (e.g., mountains, rivers, and forests). However, (Pauciullo et al., 2025) explained that genetic distance is influenced by many more complex factors, such as environmental adaptation, limited gene flow, and breeding practices. These findings may be related to the fact that some populations are more open due to easily accessible transportation routes that facilitate buffalo migration (Anggraeni et al., 2011). According to Sun et al. (2020b), swamp buffalo were domesticated gradually and in multiple places, with the Lower Yangtze region of China being the initial center of domestication. Thereafter, it spread across the region via two major migration routes, during which several additional domestication events occurred in other regions, such as the Upper Yangtze, southwest China, and northern Southeast Asia. Based on cyt b sequences, Indonesian buffaloes have two maternal lines: cluster 1: buffaloes from B. arnee, Bangladesh, China, Indonesia, Laos, Nepal, Taiwan, Thailand, and Vietnam; and cluster 2: buffaloes from Bangladesh, China, India, Indonesia, Laos, Nepal, Taiwan, Thailand, and Vietnam (Saputra et al., 2021). ConclusionThis study shows that Jambi buffaloes have low ND1 gene genetic diversity and a limited number of haplotypes. The buffalo populations from Jambi, Banten, and Riau share the same ancestors, whereas the buffalo populations from North Sumatra/Sumut show closer genetic similarities to the Indian buffaloes. These findings indicate that gene flow and breeding practices significantly influence genetic distance, surpassing the effects of geographical separation alone. Improving the genetic quality of Jambi buffaloes can be achieved by introducing buffaloes, provided that the region of origin is preserved to maintain genetic purity that has adapted to the surrounding environment. AcknowledgmentsThe authors would like to thank the Jambi Provincial Government for providing assistance through the relevant agencies in the districts of Tebo, Batanghari, Merangin, Tanjung Jabung Barat, and Kerinci. Conflict of interestThe authors have no conflicts of interest to declare. FundingThe Ministry of Education, Culture, Research, and Technology of the Republic of Indonesia provided financial support with Grant No.006/C3/DT.05.00/PL/2025 dan 23129/IT3.D10/PT.01.03/P/B/2025. Author’s contributionsAll authors have contributed to this research. PAH was responsible for data collection and drafting of the manuscript. JK, CS, and SHW contributed to the critical revision of the manuscript. All authors have read and approved the final version of the manuscript. Data availabilityAll data are included in the revised manuscript. ReferencesAnggraeni, A., Sumantri, C., Praharani, L., Dudi. and Andreas, E. 2011. Genetic distance estimation of local swamp buffaloes through morphology analysis approach. Jitv 16(3), 199–210; http://repository.ipb.ac.id/handle/123456789/76567 Aritonang, N.S., Roza, E., Rizqan. and Habsani, W. 2018. Growth traits of swamp and murrah buffalo crosses in North Sumatra, Indonesia. Arch. Anesthesiol. Crit. Care 11(10), 527–534. Fadilah, I., Sari, R.I., Ramadhani, V., Basuki, F.R. and Fitaloka, O. 2019. Ethnoscience study of the application and delivery procession of adat melayu Jambi as science learning resources. Sci. Educ. 8(2), 141–153; doi:10.24235/sc.educatia.v8i2.4428 FAO. 2023. Domestic animal diversity information system (DAD-IS). Available via https://www.fao.org/dad-is/data/en. Hall, T., Biosciences, I. and Carlsbad, C. 2011. BioEdit: an important software for molecular biology. GERF Bull. Biosci. 2, 60–61. Hendrawan, P.A., Wijaya, S.H., Sumantri, C. and Jakaria, J. 2025. Assessing the population structure and inbreeding rates of buffaloes in Batanghari District, Indonesia. World’s. Vet. J. 15(1), 49–55. Jager, D.D., Möller, M., Hoal, E., Helden, P.V., Glanzmann, B., Harper, C. and Bloomer, P. 2025. A highly divergent mitochondrial genome in extant Cape buffalo from Addo Elephant National Park, South Africa. Ecol. Evol. 15, 1–13; doi: 10.1002/ece3.70640 Jakaria, J., Musyaddad, T., Rahayu, S., Muladno, M. and Sumantri, C. 2019. Diversity of D-loop mitochondrial DNA (mtDNA) sequence in Bali and Sumba Ongole cattle breeds. J. Indones. Trop. Anim. Agric. 44(4), 335–345; doi:10.14710/jitaa.44.4.335-345 Jambi Province Central Statistics Agency [BPS]. 2024. Buffalo population by district (head). Availiable via https://jambi.bps.go.id/id/publication/2024/02/28/e697127048eeb86bba8d8d40/provinsi-jambi-dalam-angka-2024.html Kerlogue. and F. 2011. Memory and material culture a case study from Jambi, Sumatra. Indones. Malays. World 39(113), 89–101; doi:10.1080/13639811.2011.547731 Kong, S., Sánchez‐Pacheco, S.J. and Murphy, R.W. 2016. On the use of median-joining networks in evolutionary biology. Cladistics 32(6), 691–699; doi:10.1111/cla.12147 Kumar M. 2023. The importance of genetic diversity in conservation biology. J. Biodivers. Endanger Species 11(03), 10–11; doi: 10.37421/2332-2543.2023.11.477 Kumar, S., Nagarajan, M., Sandhu, J.S., Kumar, N., Behl, V. and Nishanth, G. 2007. Mitochondrial DNA analyses of Indian water buffalo support a distinct genetic origin of river and swamp buffalo. Anim. Genet. 38(3), 227–232; doi:10.1111/j.1365-2052.2007.01602.x Kumar, S., Stecher, G., Suleski, M., Sanders, M., Sharma, S. and Tamura, K. 2024. MEGA12: molecular Evolutionary Genetics Analysis version 12 for adaptive and green computing. Mol. Biol. Evol. 41(12), A1–A9; doi:10.1093/molbev/msae263 Kurniadi, M.D.K. and Putri, H.M. 2021. Bantai adat tradition: local wisdom welcoming the month of Ramadan, Jambi's Merangin community. J. Lekt. Keagamaan. 19(2), 388–418; doi:10.31291/jlka.v19i2.961 Kusumaningrum, R., Sutopo, S. and Kurnianto, E. 2020. Genetic diversity of Sragen Black Cattle based on D-Loop gene sequencing analysis. Livest. Anim. Res. 18(2), 124; doi:10.20961/lar.v18i2.42934 Mubarak, S., Nurgiartiningsih, V.M.A., Susilorini, T.E. and Agung, P.P. 2019. Genetic length and variation in mtDNA nadh dehydrogenase subunit I (ND1) of Sumba Ongole (SO) and Peranakan Ongole (PO) cattle. Int. Res. J. Adv. Eng. Sci. 4(3), 103–106. Muhaidi, M.J., Masyam, N. and Dagash, T. 2017. Determination the causative strain for hydatid cyst in Iraqi cattle by using ND1 gene. Iraqi J. Agric. Sci. 48(2), 466; doi:10.36103/IJAS.V48I2.432 Mulyati, N., Bonawati, Y. G. M., Noor, R. R., Ulum, M. F. and Jakaria. 2025. ND1 Gene Analysis in beef cattle for sustainable genetic resource management. In: IOP Conf Ser Earth Environ Sci. 1484, 1–8; doi: 10.1088/1755-1315/1484/1/012003 Nei, M. and Kumar, S. 2000. Molecular evolution and phylogenetics. Oxford, NY: University Press. Pauciullo, A., Gaspa, G., Versace, C., Cosenza, G., Piscopo, N., Gu, M., Coletta, A., Hussain, T., Seidavi, A. and Nicolae, I. 2025. New insights into genetic diversity and differentiation of 11 buffalo populations using validated SNPs for dairy improvement. Genes (Basel). 16(4), 1–13; doi:10.3390/genes16040400 Pineda, P.S., Flores, E.B., Herrera, J.R.V. and Low, W.Y. 2021. Opportunities and challenges for improving the productivity of swamp buffaloes in Southeast Asia. Front. Genet. 12, 1–8; doi:10.3389/fgene.2021.629861 Rozas, J., Ferrer-Mata, A., Sánchez-Delbarrio, J.C., Guirao-Rico, S., Librado, P., Ramos-Onsins, S.E. and Sánchez-Gracia, A. 2017. DnaSP 6: dNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34(12), 3299–3302; doi:10.1093/molbev/msx248 Rusdin, M., Solihin, D.D., Gunawan, A., Talib, C. and Sumantri, C. 2020. Genetic variation of eight indonesian swamp-buffalo populations based on cytochrome b gene marker. Trop. Anim. Sci. J. 43(1), 1–10; doi:10.5398/tasj.2020.43.1.1 Saputra, F., Anggraeni, A., Ishak, A.B.L., Hafid, A., Rusdin, M. and Sumantri, C. 2021. Haplotype diversity of swamp buffalo and river buffalo based on cytochrome b gene: a study of meta,analysis. Trop. Anim. Sci. J. 44(4), 399–407; doi:10.5398/tasj.2021.44.4.399 Saputra, F., Jakaria, J. and Sumantri, C. 2013. Genetic variation of mtDNA cytochrome oxidase subunit i (COI) in Local swamp buffaloes in Indonesia. Media Peternak. 36(3):165–170; doi:10.5398/medpet.2013.36.3.165 Singh, R., Lava Kumar, S., Mishra, S.K., Gurao, A., Niranjan, S.K., Vohra, V., Dash, S.K., Rajesh, C. and Kataria, R.S. 2020. Mitochondrial sequence-based evolutionary analysis of riverine–swamp hybrid buffaloes of India indicates novel maternal differentiation and domestication patterns. Anim. Genet. 51(3), 476–482; doi:10.1111/age.12938 Suhardi, S., Summpunn, P. and Wuthisuthimethavee, S. 2021. MtDNA D-loop sequence analysis of Kalang, Krayan, and Thale Noi buffaloes (Bubalus bubalis) in Indonesia and Thailand reveal genetic diversity. J. Indones. Trop. Anim. Agric. 46(2), 93–105; doi:10.14710/jitaa.46.2.93-105 Suhardi, S., Wibowo, A., Putra, W.P.B. and Summpunn, P. 2023. The phylogeny of bornean swamp buffalo (Bubalus bubalis) analysis based on D-loop mitochondrial DNA sequence variation. Trop. Anim. Sci. J. 46(2), 139–145; doi: 10.5398/tasj.2023.46.2.139 Sukri, A., Amin, M., Winaya, A. and Gofur, A. 2014. Substitution and haplotype diversity analysis on the partial sequence of the mitochondrial dna cyt b of Indonesian swamp buffalo (Bubalus bubalis). Biol. Med. Chem. 3(2), 59–63; doi:10.14421/biomedich.2014.32.59-63 Sun, T., Shen, J., Achilli, A., Chen, N., Chen, Q., Dang, R., Zheng, Z., Zhang, H., Zhang, X., Wang, S., Zhang, T., Lu, H., Ma, Y., Jia, Y., Capodiferro, M.R., Huang, Y., Lan, X., Chen, H., Jiang, Y. and Lei, C. 2020a. Genomic analyses reveal distinct genetic architectures and selective pressures in buffaloes. Gigascience 9(1), 1–12; doi:10.1093/gigascience/giz166 Sun, T., Wang, S., Hanif, Q., Chen, N., Chen, H. and Lei, C. 2020b. Genetic diversity of mitochondrial cytochrome b gene in swamp buffalo. Anim. Genet. 51(6), 977–981; doi:10.1111/age.12997 Winaya, A., Sukri, A., Gofur, A. and Amin, M. 2019. The genetic divergence and phylogenetic relationship of Indonesia swamp buffalo (Bubalus bubalis) based on partial sequences of cytochrome b gene of mitochondrial DNA. Int. J. Eng. Technol. 8(1.9), 96–100. Xie, N., Zhang, L., Gao, W., Huang, C., Huber, P.E., Zhou, X., Li, C., Shen, G. and Zou, B. 2020. NAD+ metabolism: pathophysiologic mechanisms and therapeutic potential. Nat. Rev. Drug Discov. 5(1), 1–35; doi:10.1038/s41392-020-00311-7 | ||
| How to Cite this Article |
| Pubmed Style Hendrawan PA, Wijaya SH, Sumantri C, Jakaria J. NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Vet. J.. 2025; 15(10): 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 Web Style Hendrawan PA, Wijaya SH, Sumantri C, Jakaria J. NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. https://www.openveterinaryjournal.com/?mno=265113 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i10.12 AMA (American Medical Association) Style Hendrawan PA, Wijaya SH, Sumantri C, Jakaria J. NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Vet. J.. 2025; 15(10): 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 Vancouver/ICMJE Style Hendrawan PA, Wijaya SH, Sumantri C, Jakaria J. NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Vet. J.. (2025), [cited January 25, 2026]; 15(10): 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 Harvard Style Hendrawan, P. A., Wijaya, . S. H., Sumantri, . C. & Jakaria, . J. (2025) NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Vet. J., 15 (10), 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 Turabian Style Hendrawan, Panca Andes, Sony Hartono Wijaya, Cece Sumantri, and Jakaria Jakaria. 2025. NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Veterinary Journal, 15 (10), 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 Chicago Style Hendrawan, Panca Andes, Sony Hartono Wijaya, Cece Sumantri, and Jakaria Jakaria. "NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity." Open Veterinary Journal 15 (2025), 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 MLA (The Modern Language Association) Style Hendrawan, Panca Andes, Sony Hartono Wijaya, Cece Sumantri, and Jakaria Jakaria. "NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity." Open Veterinary Journal 15.10 (2025), 4928-4935. Print. doi:10.5455/OVJ.2025.v15.i10.12 APA (American Psychological Association) Style Hendrawan, P. A., Wijaya, . S. H., Sumantri, . C. & Jakaria, . J. (2025) NADH dehydrogenase subunit 1 gene sequences in Jambi buffalo: Implications for genetic diversity. Open Veterinary Journal, 15 (10), 4928-4935. doi:10.5455/OVJ.2025.v15.i10.12 |