| Research Article | ||
Open Vet. J.. 2026; 16(1): 458-468 Open Veterinary Journal, (2026), Vol. 16(1): 458-468 Research Article Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, IndonesiaAlya Amaliah1, Ni Luh Putu Ika Mayasari2, Ryan Septa Kurnia3, Christian Marco Hadi Nugroho3, Muhammad Ade Putra3 and Agustin Indrawati2*1Master of Animal Biomedical Sciences, School of Veterinary Medicine and Biomedical Sciences, IPB University, Bogor, Indonesia 2Division of Medical Microbiology, School of Veterinary Medicine and Biomedical Sciences, IPB University, Bogor, Indonesia 3Research and Diagnostics Unit, PT Medika Satwa Laboratories, Bogor, Indonesia *Corresponding Author: Agustin Indrawati. Division of Medical Microbiology, School of Veterinary Medicine and Biomedical Sciences, IPB University, Bogor, Indonesia. Email: indraseta [at] apps.ipb.ac.id Submitted: 04/08/2025 Revised: 22/11/2025 Accepted: 09/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
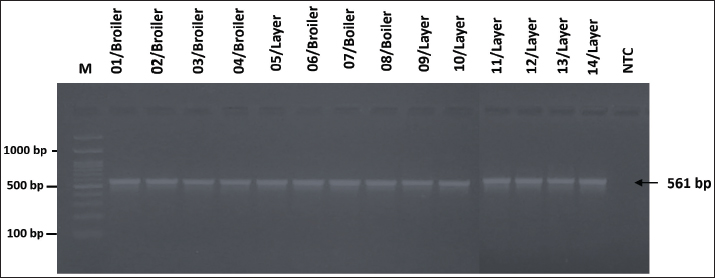
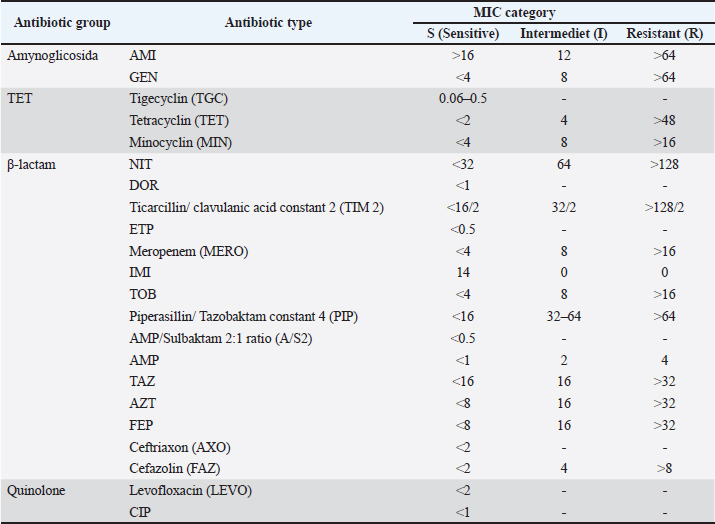
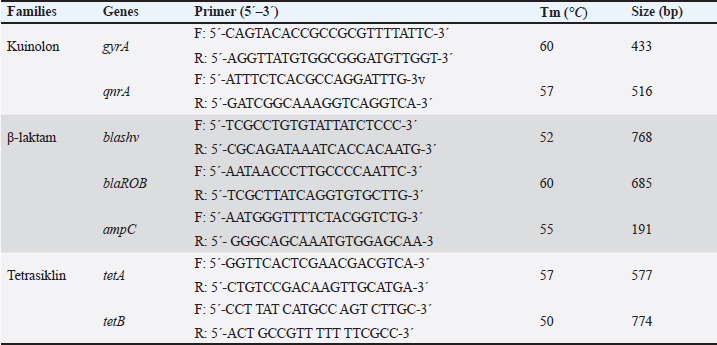
AbstractBackground: Gallibacteriosis is a disease in poultry caused by the bacterium Gallibacterium anatis that can lead to respiratory disorders or decreased production. This bacterium causes infections that are similar to those caused by other bacteria, which may lead to misdiagnosis. Infections caused by G. anatis are increasingly associated with reduced susceptibility to antibiotics. Aim: This study aimed to investigate the antibiotic resistance, antibiogram profile, and antibiotic resistance genes associated with emerging G. anatis infections in poultry. Methods: Using a targeted sampling method, G. anatis was isolated from broiler, layer, and breeder chickens exhibiting respiratory disorders and reduced egg production in eight poultry farms in West Java, Indonesia. Bacteriological isolation was performed, and the recovered isolates were subsequently tested for antimicrobial susceptibility by determining the minimum inhibitory concentration (MIC) and by polymerase chain reaction (PCR)-based detection of antibiotic resistance genes. Results: A total of 14 G. anatis isolates were obtained. The resistance profiles indicated that resistance was present in the tetracycline (TET), quinolone, and β-lactam groups. TET, ciprofloxacin, and levofloxacin antibiotics had the highest rates of resistance at 100%. Furthermore, 75.14% of the isolated G. anatis were multidrug- resistant. None of the isolates experienced resistance to amikacin, gentamicin, tigecycline, imipenem, tobramycin, and piperacillin (0%). The multiple antibiotic resistance index ranged from 0.13 to 0.59. PCR revealed that the G. anatis isolate in this study carried the following genes: gyr(A) (100%), tet(B) (92.85%), blaSHV (87.5%), and qnrA (85.71%). Conclusion: Antibiotic resistance testing of G. anatis using the MIC method revealed that many isolates were resistant to tetracyclines, β-lactams, and quinolones, which was supported by the detection of the resistance genes tet(B), blaSHV, gyrA, and qnrB. This study advances the understanding of antibiotic resistance in G. anatis strains circulating in Indonesia, highlighting resistance to antibiotics commonly used in poultry and human medicine. Keywords: Antimicrobial, Emerging, G. anatis, MDR, Resistance. IntroductionGallibacterium anatis was previously identified as Actinobacillus salpingitidis or Pasteurella salpingitidis. In 2003, it was reclassified as Gallibacterium. This bacterium is Gram-negative, non-motile, and encapsulated (Christensen et al., 2003). Gallibacterium. anatis primarily infects poultry but has also been isolated from other avian species and occasionally from mammals (Van Driessche et al., 2020). This bacterium possesses several virulence factors, including fimbriae, outer membrane vesicles, a capsule, biofilm, and repeats-in-toxin (Tang et al., 2020; Sanchez-Alonso et al., 2023). Infection with this organism in poultry can lead to reduced egg production and respiratory disorders. The disease often lacks pathognomonic clinical signs; however, peritonitis and salpingitis are frequently observed (Rzewuska et al., 2007). In broiler chickens, the clinical signs include cumulative mortality ranging from 0.06% to 4.9% per week. Such conditions have been associated with considerable economic losses to the poultry industry (Elbestawy et al., 2008; Yaman and Sahan Yapicier, 2019). Several studies on G. anatis have reported resistance to multiple antibiotic classes, including tetracycline (TET) (Hess et al., 2020), enrofloxacin, tilmicosin, amoxicillin (Kursa et al., 2023), and sulfamethoxazole (Karwańska et al., 2023). There are several categories of antimicrobial susceptibility: multidrug-resistant (MDR), extensively drug-resistant (XDR), and pandrug-resistant (PDR). MDR is defined as resistance to ≥1 antibiotic agents in ≥3 antibiotic groups. XDR refers to resistance to ≥1 antibiotic agents in all but sensitive in ≥1 groups antibiotics, while PDR is resistance to all agents in all antibiotic groups (Magiorakos et al., 2012). Antimicrobial resistance (AMR) has emerged as a critical global health challenge and is increasingly described as the next “silent pandemic” (Ahmed et al., 2024). Addressing this issue requires a comprehensive One Health perspective, as resistance genes can circulate among humans, animals, and the environment, and the complex interconnections among these sectors demand coordinated and cross-disciplinary actions (Shrestha et al., 2018; Collignon and Mcewen, 2019). The continued use of antibiotics for prophylactic purposes remains a major driver of resistance in the poultry industry, contributing to the amplification and dissemination of AMR across these interconnected domains (Liang et al., 2023). This problem is further aggravated by the emergence of MDR pathogens that can spread within communities (Duin and Peterson, 2022). The rising incidence of MDR bacteria from diverse reservoirs underscores the urgent need to strengthen our understanding of prudent antibiotic use and to advance effective strategies for managing pathogenic infections. To date, no published studies have documented the occurrence of antibiotic resistance or resistance-associated genes in emerging G. anatis isolates in Indonesia. Accordingly, this study aims to provide the first report on antimicrobial susceptibility patterns and resistance genes commonly harbored by G. anatis recovered from poultry in West Java, Indonesia. Materials and MethodsOrigin of the bacterial isolatesA total of 14 archival G. anatis isolates were used in this study. The isolates originated from poultry farms in West Java, Indonesia, between 2020 and 2023, from chickens exhibiting decreased egg production or respiratory disorders. Organ samples (trachea or ovary) were placed in sterile plastic bags, and tracheal swabs were transported in Amies transport medium (Labware Charuzu, Japan). To minimize bacterial overgrowth, all samples were delivered to the laboratory in a cold box and processed within 48 hours. The samples were cultured on blood agar, and G. anatis was molecularly confirmed using polymerase chain reaction (PCR) targeting species-specific genes. Each isolate was routinely subcultured on blood agar (HiMedia Laboratories, USA) and incubated at 37°C for 24–48 hours before use. The same isolate collection has also been used in a separate manuscript currently under review; however, the present study’s objectives, analyses, and research questions are distinct. A complete list of the isolates included in this study is provided in Additional File 1. Antimicrobial susceptibility testing (AST)The AST was performed using the minimum inhibitory concentration (MIC) method with the GN4F Sensititre™ plate (Thermo Fisher Scientific, Waltham, MA). The antibiotics included in this panel represented several pharmacological classes, namely aminoglycosides [amikacin (AMI), gentamicin (GEN), tobramycin (TOB)], tetracyclines [tigecycline (TGC), TET, minocycline (MIN)], β-lactams [ticarcillin/clavulanic acid (TIM2), ampicillin (AMP), AMP/sulbactam, piperacillin/tazobactam (PIP), aztreonam (ATM), ceftazidime (CAZ), ceftriaxone (CRO), cefazolin (CFZ), cefepime (FEP), doripenem (DOR), ertapenem (ETP), imipenem (IPM), meropenem (MEM)], and quinolones [levofloxacin (LEV), ciprofloxacin (CIP)]. Nitrofurantoin (NIT), which does not fall within these major classes, was categorized as “other.” The GN4F plate contained freeze-dried antibiotics at predefined concentration ranges (µg/ml). Gallibacterium anatis isolates cultured on blood agar were suspended in sterile distilled water and adjusted to a 0.5 McFarland turbidity standard, verified using a spectrophotometer. A 10 µl aliquot of this suspension was inoculated into Mueller–Hinton Broth (MHB) and homogenized using a vortex mixer. Subsequently, 50 µl of the MHB culture was dispensed into each well of the GN4F Sensititre™ plate using a multichannel pipette and incubated at 36°C for 24 hours. Escherichia coli ATCC 25922 was included as the quality control strain for AST. The MIC was defined as the lowest antibiotic concentration that inhibited visible bacterial growth, indicated by the absence of turbidity or pellet formation. MIC interpretations were categorized as susceptible (S), intermediate (I), or resistant (R) based on the Clinical and Laboratory Standards Institute (2018) (Additional File 2). Antibiotics included in the analysis were selected based on those approved or regulated for use in animals and/or humans. Molecular identification of genes encoding antibiotic resistanceMolecular detection of resistance-encoding genes was performed on isolates showing resistance status in MIC testing. Resistance-encoding genes were tested for three antibiotic families: TET (tet(B), tet(H), and tet(A)), β-lactam (blaSHV, ampC, and blaROB), and quinolone qnr(A) and gyr(A). The genomic DNA of G. anatis was extracted using the boiling method. The forward and reverse primers used in this study are listed in Additional File 2. The PCR reaction volume was 10 µl, comprising 5 µl of PCR master mix KAPA2G Fast Hotstart Readymix PCR Kit (KapaBiosystems, Cape Town, South Africa), 0.5 µl each of forward and reverse primer pairs with a concentration of 20 pmol, 1 µl template DNA, and 3 µl nuclease-free water. The amplification process began with pre-denaturation at 90°C for 3 minutes, followed by 35 cycles of amplification with denaturation at 94°C for 30 seconds, annealing according to the temperature specified in Additional File 2 for 40 seconds, and extension at 72°C for 1 minute. This was followed by a final extension at 72°C for 5 minutes. The PCR products were then electrophoresed on a 1.5% agarose gel stained with 10 mg/ml ethidium bromide and a 100 bp DNA ladder. Visualization bands were obtained using a UV transilluminator (Bio-Rad, USA) following electrophoresis. Statistical analysisThe multiple antibiotic resistance (MAR) index was calculated using the formula a/b, where a denotes the number of antibiotics to which the isolate exhibited resistance and b represents the total number of antibiotics tested (Catalano et al., 2022). Spearman’s correlation analysis was conducted to evaluate the relationship between phenotypic resistance patterns and the number of resistance genes detected. A correlation heatmap of antibiotic resistance was generated using SRPlot (SRPlot, Statistical data visualization platform. Retrieved from https://www.bioinformatics.com.cn/en (May 7, 2025). Ethical approvalEthical approval for this study was obtained from the Animal Ethics School of Veterinary Medicine and Biomedical Science, IPB University with number: 121/SKE/X/2023). ResultsIsolation and reconfirmation of isolated G. anatis isolate A total of 14 archival G. anatis isolates were cultured on blood agar. Macroscopic observation revealed small, non-mucoid, semi-transparent colonies with a hemolytic zone. Microscopically, the colonies were identified as Gram-negative with a pleomorphic bacilli morphology. These colonies were confirmed using the gyrB gene, which encodes both a structural component of G. anatis. All 14 isolates tested positive for the gyrB gene, showing a band at 561 bp (Fig. 1).
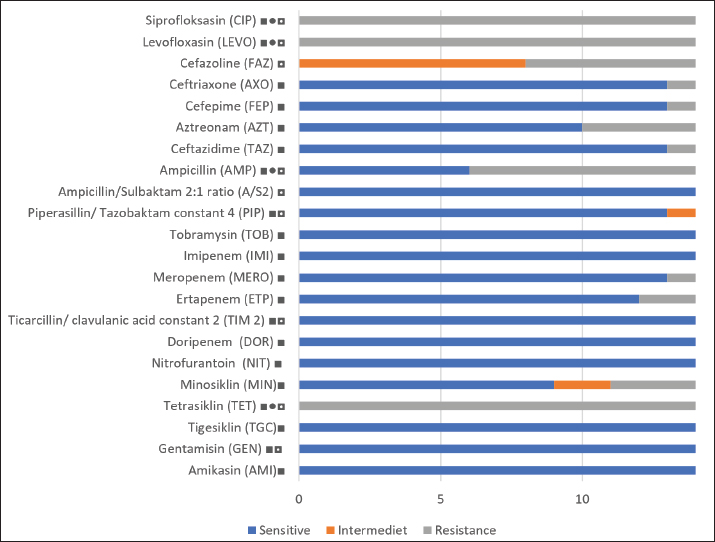
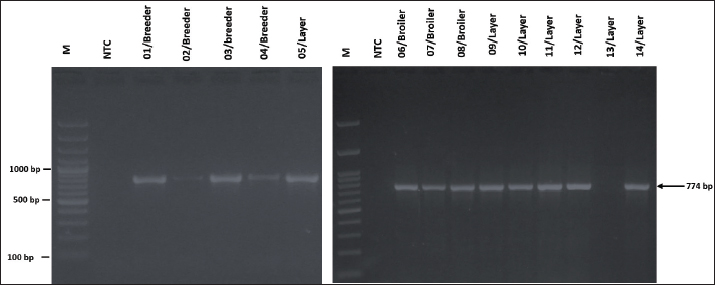
Fig. 1. Visualization of the PCR results for the gyrB gene in G. anatis isolates. Seven isolates were positive for the gyrB gene. The arrow (←) indicates the amplicon length at 561 bp. M: 100 bp marker. In vitro ASTOf the 23 samples collected, 14 isolates were confirmed as G. anatis by PCR. AST revealed that all isolates (100%) were resistant to TET, CIP, and levofloxacin. In contrast, no resistance was observed to AMI, GEN, tigecycline, IPM, TOB, or PIP (Fig. 2).
Fig. 2. Results of antibiotic resistance testing in G. anatis bacteria. ■ : antibiotics used in humans; ●: antibiotics used in poultry; ◘: antibiotics used in non-poultry animals base on Republic of Indonesia (2009, 2014). The antimicrobial susceptibility profiles of the G. anatis isolates showed diverse resistance patterns (Table 1). All isolates (n=14) were resistant to three antibiotics: TET (TET class) and CIP and levofloxacin (fluoroquinolone class). Eight antibiotics, namely TOB, IPM, ticarcillin/clavulanic acid constant 2 (TIM2), DOR, NIT, tigecycline, GEN, and AMI, showed no resistance in any isolate. Table 1. Antibiotic resistance patterns of G. anatis isolat.
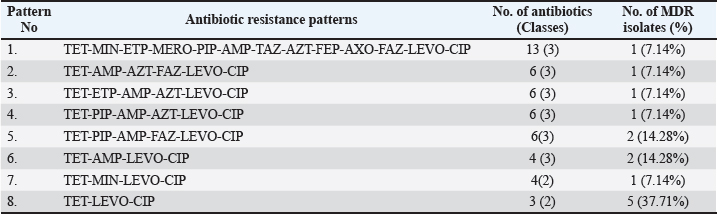
Among the antibiotics commonly used in poultry, PIP/tazobactam constant 4 (PIP) and TIM2 remained effective, as all isolates were susceptible. In contrast, several other poultry-associated antibiotics demonstrated high levels of resistance among G. anatis isolates. The resistance patterns of G. anatis isolates in this study also indicated isolates that exhibited resistance to more than two antibiotic classes, termed MDR. Eight antibiotic resistance patterns were identified, with the most common pattern being TET-LEVO-CIP, found in 37.71% of the samples. The resistance pattern containing the highest number of antibiotics was TET-MIN-ETP-MERO-PIP-AMP-TAZ-AZT-FEP-AXO-FAZ-LEVO-CIP, involving resistance to 13 antibiotics across 3 antibiotic groups (Table 2). Eight isolates (75.14%) exhibited MDR across three antibiotic classes: TGC, β-lactams, and quinolones (Table 3). No isolates met the criteria for XDR or PDR types. The occurrence of MDR among the antibiotic-resistant isolates displayed varying resistance patterns. Table 2. MDR of G. anatis isolates in antibiotic (n=14).
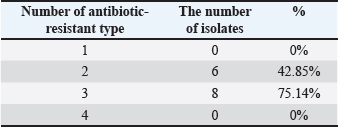
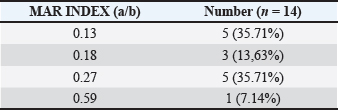
Table 3. MAR index of G. anatis isolates.
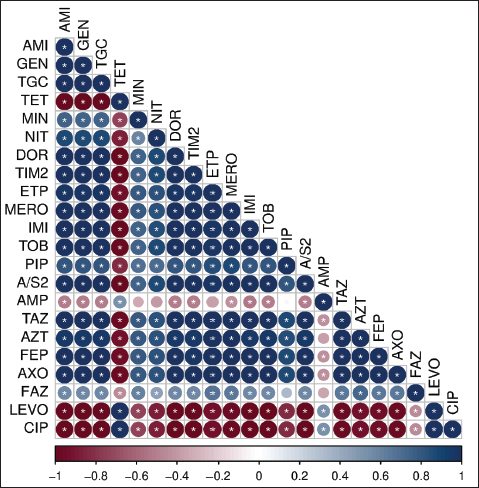
The MAR index values among the 14 G. anatis isolates ranged from 0.13 to 0.59 (Table 3). Notably, 6 of 14 isolates (42.8%) had MAR index values above 0.2, suggesting exposure to environments with high antibiotic pressure. Higher MAR values were associated with resistance to multiple antibiotic classes. Spearman’s correlation analysis using 22 types of antibiotics revealed several relationships, from strong positive to weak negative correlations. In general, most antibiotic pairs exhibited significant positive correlations, as indicated by dark blue shading and the presence of asterisks (*) on the heatmap (Fig. 3), with a significance level of p < 0.05. Aminoglycoside antibiotics, such as AMI and GEN, showed a high positive correlation, which is expected as they belong to the same antibiotic class—aminoglycosides. In contrast, AMI and GEN were strongly negatively correlated with TET, levofloxacin, and CIP, which belong to different antibiotic classes, namely TGC and quinolones.
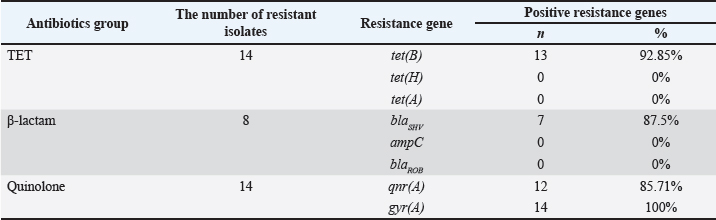
Fig. 3. Spearman’s correlation coefficient graphic between antibiotics. Color-coded values range from -1=negative correlation (red) to 1=positive correlation (blues), p significant level: 0.05; (*) are significant, without (*) are not significant. Conversely, several antibiotic pairs demonstrated low or statistically insignificant correlations (indicated by the absence of an asterisk), regardless of whether the relationship was positive or negative. In this research, weak or non-significant correlations were observed between PIP and AMP, ETP and AMP, MIN and AMP, PIP and cefazolin (FAZ), and FAZ and AMP. Antibiotic resistance-encoding genesThe resistance-encoding genes tet(B), tet(A), and tet(H) were detected in 14 G. anatis isolates resistant to TET antibiotics revealed that 13 out of 14 (93.33%) isolates tested positive for tet(B) with an amplicon size of 744 bp (Fig. 4), whereas none (0%) tested positive for the tet(A) and tet(H) encoding genes (Table 4).
Fig. 4. Visualization of the tet(B) resistance-encoding gene in G. anatis bacterial isolates using PCR. Thirteen isolates tested positive for tet(B). Arrow (←) indicates the amplicon length at 744 bp. M: 100 bp marker; NTC: non-template control. Table 4. Precentage entibiotic encoding resistance genes (n=14).
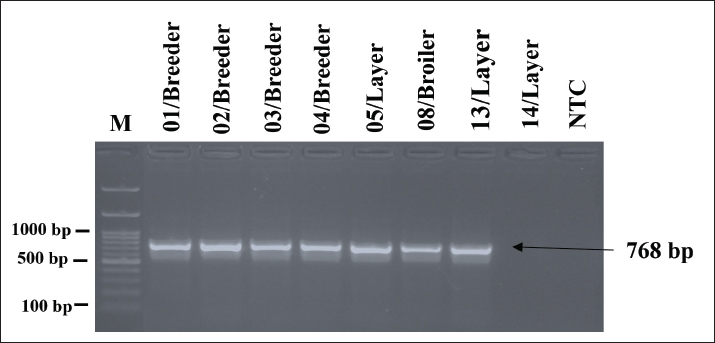
The results of detecting resistance-encoding genes blaSHV, ampC, and blaROB in eight G. anatis isolates resistant to β-lactam antibiotics revealed that seven out of eight (87.5%) isolates tested positive for blaSHV with an amplicon size of 768 bp (Fig. 5), while no isolates (0%) tested positive for the ampC and blaROB encoding genes. No isolates tested positive for the ampC or blaROB genes, indicating that blaSHV may play a primary role in mediating β-lactam resistance in these isolates.
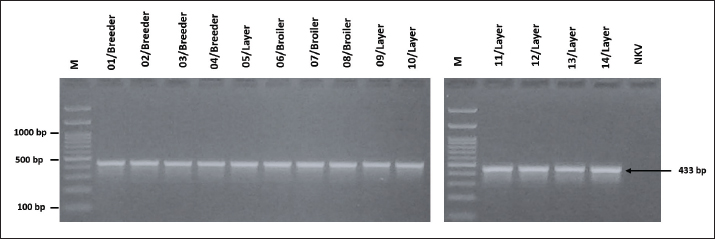
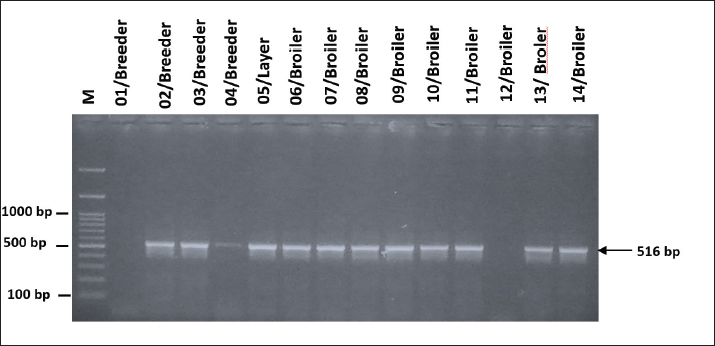
Fig. 5. Visualization of PCR results for the blaSHV resistance-encoding gene in G. anatis isolates. Seven isolates were positive for the blaSHV gene. The arrow (←) indicates the amplicon length at 768 bp. M: 100 bp marker. The detection of resistance-encoding genes gyr(A) and qnr(A) in 14 G. anatis isolates resistant to quinolone antibiotics revealed that all 14 isolates (100%) tested positive for gyr(A) with an amplicon size of 433 bp (Fig. 6), and a total of 12 out of 14 isolates (92.85%) tested positive for the qnr(A) encoding gene with an amplicon size of 516 bp (Fig. 7). The high prevalence of these genes reflects strong genetic support for the phenotypic resistance to quinolones observed in the isolates.
Fig. 6. Visualization of the gyr(A) resistance-encoding gene in G. anatis bacterial isolates using PCR. All isolates (n=14) were positive for gyr(A). Arrow (←) indicates the amplicon length of 433 bp. M: 100 bp marker; NTC: non-template control.
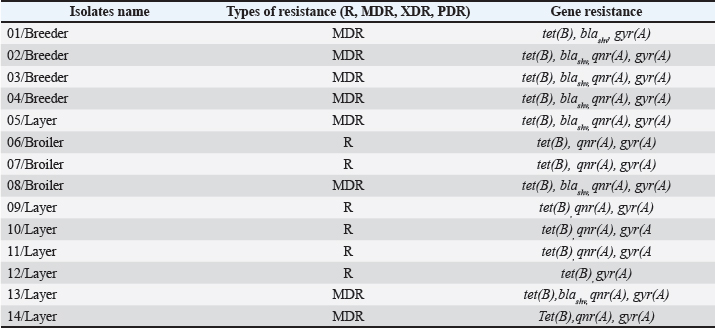
Fig. 7. Visualization of qnr(A) resistance-encoding gene in G. anatis bacterial isolates using PCR. Of the 14 isolates, 13 were positive for the qnr(A) gene. Arrow (←) indicates the amplicon length at 516 bp. M: 100 bp marker. A comparison between the phenotypic and genotypic profiles of AMR revealed a strong positive correlation (r=0.82), as shown in Table 5. All isolates derived from breeder chickens were classified as MDR and harbored the same four resistance genes: tet(B), blaSHV, qnr(A), and gyr(A), except for isolate 01/Breeder, which did not carry the qnr(A) gene. Among the layer chicken isolates, three were identified as MDR; however, isolate 14/Layer did not carry the blaSHV gene. In addition, four other isolates from layer chickens were categorized as resistant (R). Of these, three isolates (09–11/Layer) carried identical resistance genes—tet(B), qnr(A), and gyr(A)—while isolate 12/Layer lacked the qnr(A) gene. In broiler chickens, only isolate 08/Broiler met the criteria for MDR, possessing the tet(B), blaSHV, qnr(A), and gyr(A) genes. The remaining two broiler isolates (06/Broiler and 07/Broiler) were classified as resistant and carried tet (B), qnr (A), and gyr (A). These findings highlight the consistency between phenotypic resistance and the presence of specific resistance genes across different types of poultry. Table 5. Comparison of phenotypic and genotypic of AMR.
DiscussionThis study investigated the AMR profiles of G. anatis isolates and screened for resistance genes using PCR. The isolates originated from chickens experiencing decreased egg production and/or respiratory disorders. Although all archival isolates were successfully re-cultured on blood agar—characterized by hemolysis, small colony size, and a mucoid appearance—they consistently exhibited slow growth. Resistance testing against 22 antibiotics revealed that all isolates were TET-resistant, CIP, and levofloxacin. The high level of resistance to TET and fluoroquinolones is likely associated with their widespread use in poultry production (Shrestha et al., 2022). This finding aligns with Kursa et al. (2023)who reported 100% and 92% resistance to enrofloxacin and TET, respectively, among G. anatis isolates. MDR was primarily observed in TET, fluoroquinolone, and β-lactam antibiotic classes. MDR was more frequent in isolates from layer and breeder chickens, which generally have longer lifespans. This suggests that prolonged or repeated exposure to antibiotics—whether through direct administration or environmental contamination—may contribute to the emergence of MDR strains. The diversity of resistance patterns across isolates, as reflected in the MAR index values, further supports this observation. A MAR index greater than 0.2 indicates exposure to environments with intensive antibiotic usage, consistent with previous reports describing high MAR indices in G. anatis (Algammal et al., 2022). MDR in poultry pathogens represents a global concern, contributing to increased morbidity and mortality in humans, aquaculture, and livestock (Catalano et al., 2022). These findings highlight the need for stricter regulation and effective antimicrobial stewardship in poultry production systems. Aminoglycosides demonstrated the highest susceptibility levels, with no resistance detected in any isolate. This may be associated with the relatively limited use of aminoglycosides in Indonesian poultry production, as their application is restricted under national regulations (Government Regulation No. 18/2009; Regulation No. 41/2014). Resistance to antibiotics not designated for poultry use—such as MIN, ETP, MEM, CAZ, ATM, FEP, and CRO—was observed in 7%–28.75% of the isolates. This suggests possible off-label or inappropriate antibiotic use, or the circulation of resistant bacteria in the farm environment leading to cross-exposure. Genotypic screening identified tet(B) as a TET resistance gene in several isolates. This gene is commonly associated with TET resistance in Gram-negative bacteria, including G. anatis (Bojesen et al., 2011). No resistance genes were detected in isolate 13/Layer, indicating that other tet genes—of which more than 30 exist (Roberts, 2003)—may contribute to the resistance phenotype but were not included in the present PCR panel. Within the β-lactam class, blaSHV was the only resistance gene detected. This gene encodes extended-spectrum β-lactamase activity and is typically associated with resistance to penicillins and third-generation cephalosporins (Zahedi et al., 2018). These results correspond well with the observed phenotypic resistance patterns, particularly the high resistance to penicillin derivatives. In contrast, blaROB was not detected, which differs from the findings of Algammal et al. (2022) who reported a high prevalence of blaROB in G. anatis. Geographic variation and differences in farm microbiomes may account for this discrepancy. qnr(A) and gyr(A) were successfully identified in the fluoroquinolone group. Notably, two isolates (01/Breeder and 12/Layer) were negative for qnr(A), which differed from the findings of Zahedi et al. (2018) who found qnr(A) to be the major plasmid-mediated determinant of CIP resistance in E. coli. This suggests that resistance mechanisms in G. anatis may differ between bacterial species or involve alternative plasmid or chromosomal factors. Analysis of eight resistance genes revealed that each G. anatis isolate carried at least three resistance determinants, and six isolates possessed genes from three antibiotic classes, consistent with their phenotypic MDR profiles. The positive correlation between phenotypic and genotypic resistance supports the reliability of MIC-based AST as an indicator of underlying resistance mechanisms. Antibiotic resistance in G. anatis represents a significant threat to poultry health and productivity. The high levels of resistance—particularly to commonly used antibiotics—and the detection of resistance to antibiotics not authorized for poultry use emphasize the urgent need for appropriate antibiotic stewardship. Routine susceptibility testing and adherence to national regulations are essential to ensure effective treatment and reduce the emergence of MDR strains. ConclusionTo the best of the author’s knowledge, this is the first study to report the AMR status of G. anatis isolated from chickens in Indonesia. Antibiotic susceptibility testing using MIC revealed that MDR occurrences are highly common in field isolates of G. anatis. The occurrence of antibiotic resistance in this study also shows resistance to drugs that should not be used in poultry. This result is further supported by the high MAR index in these isolates. This finding serves as an alarm for Indonesia’s poultry farming industry and public health in general. The study also indicated resistance genes in the three antibiotic classes. AcknowledgmentsThe authors thank the United States Agency for International Development (USAID) through the SEAOHUN One Health Scholarship Program for funding this research and anonymous referees for their kind advice. Conflict of interestThe authors declare that there is no conflict of interest. FundingThe United States Agency for International Development (USAID) granted the publication of this journal through the SEAOHUN One Health Scholarship Program. The contents are the responsibility of the author(s) and do not necessarily reflect the views of USAID or the United States Government. Authors’ contributionsAlya Amaliah contributed to the conceptualization, sample collection, investigation, data analysis, and manuscript preparation. Agustin Indrawati and Ni Luh Putu Ika Mayasari contributed to the conceptualization, supervision, validation, data curation, and manuscript review and editing. Ryan Septa Kurnia contributed to the conceptualization, data curation, formal analysis, methodology, supervision, validation, manuscript review, and editing. Christian Marco Adi Nugroho contributed to the conceptualization, formal analysis, investigation, supervision, validation, review, and editing of the manuscript. Muhammad Ade Putra contributed to data curation, investigation, and manuscript review and editing. Data availabilityAll data were provided in the manuscript. ReferencesAhmed, S.K., Hussein, S., Qurbani, K., Ibrahim, R.H., Fareeq, A., Mahmood, K.A. and Mohamed, M.G. 2024. Antimicrobial resistance: impacts, challenges, and future prospects. J. Med. Surg. Public Health 2(2), 1000081. Algammal, A.M., Abo Hashem, M.E., Alfifi, K.J., Al-Otaibi, A.S., Alatawy, M., Eltarabili, R.M., Abd El-ghany, W.A., Hetta, H.F., Hamouda, A.M., Elewa, A.A. and Azab, M.M. 2022. Sequence analysis, antibiogram profile, virulence and antibiotic resistance genes of XDR and MDR Gallibacterium anatis isolated from layer chickens in Egypt. Infect. Drug Resist. 15, 4321–4334. Bojesen, A.M., Vazquez, M.E., Bager, R.J., Ifrah, D., Gonzalez, C. and Aarestrup, F.M. 2011. Antimicrobial susceptibility and tetracycline resistance determinant genotyping of Gallibacterium anatis. Vet. Microbiol. 148, 105–110. Catalano, A., Iacopetta, D., Ceramella, J., Scumaci, D., Giuzio, F., Saturnino, C., Aquaro, S., Rosano, C. and Sinicropi, M.S. 2022. Multidrug resistance (MDR): a widespread phenomenon in pharmacological therapies. Molecules 27(3), 616. Christensen, H., Bisgaard, M., Bojesen, A.M., Mutters, R. and Olsen, J.E. 2003. Genetic relationships among avian isolates classified as Pasteurella haemolytica, 'Actinobacillus salpingitidis' or Pasteurella anatis with proposal of Gallibacterium anatis gen. nov., comb. nov. and description of additional genomospecies within Gallibacterium gen. nov. Int. J. Syst. Evol. Microbiol. 53 (1), 275–287. Clinical and Laboratory Standards Institute. 2018. Performance standards for antimicrobial susceptibility testing, 31st ed.. Wayne, PA: CLSI. Collignon, P.J. and Mcewen, S.A. 2019. One Health—its importance in helping to better control antimicrobial resistance. Trop. Med. Infect. Dis. 4(1), 22. Duin, D.V. and Paterson, D.L. 2022. Multidrug resistant bacteria in the community: an update. Infect. Dis. Clin. North Am. 34(4), 709–722. Elbestawy, A.R., Ellakany, H.F., Abd El-hamid, H.S., Bekheet, A.A., Mataried, N.E., Nasr, S.M. and Amarin, N.M. 2008. Isolation, characterization, and antibiotic sensitivity assessment of Gallibacterium anatis biovar haemolytica, from diseased Egyptian chicken flocks during the years 2013 and 2015. Poult. Sci. 1519, 1519–1525. Hess, C., Grafl, B., Bagheri, S., Kaesbohrer, A., Zloch, A. and Hess, M. 2020. Antimicrobial resistance profiling of Gallibacterium anatis from layers reveals high number of multiresistant strains and substantial variability even between isolates from the same organ. Microbial. Drug. Resist. 26, 169–177. Karwańska, M., Wieliczko, A., Bojesen, A.M., Villumsen, K.R., Krzyżewska-Dudek, E. and Woźniak-Biel, A. 2023. Isolation and characterization of multidrug resistant Gallibacterium anatis biovar haemolytica strains from Polish geese and hens. Vet. Res. 54, 67. Kursa, O., Tomczyk, G., Sieczkowska, A. and Sawicka-Durkalec, A. 2023. Prevalence, identification and antibiotic resistance of Gallibacterium anatis isolates from chickens in Poland. Pathogens 12(8), 992. Liang, X., Zhang, Z., Wang, H., Lu, X., Li, W., Lu, H., Roy, A., Shen, X., Irwin, D.M. and Shen, Y. 2023. Early-life prophylactic antibiotic treatment disturbs the stability of the gut microbiota and increases susceptibility to H9N2 AIV in chicks. Microbiome 11, 163. Magiorakos, A.P., Srinivasan, A., Carey, R.B., Carmeli, Y., Falagas, M.E., Giske, C.G., Harbarth, S., Hindler, J.F., Kahlmeter, G., Olsson-Liljequist, B., Paterson, D.L., Rice, L.B., Stelling, J., Struelens, M.J., Vatopoulos, A., Weber, J.T. and Monnet, D.L. 2012. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281. Republic of Indonesia. 2009. Law Number 18 of 2009 concerning Livestock and Animal Health. State Gazette of the Republic of Indonesia No. 84 of 2009. Republic of Indonesia. 2014. Government regulation number 41 of 2014 concerning amendments to government regulation number 47 of 2011 on cultivation area management. State Gazette of the Republic of Indonesia No. 123 of 2014. Roberts, M. 2003. Acquired tetracycline and/or macrolide-lincosamides-streptogramin resistance in anaerobes. Anaerobe 9, 63–69. Rzewuska, M.E.K., Szeleszczuk, P. and Binek, M. 2007. Isolation of Gallibacterium spp. from peacocks with respiratory tract infections. Medycyna. Weterynaryjna. 63, 1431–1433. Sanchez-Alonso, P., Cobos-Justo, E., Avalos-Rangel, M.A., López-Reyes, L., Paniagua-Contreras, G.L., Vaca-Paniagua, F., Anastacio-Marcelino,E., López-Ochoa, A.J., Pérez Marquez, V.M., Negrete-Abascal, E, Vázquez-Cruz, C. 2023. A Maverick-like cluster in the genome of a pathogenic, moderately virulent strain of Gallibacterium anatis, ESV200, a transient biofilm producer. Front. Microbiol. 14, 1084766. Shrestha, K., Acharya, K.P. and Shrestha, S. 2018. One Health: the interface between veterinary and human health. Int. J. One Health 4 (2), 8–14. Shrestha, N., Layalu, S., Amatya, S., Shrestha, S., Basnet, S., Shrestha, M. and Shrestha, U.T. 2022. Higher level of quinolone residues in poultry meat and eggs: an alarming public health issue in Nepal. medRxiv [Preprint]. doi:10.1101/2022.09.03.22279573 Tang, B., Pors, S.E., Kristensen, B.M., Skjerning, R.B.J., Olsen, R.H. and Bojesen, A.M. 2020. GtxA is a virulence factor that promotes a Th2-like response during Gallibacterium anatis infection in laying hens. Vet. Res. 51(1), 1–12. Van Driessche, L., Vanneste, K., Bogaerts, B., De Keersmaecker, S.C.J., Roosens, N.H., Haesebrouck, F., De Cremer, L., Deprez, P., Pardon, B. and Boyen, F. 2020. Isolation of drug-resistant Gallibacterium anatis from calves with unresponsive Bronchopneumonia, Belgium. Emerg. Infect. Dis. 26, 721–730. Yaman, S. and Sahan Yapicier, O. 2019. Diagnosis of Gallibacterium anatis in layers: first report in Turkey. Braz. J. Poultry Sci. 21(03), 001–008. Zahedi, Z., Hakemi, V.M. and Bagheri, B.F. 2018. Contribution of gyrA and qnrA genes in ciprofloxacin resistant Escherichia coli isolates from patients with urinary tract infections of Imam Khomeini Hospital of Tehran. Arch. Pediatr. Infect. Dis. 6(4), e62129. Additional File 1. List of samples in the studies.
Additional File 2. List of primers used in this study for PCR amplification.
| ||
| How to Cite this Article |
| Pubmed Style Amaliah A, Mayasari NLPI, Kurnia RS, Nugroho CMH, Putra MA, Indrawati A. Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Vet. J.. 2026; 16(1): 458-468. doi:10.5455/OVJ.2026.v16.i1.43 Web Style Amaliah A, Mayasari NLPI, Kurnia RS, Nugroho CMH, Putra MA, Indrawati A. Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. https://www.openveterinaryjournal.com/?mno=275180 [Access: January 31, 2026]. doi:10.5455/OVJ.2026.v16.i1.43 AMA (American Medical Association) Style Amaliah A, Mayasari NLPI, Kurnia RS, Nugroho CMH, Putra MA, Indrawati A. Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Vet. J.. 2026; 16(1): 458-468. doi:10.5455/OVJ.2026.v16.i1.43 Vancouver/ICMJE Style Amaliah A, Mayasari NLPI, Kurnia RS, Nugroho CMH, Putra MA, Indrawati A. Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Vet. J.. (2026), [cited January 31, 2026]; 16(1): 458-468. doi:10.5455/OVJ.2026.v16.i1.43 Harvard Style Amaliah, A., Mayasari, . N. L. P. I., Kurnia, . R. S., Nugroho, . C. M. H., Putra, . M. A. & Indrawati, . A. (2026) Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Vet. J., 16 (1), 458-468. doi:10.5455/OVJ.2026.v16.i1.43 Turabian Style Amaliah, Alya, Ni Luh Putu Ika Mayasari, Ryan Septa Kurnia, Christian Marco Hadi Nugroho, Muhammad Ade Putra, and Agustin Indrawati. 2026. Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Veterinary Journal, 16 (1), 458-468. doi:10.5455/OVJ.2026.v16.i1.43 Chicago Style Amaliah, Alya, Ni Luh Putu Ika Mayasari, Ryan Septa Kurnia, Christian Marco Hadi Nugroho, Muhammad Ade Putra, and Agustin Indrawati. "Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia." Open Veterinary Journal 16 (2026), 458-468. doi:10.5455/OVJ.2026.v16.i1.43 MLA (The Modern Language Association) Style Amaliah, Alya, Ni Luh Putu Ika Mayasari, Ryan Septa Kurnia, Christian Marco Hadi Nugroho, Muhammad Ade Putra, and Agustin Indrawati. "Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia." Open Veterinary Journal 16.1 (2026), 458-468. Print. doi:10.5455/OVJ.2026.v16.i1.43 APA (American Psychological Association) Style Amaliah, A., Mayasari, . N. L. P. I., Kurnia, . R. S., Nugroho, . C. M. H., Putra, . M. A. & Indrawati, . A. (2026) Antimicrobial resistance patterns and genetic determinants of emerging Gallibacterium anatis isolates from poultry in West Java, Indonesia. Open Veterinary Journal, 16 (1), 458-468. doi:10.5455/OVJ.2026.v16.i1.43 |