Open Veterinary Journal, (2026), Vol. 16(1): 105-117
Research Article
10.5455/OVJ.2026.v16.i1.10
Chitosan nanoparticles as a sustainable antibacterial approach
for inhibiting Escherichia coli and Salmonella spp. in mastitic dairy cattle and buffaloes
Mahmoud Abd El-Mongy1, Manar M. Farouk2*, Amani Abd El Latif Mosleh3 and Shaimaa Moawad Nada3
1Department of Microbial Biotechnology, Faculty of Genetic Engineering and Biotechnology Research, University of Sadat City, Sadat City, Egypt
2Department of Internal Medicine and Infectious Diseases, Faculty of Veterinary Medicine, Cairo University, Giza, Egypt
3Animal Health Research Institute, Shibin El-Kom Branch, ARC, El-Menufia, Egypt
*Corresponding Author: Manar M. Farouk. Department of Internal Medicine and Infectious Diseases, Faculty of Veterinary Medicine, Cairo University, Giza, Egypt. Email: manarmagdy [at] cu.edu.eg
Submitted: 29/09/2025 Revised: 30/11/2025 Accepted: 10/12/2025 Published: 31/01/2026
© 2025 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Background: Mastitis poses a significant economic impact in the dairy sector and serves as a potential source of infection for healthy cows, impeding attempts to become self-sufficient in milk production. Treatment failure due to the development of multidrug-resistant pathogens raises concern about the need to investigate antimicrobial alternatives. Nanoparticles are attracting attention as a therapeutic approach for various infectious disorders. Despite the importance of this issue, limited research has been conducted on resistant bacteria that cause mastitis.
Aim: This study aimed to investigate the occurrence and antimicrobial susceptibility profile of Escherichia coli and Salmonella spp. as environmental pathogens causing mastitis and the antibacterial activity of chitosan nanoparticles (ChNPs) against multidrug-resistant pathogens.
Methods: A total of 100 milk samples were collected from dairy cattle and buffaloes with clinical and subclinical mastitis using a convenience sampling approach, with no randomization. As such, the sample may not fully represent all dairy sources in the study area. Mastitis was initially detected using the California mastitis test. Following standard biochemical tests and bacteriological culture, suspected positive samples were molecularly identified using polymerase chain reaction and phylogenetic analysis of 16S rRNA gene sequences. Along with antimicrobial susceptibility testing. The qualitative antimicrobial activity of ChNPs was assessed using the disc diffusion method along with quantitative determination methods of minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC). Additionally, the effect of ChNPs on the bacterial cells was evaluated through TEM imaging.
Results: Results revealed that E. coli was recovered with percentages of 56%, 32%, 22%, and 24%, and Salmonella spp. was recovered with percentages of 10.5%, 8.1%, 2.1%, and 0% from clinical and subclinical cases of cattle and buffaloes, respectively. The tested E. coli and Salmonella spp. isolates displayed variable sensitivity and resistance patterns toward the evaluated antimicrobials. The ChNPs exhibited potent MIC and MBC effects at 2.5 and 10 mg/ml, respectively, against both multidrug-resistant E. coli and Salmonella spp. TEM imaging revealed reduced bacterial cell viability by disrupting the membrane integrity.
Conclusion: The outcomes of this study collectively provide a crucial starting point for developing a novel treatment plan for bovine mastitis.
Keywords: Antimicrobial resistance, Bovine mastitis, E. coli, Nanoparticles, Salmonella.
Introduction
Mastitis seems to be a big concern for dairy production, and measures have been researched and recommended for improving dairy cattle well-being (Dingwell et al., 2003; Crispie et al., 2004). It causes financial losses such as a decline in milk yield, an extraordinary rate of culling, veterinary interventions, and infrequent mortality in challenging or non-treated severe cases (Aghamohammadi et al., 2018).
Intramammary infections may result from two major groups of environmental and contagious pathogens (Chehabi et al., 2019). Coliforms are Gram-negative bacteria that have been identified as a major cause of bovine environmental mastitis (Batool, et al., 2012). Furthermore, mastitis is considered one of the major causes of antibiotic use in veterinary practices (Osman et al., 2014). Owing to the upward concern for antimicrobial resistance, national and international protocols have been executed to minimize unnecessary use of antibiotics (Barlam et al., 2016).
Nanotechnology is a highly developed technology that plays an eminent role in various applicable therapeutic fields (Youssef et al., 2019). Recently, nanoparticles have been used as an antimicrobial alternative and a therapeutic approach for controlling bacterial illnesses caused by multidrug-resistant pathogens in animals (Farouk et al., 2015).
Because of their unique characteristics, chitosan nanoparticles (ChNPs) displayed a strong antimicrobial effect against both Gram-positive and Gram-negative microorganisms (Abdel-Razek 2019). The rapid advancement of sustainable chemicals has increased the appeal of ChNPs as an antibacterial agent (Abdel-Ghany and Salem 2020).
Nanotechnology, particularly ChNPs, offers a promising alternative. ChNPs are biodegradable, nontoxic, and positively charged, enabling electrostatic interactions with negatively charged bacterial membranes to disrupt integrity and kill cells. ChNPs show strong antimicrobial and antibiofilm activity against Staphylococcus aureus strains isolated from bovine mastitis milk (Godoy et al., 2025). Moreover, ChNPs outperform native chitosan against mastitis pathogens while maintaining mammalian cell viability (Orellano et al., 2019). In addition, antibiotic-loaded ChNPs (e.g., cloxacillin) can further reduce minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values against biofilm-producing S. aureus, suggesting enhanced therapeutic efficacy (Eskandari et al., 2025). ChNPs have also been reported to inhibit Pseudomonas species from mastitic milk and disrupt their biofilms (Rivera Aguayo et al., 2020). Importantly, in vitro safety assessments indicate that ChNP formulations show minimal cytotoxicity to bovine cells, supporting their potential for use in intramammary applications (Castro-Valenzuela et al., 2025).
The current investigation aimed to determine the prevalence of Escherichia coli and Salmonella spp. as causes of environmental mastitis in both cattle and buffaloes, characterize the antibiogram profile of the recovered isolates, and evaluate the possible antibacterial effectiveness of commercially available synthetically manufactured ChNPs against strains that have been found to be resistant to multiple antimicrobials. The findings of this investigation may pave the way to identify an alternative approach to control environmental mastitis infections in dairy farms and establish an innovative strategy to reduce antibiotic resistance in animal bacterial diseases.
Materials and Methods
Study design
A pilot study targeting 100 buffaloes and cows admitted to governmental veterinary units in El-Menoufia Governorate, Egypt, was conducted between February and July 2024. Among the inclusion criteria were any milking buffaloes or cows that could have clinical or subclinical mastitis admitted to veterinary units. The cases were included after they had been confirmed to have mastitis using the California Mastitis Test (CMT) according to the method of Kandeel et al. (2018). Buffaloes and cows with negative CMT results were excluded from the study.
Sampling
The sampling procedure was conducted from February 2024 to July 2024. A total of 100 milk samples were collected, including 50 (25 for each cattle and buffalo species) from clinical mastitis cases and 50 (25 for each cattle and buffalo species) from subclinical mastitis cases. Following the pre-milking teat dip, the teat ends were cleansed, and each quarter’s first two to three milk strips were discarded. The CMT was then tested using a five-point rating system (CMT: Negative, the mixture remained liquid, with no evidence of precipitation; Trace, the mixture had slight precipitation; 1, a clear precipitate formed with no tendency for gel formation; 2, the mixture thickened with some gel formed; 3, the mixture thickened with complete gel formation and stuck to the bottom of the paddle) according to Kandeel et al. (2018). Samples with a CMT score of 3 and the presence of abnormal milk (e.g., clots, watery secretion) or udder changes (swelling, heat, pain) were classified as CMT (Schalm and Noorlander, 1957; Ruegg 2002; Dohoo et al., 2009).
Positive CMT cases were enrolled, and negative cases were excluded. After the enrollment process, 1 ml of milk was collected aseptically from clinically affected quarters in sterile containers, and the animal’s identifying number, date, and time of collection were each prominently recorded. The samples were promptly transported to the laboratory in a cool ice box for further bacteriological analysis within 24 hours.
Isolation and identification of E. coli and Salmonella species
After receiving the milk samples, all cups were gently mixed, and approximately 1 ml from each milk sample was inoculated into brain heart infusion (BHI) broth (Oxoid, UK) and incubated at 37°C for 24 hours to promote E. coli propagation. The inoculated samples were plated on Eosine Methylene Blue (EMB) agar medium (Oxoid, UK) and incubated for 24 hours at 37°C. The presumptive 2–3 mm circular, convex, metallic green colonies were purified by subculturing on EMB agar plates and biochemically identified based on the indole, methyl red, Voges-Proskauer, and citrate utilization tests, and triple sugar iron agar media (Oxoid, UK) (Cheesbrough 2006 and Lebofe and Pierce 2012). Salmonella spp. isolation was performed following ISO-6579 standard (ISO-6579) and biochemically identified according to Quin et al. (2002). After routine biochemical identification, presumptive E. coli and Salmonella spp. isolates were cultivated in BHI broth and kept at 37°C overnight. Subsequently, the cultured broths were analyzed.
Serological identification of E. coli and Salmonella species
The E. coli isolates were serologically identified according to Kok et al. (1996) using rapid diagnostic E. coli antisera sets (DENKA SEIKEN Co., Japan) for the diagnosis of enteropathogenic types. The Salmonella species isolates were serotyped following the Kauffman-White scheme (Grimont and Weill, 2007) using a rapid diagnostic Salmonella antisera kit (Welcome Diagnostic, a Division of the Wellcome Foundation Limited, Dartford, England DA15 AH) at the Animal Health Institute, Dokki, Giza.
Molecular identification and phylogenetic analysis of recovered E. coli and Salmonella isolates
Deoxyribonucleic acid (DNA) extraction from recovered isolates
Genomic DNA was extracted from overnight cultures on tryptone soy agar (Oxoid®, USA) (QIAamp DNA Mini Kit, Cat. No. 51304, QIAGEN GmbH, Hilden, Germany) according to the manufacturer’s instructions. The bacterial DNA was used as a DNA template for the polymerase chain reaction (PCR).
Molecular identification
The phoA gene is one of the highly conserved genes selected for the molecular identification of E. coli isolates using PCR. This gene encodes alkaline phosphatase, an enzyme in E. coli that removes phosphate groups from molecules, particularly under phosphate-limited conditions. It is specific enough to help distinguish E. coli from many other Enterobacteriaceae.
Meanwhile, conventional PCR was conducted to identify the Salmonella species isolates using the invA gene. (invasion-associated gene) as a highly conserved marker for genus-level confirmation.
A 25 μl reaction mixture, which included 5 μl of DNA template and 5 μl of 5 × TaqMaster mix (Jena Bioscience, Germany), 1 μl of each forward and reverse primer (10 pmol/μl), with the volume adjusted to 25 μl using PCR-grade water (Jena Bioscience, Germany), was used for PCR. Electrophoresis was performed using 1.5% (wt/vol) agarose stained with 0.5 μg/ml ethidium bromide, after which the PCR amplicons were visualized under a UV illumination system. Gel images were captured using a GelDoc 1000 fluorescent imaging system (Bio-Rad) and analyzed using Gel-Pro Analyzer® version 4 (Media Cybernetics, Silver Spring, MD, USA). The PCR thermal profiles, primer set sequences, and amplicon size for each invA and phoA gene are consistent with those of Hu et al. (2011) and Oliveira et al. (2003).
Phylogenetic analysis
Salmonella spp. and E. coli isolates were detected using 16S rRNA gene universal primers: forward (5-AGAGTTTGATCMTGGCTCAG-3) and reverse (5-TACGGYTACCTTGTTACGACTT-3) (Metabion, Germany). The reactions were performed in a T3 Thermal cycler (Biometra, Germany) to amplify the 16S rRNA gene in accordance with Lagacé et al. (2004). Images of the amplified DNA were also captured using agar gel electrophoresis. The unassembled raw sequences of our investigation were combined and compared to the other sequences in GenBank using BLASTN. Nucleotide sequences deposited in GenBank were identified using the retrieved accession numbers. A phylogenetic tree was generated using the neighbor-joining method in MEGA X.11 software to determine the genetic distance. At 1,000 repetitions, the confidence level for each branch was evaluated using bootstrap analysis (Felsenstein, 1985).
Antibiogram analysis and multiple antimicrobial resistance index estimation
The susceptibility of 10 out of 39 randomly selected E. coli isolates from both clinical and subclinical cases and all 4 recovered Salmonella isolates to the following antibiotics was inspected using the Kirby-Bauer disc diffusion method: ofloxacin (OFX, 5 µg), clindamycin (DA, 2 µg), ceftriaxone (CRO, 30 µg), ampicillin/sulbactam (SAM, 20 µg), and tetracycline (TE, 20 µg). The findings were assessed in accordance with the breakpoint guidelines of the Clinical and Laboratory Standards Institute (CLSI 2020) because that was in force when the experiments were performed.
The MAR index of each tested isolate was evaluated using the formula of Paul et al. (1997): MAR index=(number of antibiotics to which the isolate is resistant) / (total number of antibiotics tested). In accordance with Christopher et al. (2013), isolates were categorized using the MAR index results as follows: isolates are deemed to have narrow drug resistance (NDR) if their index is less than 0.3; multi drug resistant (MDR) is considered as those with an index ≥ 0.3–0.7; XDR isolates are indicated by an index of 0.8–0.9; and isolates identified as pan drug-resistant exhibited resistance to all antibiotics tested.
Antibacterial efficacy of chitosan against MDR isolates of E. coli and Salmonella
Characterization of chitosan nanoparticles
Chemically synthesized CNPs were purchased from the Nanotech Center, 6th October, Giza, Egypt. The nanoparticle powder was dissolved (1000 μg/ml) in deionized water to create a nanoparticle suspension. Some characteristics of ChNPs were determined. Briefly, the particle size distribution was determined using dynamic light scattering (DLS) or a Zetasizer instrument (Malvern). The elemental contents of the chitosan samples were evaluated using energy dispersive X-ray spectroscopy (JED-2300 analysis station, Joel) and X-ray diffraction (XRD) techniques.
Minimum inhibitory concentration and minimum bactericidal concentration estimation
The synthesized sonicated chitosan nanoparticle solution was prepared by mixing it in sterile normal saline and stirring until a consistent colloidal suspension (CS) with a final concentration of 1,000 μg/ml was obtained. The resulting solution was stored at 4°C until needed. The MIC of the synthesized sonicated CS group with NP against E. coli and Salmonella spp. isolates was determined using the tube dilution method. This was achieved by preparing various dilutions (10, 5, 2.5, 1.25, and 0.6 mg/ml) from the stock nanoparticles. Then, 50 µl of the overnight incubated E.coli and Salmonella spp. bacterial suspensions were incubated overnight at 37°C with growth adjusted to 0.5 McFarland (equivalent to 1.5 × 108 CFU) and added to each individual tube in two distinct groups. After that, for 24 hours, the tubes were incubated at 37°C. The MIC is the lowest concentration of chitosan nanoparticles that terminated visible bacterial growth after 24 hours of incubation. Meanwhile, the MBC was determined by plating 50 μl from the tested tubes that lacked growth or cloudiness on Müller-Hinton agar and incubating at 37°C for 24 hours. The plates were examined for macroscopic bacterial growth, and the presence or absence of growth was determined. The MBC endpoint occurs when the lowest concentration of a nanoparticle agent results in the death of 99.9% of the bacterial population (Parvekar et al., 2020).
Evaluation of the antimicrobial action of CNPs against E. coli and Salmonella using transmission electron microscopy
Two bacterial isolates were subcultured with 20 μg/ml of CNPs in separate Falcon tubes containing tryptic soy broth. Twenty-four hours of incubation at 30°C was followed by centrifugation (3,000 rpm) for 15 minutes to remove the supernatants. The precipitate was collected for examination via TEM (JEOL, USA) at the TEM unit, Assuit University, Egypt, according to Abdel-Razek (2019).
Statistical analysis
Statistical analyses were performed using the Statistical Package for the Social Sciences (version 18.0). The chi-square test was used to compare categorical variables (Pearson, 1900). For quantitative measurements, data were first assessed for normality using the Shapiro–Wilk test (Shapiro and Wilk, 1965). Normally distributed data were analyzed using one-way analysis of variance (Fisher, 1925), whereas non-normal data were analyzed using the Kruskal–Wallis test (Kruskal and Wallis, 1952). At a probability value of less than 0.05 (p ˂ 0.05), the significance level was established. The conducted analyses were based on the clinical or subclinical status of the investigated cases for each separate bacterial species.
Ethical approval
The Institutional Animal Care and Use Committee of the Cairo University Faculty of Veterinary Medicine (VETCU-IACUC) approved all procedures executed during this investigation (VET CU 110520251134). All procedures were in accordance with the ARRIVE standards and closely obeyed the rules and regulations Approval Number: Vet CU110520251134. Approval Date: 11/05/2025. Date of Expiration: 10/05/2026.
Results
Prevalence of E. coli and Salmonella spp. in cattle and buffaloes
Regarding E. coli isolation, a total of 39 isolates were biochemically identified and molecularly confirmed using a PCR assay targeting E. coli phoA-specific primer. Of the 39 isolates, 14 were from clinical cattle, 8 from clinical buffaloes, 11 from subclinical cattle, and 6 from subclinical buffaloes.
The overall E. coli prevalence was 44% for clinical and 34% for subclinical cases (Table 1). Ten randomly selected out of 39 recovered E. coli isolates were serotyped, comprising 3, 2, 2, and 3 isolates from cattle and buffaloes suffering from clinical and subclinical mastitis, respectively. The identified serotypes were O111:K58, O164:K-, and O114:K90 from clinically affected cattle and O118:K- and O164:K- from subclinically affected cattle. Meanwhile, the O118:K- and O44:K80 serotypes were obtained from clinically affected buffaloes, and the O78:K80, O44:K80, and O114:K90 serotypes were obtained from subclinical mastitis cases of buffaloes.
Table 1. Prevalence of E-coli species recovered from milk samples of cattle and buffaloes with clinical and subclinical mastitis.
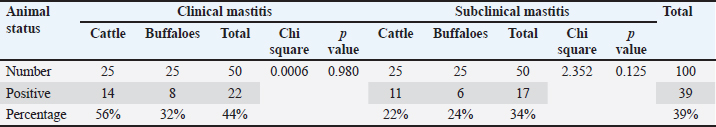
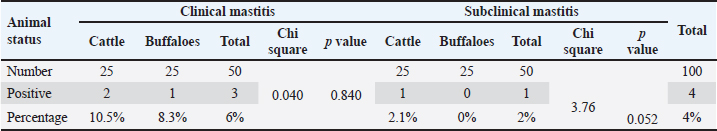
A total of 4 Salmonella spp. isolates with an overall prevalence of 4% (4/100) were detected and identified via routine bacteriological isolation and biochemical identification. Positive Salmonella colonies appeared pink with a black center and a highly transparent reddish zone on xylose lysine deoxycholate media. The prevalence of Salmonella spp. among the investigated animals was 10.5%, 8.3%, 2.1%, and 0% in the clinical and subclinical cases of cattle and buffaloes, respectively (Table 2). Serotyping of the 4 strains revealed that 3 out of the recovered serovars were Salmonella typhimurium and 1 Salmonella Entertidis strain recovered from cattle with subclinical mastitis. The other three recovered S. typhimurium strains, comprising 1 recovered from buffalo with clinical mastitis and 2 from cattle with clinical mastitis. The statistical analyses revealed no significant difference (p > 0.05) among the investigated clinical or subclinical cases of both cattle and buffaloes in the detection of E. coli and Salmonella spp., as shown in Tables 1 and 2.
Table 2. Prevalence of Salmonella spp recovered from milk samples of cattle and buffaloes with clinical and subclinical mastitis.
Phylogenetic analysis of 16S rRNA sequence
The partial codon sequences of the amplified fragments of 16S rRNA of amplicon size 1485 bp for each selected E. coli and S. Enteritidis isolate were deposited in GenBank. The accession numbers of the deposited sequences were PP425900 and PP430329 for E. coli and S. Enteritidis, respectively. The maximum likelihood analysis of the 16S rRNA gene revealed the genetic relatedness of the E. coli sequence (PP425900) to other E. coli strains retrieved from GenBank (Fig. 1). Furthermore, the analysis revealed that the PP430329 sequence shared the same clade with other S. Enteritidis strains retrieved from GenBank (Fig. 2).
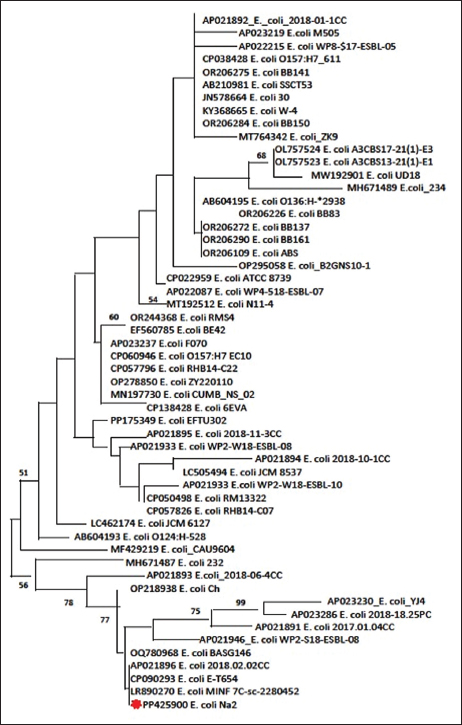
Fig. 1. Phylogenetic tree of Escherichia coli.
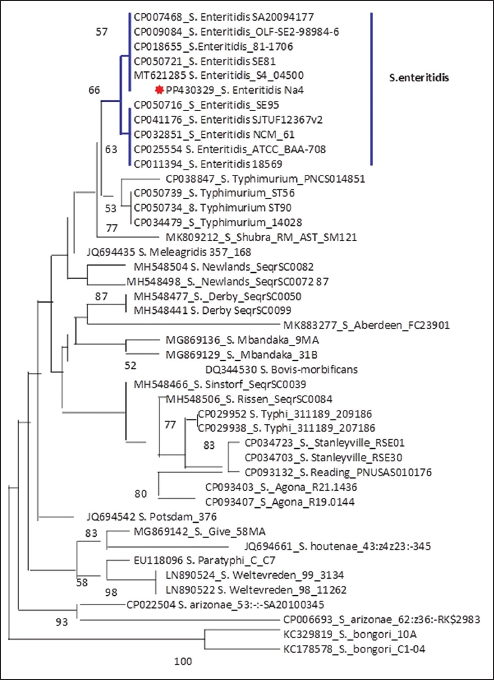

Fig. 2. Phylogenetic tree of Salmonella.
Antibiogram and the MAR index
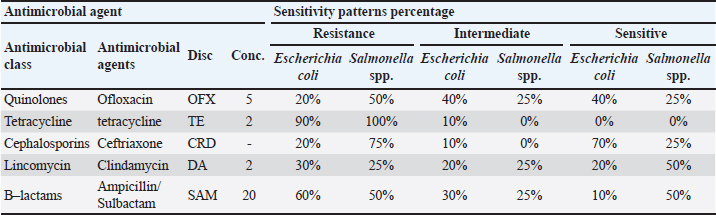
The tested E. coli and Salmonella spp. isolates displayed variable sensitivity and resistance patterns toward the evaluated antimicrobials (Table 3). Furthermore, all 4 tested Salmonella spp isolates were identified as MDR Salmonella spp, with an index range 0.4–0.8. The MDR and NDR were exhibited by 7 and 3 of the tested E. coli isolates, respectively. The isolates with a MAR index of less than 0.3 MAR index were identified as NDR, and those of ≥ 0.3–0.7 are considered MDR.
Table 3. Antimicrobial susceptibility patterns of Escherichia coli and Salmonella spp. recovered from cattle and buffaloes.
Characteristics of ChNPs
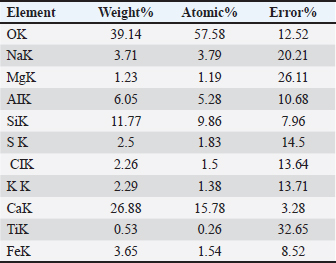
DLS showed that the average size distribution range of ChNPs was 20.1–75.2 nm. The results of the elemental EDAX analysis are displayed in Table 4 and Figure 3. Meanwhile, the XRD analysis showed a stronger peak at a maximum reflection at 13.1 (32.2 count/s) (Fig. 4).
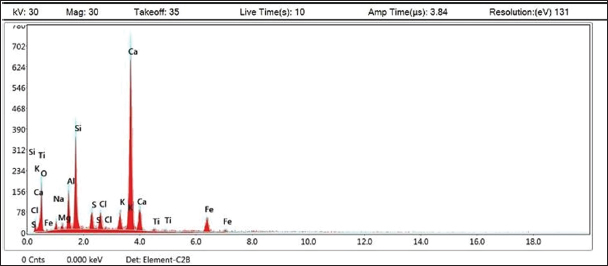
Fig. 3. EDAX analysis of chitosan nanoparticles.
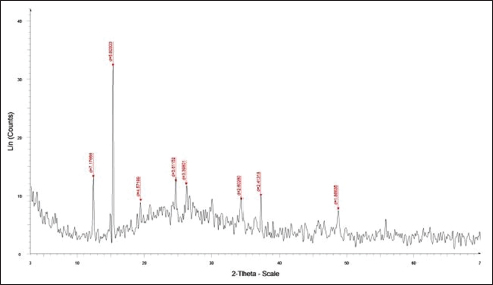
Fig. 4. X-ray diffraction pattern (XDR) of chitosan nanoparticles.
Table 4. Elementary EDAX analysis for chitosan nanoparticles.
In vitro antibacterial activity of ChNPs
The antimicrobial properties of ChNPs against various E. coli and Salmonella spp. isolates were evaluated using the agar well diffusion method along with their MIC and MBC determinations. ChNPs exhibited a potent inhibitory effect, resulting in a growth inhibition zone that extended from 8 to 10 mm. The MIC and MBC for ChNPs were the same for the tested Salmonella spp. and E. coli isolates, whereas the MIC and MBC were 2.5 and 10 mg/ml, respectively.
Antibacterial activity of ChNPs via TEM
ChNP-treated bacterial culture revealed ChNPs in the cell cytoplasm. There were empty areas in the cytoplasm of particular bacterial cells. (Fig. 5A). The TEM images of bacteria cultivated without ChNPs appeared to be in their usual shape (Fig. 5B).
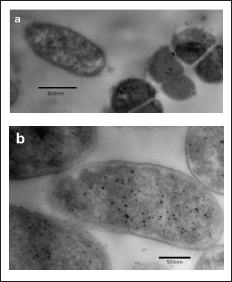
Fig. 5. Treated and untreated Escherichia coli under an electron microscope.
Discussion
Mastitis has a serious economic impact on the dairy industry, resulting in adverse effects on milk quality and animal health (Patil et al., 2021). Although it is not a commonly fatal disease, it has serious financial consequences for both the dairy industry and farmers. Additionally, it is highly prevalent in cattle and buffaloes with almost a 25% drop in their milk production, and it is of public health importance due to its associated hazard of pathogenic bacteria transmission and its heat-stable toxins (Ashraf and Imran, 2020).
In the current investigation, E. coli was found in only 14 (56%), 8 (32%), 11 (22%), and 6 (24%) clinical and subclinical mastitis cases of cattle and buffaloes, respectively. Furthermore, 10.5%, 8.1%, 2.1%, and 0% of Salmonella spp. were recovered from clinical and subclinical cases of cattle and buffaloes, respectively. There was no significant increase in the prevalence of Salmonella spp. in subclinical affected buffaloes compared with subclinical cattle cases. There was no significant difference in E. coli detection among the investigated clinical or subclinical cases of both cattle and buffaloes.
The difference in the prevalence of the detected pathogens in the investigated clinical and subclinical mastitis cases of both cattle and buffaloes may be attributed to the low sample size of the examined animals and different rearing locations, milking, and management practices.
In addition to bacterial identification using species-specific PCR assays, 16S rRNA sequencing was performed.
The maximum likelihood phylogeny of two representative E. coli and Salmonella spp. strains was conducted based on the similarities of the 16S rRNA oligonucleotide and revealed a very significant relationship between the study strains and the deposited GenBank strains, which were found to exist for the Escherichia and Salmonella bacteria genera.
In the present study, serotyping of 10 out of 39 recovered E. coli strains revealed 6 different serotypes: O118:K-, O164:K-, O44:K80, O78:K80, O44:K80, and O114:K90. Furthermore, the four recovered Salmonella spp. serovars were identified as three S. typhimurium and 1 S. Entertidis serovar. The results revealed that both recovered E. coli and Salmonella spp. pathotypes are of public health importance, and this triggers the need for their control to limit their dissemination to humans, especially if they are characterized as multidrug-resistant pathotypes.
Concerning the randomly selected E. coli and Salmonella spp. antibiogram profiles, the findings revealed variable exhibited sensitivity and resistance patterns toward the five investigated antimicrobial groups, as shown in Table 3. The highest resistance rates for both E. coli (90%) and Salmonella spp. (100%) were exhibited toward TC antibiotics. Meanwhile, the highest exhibited sensitivity patterns were toward ceftriaxone (70%) and 50% for both clindamycin and ampicillin/sulbactam by E. coli and Salmonella spp., respectively. Precisely, the results of antibiograms are different between different pathogens within the same studies, as well as between various studies, and this may be attributed to the different methods used for antibiogram evaluation, which is considered unacceptable for adequate comparison (Schwarz et al., 2010).
The MDR index was displayed by all 4 tested Salmonella spp isolates and 7 E. coli isolates. However, 3 of the tested E. coli isolates were classified as NDR. The emergence of MDR E. coli and Salmonella serovars is a crucial factor in treatment failure of both animal and human infections (Kimang’a, 2012). Consequently, antimicrobial therapy and mass vaccination have become of low or limited significance in the control of bovine mastitis (Munoz et al., 2007).
Likewise, the emergence of multidrug-resistant pathogens results in further upsurges in antibiotic usage, which will not only result in environmental consequences but also a public health concern (Han et al., 2022) and establishes a great challenge for these pathogens, and advocates for the necessity to explore antimicrobial alternatives as an alternative method of control (Farouk et al., 2015).
Nanoparticles have the potential to be applied as an admirable alternative to antimicrobials for the treatment of veterinary bacterial diseases (Youssef et al., 2019). Chitosan nanoparticles present promising potential for field application in both agricultural and food-safety contexts due to their biodegradability, low toxicity, and strong antimicrobial properties. In agriculture, ChNPs have been proposed as sustainable agronanochemicals: they can be used to deliver biocontrol agents or as biopesticides, enhancing plant growth and protection while reducing the reliance on conventional agrochemicals. (Maluin and Hussein, 2020). ChNPs have been evaluated as natural preservatives in food systems; for example, spray applications on fishery products significantly reduced bacterial contamination without compromising sensory qualities (Elmotyam et al., 2023). In vivo toxicity studies have indicated that chitosan-based nanoparticles generally have favorable biocompatibility, with high LD50 values (often > 5,000 mg/kg) and no significant adverse effects in subacute studies (Salsabila et al., 2025). Aqueous suspensions of ChNPs showed minimal cytotoxicity in toxicity assessments in fish cell lines at concentrations relevant to antimicrobial applications (Ahmed et al., 2020). Moreover, comprehensive literature reviews highlight that ChNPs exhibit negligible toxicity in normal mammalian cells, though toxicity can depend on factors such as dose, exposure time, and particle size (Zoe et al., 2023). Collectively, these data support the feasibility of using ChNPs in practical settings, but also underscore the need for detailed in vivo field trials to establish optimal dosing, long-term safety, and environmental behavior.
In this study, the use of ChNPs is described as a sustainable antibacterial approach due to several inherent properties of chitosan. First, chitosan is a naturally derived, biodegradable, and environmentally safe biopolymer obtained from renewable biomass resources, which minimizes ecological accumulation and reduces environmental impact compared with synthetic antibacterial agents (Rinaudo, 2006; Dash et al., 2011). Moreover, ChNPs exhibit a multi-target antibacterial mechanism—including disruption of the cell membrane, chelation of essential metal ions, and interference with intracellular processes—which collectively lower the likelihood of developing bacterial resistance relative to conventional antibiotics (Goy et al., 2009; Kong et al., 2010). Additionally, chitosan production relies on abundant waste materials, such as crustacean shells, aligning with circular-economy principles and supporting sustainable large-scale utilization (Aranaz et al., 2009). Collectively, these characteristics justify the use of ChNPs as a sustainable and environmentally responsible antibacterial alternative.
Effective antimicrobial effects of chitosan nanoparticles have been exhibited against numerous pathogens, including Gram-negative bacteria (S. typhi, E. coli, and Pseudomonas aeruginosa) and Gram-positive microorganisms (Streptococcus pyogenes and Staphylococcus aureus) (Hassan et al., 2016).
In the current study, the in vitro antibacterial activity of ChNPs with an average size of 20.1–75.2 nm was evaluated against both MDR E. coli and S. Enteritidis strains recovered from bovine clinical mastitis cases. The MIC and MBC values of the tested ChNPs were estimated. Moreover, the mechanism by which ChNPs have antimicrobial effects was investigated based on the TEM imaging assessment of the bacterial cell viability.
ChNPs exhibited a potent visible inhibitory effect at 2.5 mg/ml for both MDR E. coli and Salmonella spp. Meanwhile, the exhibited MBC for both tested strains was at 10 mg/ml concentration. These findings may be different from those of various reported studies concerning these two pathogens. This might be explained by the varying nanoparticle sizes, bacterial strain under test, and pathogen isolation source (Fadl et al., 2021).
Furthermore, when comparing our MIC (2.5 mg/ml) and MBC (10 mg/ml) values for ChNPs against Salmonella spp. and E. coli, these concentrations are higher than those previously reported. For instance, Qi et al. (2004) found MIC values of < 0.25 µg/ml and MBC of ~1 µg/ml for E. coli and S. spp. typhimurium using ChNPs prepared via ionic gelation. Xing et al. (2021) reported an MIC of 0.625 mg/ml and MBC of 1.25 mg/ml against E. coli. Zahediyeghaneh et al. (2014) reported MIC/MBC of 0.25–1 mg/ml for E. coli. These differences may stem from variations in nanoparticle synthesis methods (e.g., degree of deacetylation, molecular weight, size, and surface charge) and assay conditions (pH, inoculum size, and incubation time). Such variability underscores the importance of characterizing the physicochemical properties of ChNPs when interpreting their efficacy.
Although the ChNP concentrations used in our antimicrobial assays (2.5–10 mg/ml) may seem high, similar studies have reported bactericidal activity of the same order of magnitude. For example, Ahmed et al. (2020) reported MBC values ranging from 1 to 7 mg/ml for chitosan nanoparticles against fish-pathogenic bacteria. In terms of safety, recent toxicity evaluations have shown that chitosan nanoparticles can be relatively well tolerated: a comprehensive review reported minimal cytotoxicity in non-cancerous cells at concentrations relevant for biological applications (Zoe et al., 2023).
The lower concentration used for TEM imaging (20 µg/ml) is consistent with common practice: TEM sample preparation typically requires dilute suspensions to avoid aggregation and to clearly resolve individual nanoparticles. Des Bouillons-Gamboa et al. (2024) studied chitosan nanoparticles and noted inhibitory effects at similar low-µg/ml concentrations in MIC assays, while preparing TEM samples at much lower concentrations for visualization.
Interestingly, the high concentrations in our antimicrobial tests reflect the efficacious doses required under biological conditions, whereas the low concentration for TEM allows for optimal imaging. Nonetheless, we acknowledge that cytotoxicity was not directly assessed in this study, and future work should evaluate cell viability across the concentration range used.
Concerning the antibacterial efficacy evaluation using TEM, the bacterial culture treated with ChNPs showed clear ChNPs inside the cell cytoplasm. The observed changes may be attributed to various ChNP interactions with bacterial cells. Owing to the polycationic nature of chitosan and the presence of amino groups, it can interact with the negatively charged components of microbial cell membranes, including lipopolysaccharides in gram-negative bacteria. These interactions disrupt the integrity and permeability of the cell membrane, leading to the leakage of intracellular contents, chelation of essential nutrients, and enhanced cellular uptake due to the nanoscale size of the particles and eventual cell death. Moreover, chitosan can infiltrate the cytoplasmic membrane, thereby interfering with vital cellular processes and hindering microbial growth (Nasaj et al., 2024).
The use of convenience sampling is a key limitation of this study, which may introduce selection bias and limit the generalizability of the results. The modest sample size further underscores the need for cautious interpretation and future studies employing randomized sampling strategies. However, this study was conducted as a preliminary exploration to provide insights into the situation of the investigated pathogens in dairy bovines in Egypt. Its results cannot be extrapolated to the entire populations of Egyptian dairy cattle and buffaloes. Therefore, a large-scale epidemiological study is necessary to fully estimate the occurrence of pathogens implicated in the development of environmental mastitis in dairy herds in Egypt.
Another limitation of this study is that a limited number (10 out of 39) of randomly selected recovered E. coli strains were selected for applying the antibiogram due to a lack of financial resources. Although these isolates were selected to represent different animals, sample sources, and colony morphotypes, the smaller subset may not fully capture all isolates’ broader resistance patterns. Consequently, the findings should be interpreted with caution because the antimicrobial profiles reported here may underestimate or overestimate the true prevalence of resistance within the full isolate population. Future studies using a larger or complete set of isolates would provide more robust and representative resistance data.
In addition, one selected MDR S. Enteritidis and E. coli strains were selected for sequencing and investigation of their inhibiting efficacy by ChNPs.
Another limitation of the current study is that the experiments, including inhibition zone measurements and MIC/MBC determinations, were performed without multiple replicates, preventing the calculation of standard deviation or standard error. The precision and variability of these measurements could not be fully assessed. Future studies should include antimicrobial assays based on multiple replicates to improve the statistical robustness of the findings.
Conclusion
The accomplished results clarify that ChNPs have garnered attention as a promising therapeutic strategy for bovine bacterial mastitis; however, further investigations are needed to assess their therapeutic significance in vivo.
Conflict of interest
The authors have no relevant financial or nonfinancial interests to disclose.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Authors’ contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by (Amany Abd El latif Moselh), (Mahmoud Abd El-Mongy), (Nadia Saed Arafa), and (Shaimaa Moawad Nada). The first draft of the manuscript was written by (Manar M. Farouk), and all authors commented on the previous versions of the manuscript. All authors have read and approved the final version of the manuscript.
Data availability
The datasets generated during and/or analyzed during the current study are available in the (Salmonella enterica subsp. enterica serovar Enteritidis strain Na4 16S ribosomal RNA gene, partial sequence GenBank: PP430329.1& Escherichia coli strain Na2 16S ribosomal RNA gene, partial sequence GenBank: PP425900.1) repository, (https://www.ncbi.nlm.nih.gov/nuccore/PP430329 and https://www.ncbi.nlm.nih.gov/nuccore/PP425900).
References
Abdel-Ghany, H.M. and Salem, M.S. 2020. Effects of dietary chitosan supplementation on farmed fish; a review. Rev. Aquacl. 12, 438–452.
Abdel-Razek, N. 2019. Antimicrobial activities of chitosan nanoparticles against pathogenic microorganisms in Nile tilapia, Oreochromis niloticus. Aquac. Intern. 27(5), 1315–1330.
Aghamohammadi, M., Haine, D., Kelton, F.D., Barkema, W.H., Hogeveen, H., Keefe, P.G. and Dufour, S. 2018. Herd-Level Mastitis-Associated Costs on Canadian Dairy Farms. Front. Vet. Med. 5, 100; doi:10.3389/fvets.2018.00100
Ahmed, F., Soliman, F.M., Adly, M.A., Soliman, H.M., El-Matbouli, M. and Saleh, M. 2020. In vitro assessment of chitosan nanoparticles’ antimicrobial efficacy against major fish pathogens and their cytotoxicity to fish cell lines. J. Fish Dis. 43, 1049–1063.
Aranaz, I., Harris, R. and Heras, A. 2009. Amphiphilic chitosan derivatives: chemistry and applications. Curr. Organic Chem. 13, 1050–1071.
Ashraf, A. and Imran, M. 2020. Causes, types, etiological agents, prevalence, diagnosis, treatment, prevention, effects on human health and future aspects of bovine mastitis. Anim. Health. Res. Rev. 21, 36–49.
Barlam, T.F., Cosgrove, S.E., Abbo, L.M., Macdougall, C., Schuetz, A.N., Septimus, E.J., Srinivasan, A., Dellit, T.H., Falck-Ytter, Y.T., Fishman, N.O., Hamilton, C.W., Jenkins, T.C., Lipsett, P.A., Malani, P.N., May, L.S., Moran, G.J., Neuhauser, M.M., Newland, J.G., Ohl, C.A., Samore, M.H., Seo, S.K. and Trivedi, K.K. 2016. Implementing an Antibiotic Stewardship Program: guidelines by the Infectious Diseases Society of America and the Society for Health care Epidemiology of America. Clin. Infect. Dis. 62, e51–e77; doi:10.1016/j.clindis.2012.01.013
Batool, S.A., Kalsoom, R., Rauf, N., Tahir, S.S. and Hussain, F. 2012. Microbial and physicochemical quality assessment of raw and pasteurized milk supplied in the Twin City of Pakistan. Internet. J. Food. Saf. 14, 17–22.
Castro-Valenzuela, B.E., Franco-Molina, M.A. and Rodríguez-Padilla, C. 2025. Nanoparticles as an alternative treatment for bovine mastitis: a review. Anim. Biosci. 38(7), 1291–1304.
Cheesbrough, M. 2006. District Laboratory Practice in Tropical Countries. Cambridge: cambridge University Press. Cambridge, UK: Cambridge University Press.
Chehabi, C.N., Nonnemann, B., Astrup, L.B., Farre, M. and Pedersen, K. 2019. In vitro Antimicrobial Resistance of Causative Agents to Clinical Mastitis in Danish Dairy Cows. Foodborne Pathog. Dis. 16, 562–572.
Christopher, A.F., Hora, S. and Ali, Z. 2013. Investigation of plasmid profile and antibiotic susceptibility pattern multiple antibiotic resistance index calculation of Escherichia coli isolates obtained from different human clinical specimens at a tertiary care hospital in Bareilly, India. Ann. Trop. Med. PH. 6, 285–289.
CLSI (2020). Performance standards for antimicrobial susceptibility testing.Clinical Lab Standards Institute.
Crispie, F., Flynn, J., Ross, R.P., Hill, C. and Meaney, W.J. 2004. Dry Cow Therapy with a Non-Antibiotic Intramammary teat Seal—A Review. Int. Vet. J. 57, 412.
Dash, M., Chiellini, F., Ottenbrite, R.M. and Chiellini, E. 2011. Chitosan—A versatile semi-synthetic polymer in biomedical applications. Prog. Polym. Sci. 36, 981–1014.
Des Bouillons-gamboa, R.E., Montes De Oca, G., Baudrit Jrv, R., íos Duarte LC, Lopretti., Renter, M., Ía Urquiza, M.Z., úñiga-Umaña JM Barreiro. and Vázquez, P. 2024. Synthesis of chitosan nanoparticles (CSNPs): effect of the CHCH-TPP ratio on the size and stability of NPs. Front. Chem. 12, 1469271.
Dingwell, R.T., Kelton, D.F. and Leslie, K.E. 2003. Management of the dry cow in control of peripartum disease and mastitis. Vet. Clin. N. Am. Food. Anim. Pract. 19, 235–265.
Dohoo W Martin. and Stryhn. 2009. Veterinary Epidemiologic Research. Charlottetown, Prince Edward Island, Canada: vER Inc. Canada.
Elmotyam, H.A., Belal, A.M., Fouad, H.M., Mohamed, A.N., Elkasas, E.N. and Aboul-Ela, M.H. 2023. Potential Application of Chitosan Nanoparticles as a Preservative Agent for Fishery Products Egypt. J. Aquatic. Biol. Fish. 27(4), 785–799.
Eskandari, M., Abdolmaleki, Z., Moosakhani, F. and Eslampour, M.A. 2025. Antimicrobial Efficacy of Cloxacillin-Loaded Chitosan Nanoparticles against Staphylococcus aureus Biofilms in Subclinical Mastitis. Curr. Microbiol. 3(3), 65.
Fadl, A.F., Abu Aita, A.N., Abelaziz, A.M. and Mohamed, A. 2021. In vitro assessment of the antimicrobial activity of zinc oxide nanoparticles against pathogenic Streptococcus parauberis. Adv. Anim. Vet. Sci. 9(6), 913.
Farouk, M.M., El-Molla, A., Salib, A.F., Soliman, A.Y. and Shaalan, M. 2015. Role of Silver Nanoparticles in the Treatment of Multidrug-Resistant Salmonella Species Isolates. Int. J. Nanomed. 15, 6993–7011.
Felsenstein, J. 1985. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39(4), 783–791.
Fisher. 1925. Statistical methods used by research workers. Edinburgh, UK: Oliver and Boyd.
Godoy, C.A., Balic, I., Moreno, A.A., Diaz, O., Arenas Colarte, C., Bruna Larenas, T., Gamboa, A. and Caro Fuentes, N. 2025. Antimicrobial and Antibiofilm Activity of Chitosan Nanoparticles Against Staphylococcus aureus Strains Isolated from Bovine Mastitis Milk. Pharmaceutics 17(2), 186; doi:10.1016/j.pharmaceutics.2017.09.010
Goy, R.C., Britto, D.D. and Assis, O.B.G. 2009. A review of the antimicrobial activity of chitosan. Polímeros 19, 241–247.
Grimont, P.A., & Weill, F.X. (2007). WHO Collaborating Center for Reference and Research on Salmonella “Antigenic Formulae of the Salmonella Serovars.” 9th ed. Paris, France: Institut Pasteur; ed.
Han, G., Zhang, B., Luo, Z., Luo, B., Luo, Z., Zhang, J. and Zhang, J. 2022. 9*+-Molecular typing and prevalence of antibiotic resistance and virulence genes in Streptococcus agalactiae isolated from Chinese dairy cows with clinical mastitis. PLos One 17(5), 268262.
Hassan, I.M., Mohamed, F.A., Taher, A.F. and Kamel, R.M. 2016. Antimicrobial activities of chitosan nanoparticles prepared from Lucilia Cuprina maggots (Diptera: calliphoridae). J. Egypt. Soc. Parasitol. (JESP). 46(3), 563–570.
Hu, Q.H., Tu, J., Han, X.G., Zhu, Y.Y., Ding, C. and Yu, S.Q. 2011. Development of multiplex PCR assay for rapid detection of Riemerella anatipestifer, Escherichia coli, and Salmonella enterica simultaneously from ducks. Methods Mol. Biol. 87, 64–69.
International Organization for Standardization −6579. (1998). Microbiology of food and animal feeding stuff-horizontal method for the detection of Salmonella. Geneva: ISO; 1998.
Kandeel, S.A., Morin, D.E., Calloway, C.D. and Constable, P.D. 2018. Association of California Mastitis Test Scores with Intramammary Infection Status in Lactating Dairy Cows Admitted to a Veterinary Teaching Hospital. Intern. Med. 32, 497–505.
Kimang’a, A.N. 2012. Situational analysis of antimicrobial drug resistance in Africa: are we losing the battle?. Ethiop. J. Health. Sci. 22, 135–143.
Kok, T., Worswich, D. and Gowans, E. 1996. Some serological techniques for microbial and viral infections.In Practical Medical Microbiology. Collee, J., Fraser, A., Marmion, B. and Simmons, A 14th, Edinburgh, UK: Churchill Livingstone.
Kong M Chen., Xing, X.G. and Park, H.J. 2010. Antimicrobial properties of chitosan and its mode of action: a state-of-the-art review. Int. J. Food. Microbiol. 144, 51–63; doi:10.1016/j.ijfm.2014.01.010
Kruskal, W.H. and Wallis, W.A. 1952. Use of ranks in the one-criterion variance analysis. J. Am. Stat. Assoc. 47(260), 583–621.
Lagacé, L., Pitre, M., Jacques, M. and Roy, D. 2004. Identification of the bacterial community of maple sap by using amplified ribosomal DNA (rDNA) restriction analysis and rDNA sequencing. Appl. Environ. Microbiol. 70(70), 2052–2060; doi:10.1016/j.applenviron.2012.06.020
Lebofe, M.J. and Pierce, B.E. 2012. A photographic atlas for the microbiology laboratory. Englewood, USA: Morton Publishing Company.
Maluin, F.N. and Hussein, M.Z. 2020. Chitosan-based agronanochemicals as a sustainable alternative in crop protection. Molecules 25, 1611.
Munoz, M.A., Welcome, F.L., Schukken, Y.H. and Zadoks, R.N. 2007. Molecular epidemiology of two outbreaks of Klebsiella pneumoniae mastitis on a dairy farm in New York State. J. Clin. Microbiol. 45(12), 3964–3971.
Nasaj, M., Chehelgerdi, M., Asghari, B., Ahmadieh-Yazdi, A., Asgari, M., Saber Samani, K., Sharifi, E. and Arabestani, M. 2024. Factors influencing the antimicrobial action of chitosan and its derivatives: a review. Int. J. Biol. Macromolecules. 277, 134321.
Oliveira, S.D., Rodenbusch, C.R., Ce, M.C., Rocha, S.L.S. and Canal, C.W. 2003. Evaluation of selective and non-selective PCR enrichment. Lett. Appl. Microbiol. 36(4), 217–221.
Orellano, M.S., Isaac, P., Breser, M.L., Bohl, L.P., Conesa, A., Falcone, R.D. and Porporatto, C. 2019. Chitosan NPs enhance the antibacterial activity of the native polymer against bovine mastitis pathogens. Carbohydr. Polym. 213(1), 1–9.
Osman, M.K., Hassan, M.H., Orabi, A. and Abdelhafez, S.A. 2014. Phenotypic, antimicrobial susceptibility profile and virulence factors of Klebsiella pneumoniae isolated from buffalo and cow mastitic. Pathog. Glob. Health. 108, 191–199.
Parvekar, P., Palaskar, J., Metgud, S., Maria, R. and Dutta, S. 2020. Minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of silver nanoparticles against S. aureus. Biomater. Investig. Dent. 23(71), 105–109.
Patil, N.A., Satbige, S.A., Awati, B. and Halmandge, S. 2021. Therapeutic management of subclinical mastitis in buffaloes. Bufalo Bull. 40(1), 157–160.
Paul, S., Bezbaruah, R.L., Roy, M.K. and Ghosh AC. 1997. Multiple antibiotic resistance index and its reversion in Pseudomonas aeruginosa. Lett. Appl. Microbiol. 24, 169–171.
Pearson, K. 1900. On the criterion that a given system of deviations from the probable in the case of a correlated system of variables is such that it can be reasonably supposed to have arisen from random sampling. London. Edinburgh. Dublin Phil. Mag. J. Sci. 50(302), 157–175.
Qi, L., Xu X Jiang., Hu. and Zou. 2004. Preparation and antibacterial activity of CSNPs. Carbohydrate. Res. 339, 2693–2700.
Quin, P. J., marquee, B. K., Carter, M. E., et al (2002). Veterinary microbiology and microbial diseases Bodmin, Cornwall, UK: Books Ltd; 2002:114–118.
Rinaudo. and M. 2006. Chitin and chitosan: properties and applications. Prog. Polym. Sci. 31, 603–632.
Rivera Aguayo, P., Bruna Larenas, T., Alarcón Godoy, C., Cayupe Rivas, B., González-Casanova, J., Rojas-Gómez, D., and Caro Fuentes, N. (2020). Antimicrobial and Antibiofilm Capacity of Chitosan Nanoparticles against Pseudomonas sp. Isolated from milk of cows diagnosed with bovine mastitis. Antibiotics 2009; 9: 551.
Ruegg, P.L. 2002. Investigation of mastitis problems in farms. Vet. Clinics. North. Amer. Food. Anim. Pract. 18(2), 347–368.
Salsabila, S., Khairinisa, M.A., Wathoni, N., Sufiawati, I., Mohd Fuad, W.E., Khairul Ikram, N.K. and Muchtaridi, M. 2025. In vivo toxicity of chitosan-based nanoparticles: a systematic review. Artif. Cells. Nanomed. Biotechnol. 53(1), 1–15.
Schalm, O.W. and Noorlander, D.O. 1957. Experiments and observations leading to the development of the CAMT. J. Am. Vet. Med. Assoc. 130, 199–204.
Schwarz, D., Diesterbeck, U.S., Failing, K., König, S., Brügemann, K., Zschöck, M., Wolter, W. and Czerny, C.P. 2010. Somatic cell counts and bacteriological status in quarter foremilk samples of cows in Hesse, Germany: a longitudinal study. J. Dairy. Sci. 93, 5716–5728.
Shapiro, S.S. and Wilk, M.B. 1965. An analysis of variance test for normality (complete samples). Biometrika 52(3-4), 591–611.
Xing, Y., Wang, X., Guo, X., Yang, P., Yu, J., Shui, Y., Chen, C., Li, X., Xu, Q., Xu, L. and Bi, X. 2021. Comparison of antimicrobial activity of chitosan nanoparticles. Coatings 11(7), 769.
Youssef, S.F., El-Banna, A.H., Elzorba, Y.H. and Galal, M.A. 2019. Application of nanoparticles in veterinary medicine. Int. J. Vet. Sci. Med. 7(1), 78–93.
Zahediyeghaneh, Z. and Hadizadeh, M. 2014. Comparison of the antibacterial property of chitosan nanoparticles against Escherichia coli and Staphylococcus aureus. J. Inflamm. Diseases 19(6), 21–28.
Zoe, L.H., David, S.R. and Rajabalaya, R. 2023. Toxicity of chitosan nanoparticles: a comprehensive literature review of in vivo and in vitro assessments for medical applications. Toxicol. Rep. 11, 83–106.

















