| Research Article | ||
Open Vet. J.. 2026; 16(2): 847-860 Open Veterinary Journal, (2026), Vol. 16(2): 847-860 Research Article Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutritionIsmah Ulfiyah Azis1*, Athhar Manabi Diansyah1, Andi Nimahtul Churriyah2, Muhammad Ihsan Andi Dagong1, Fahrul Irawan1, Erni Damayanti1, Masturi Masturi1, Rusli Fidriyanto3, Tulus Maulana3, Aeni Nurlatifah4 and Sitti Nurhaliza51Faculty of Animal Science, Hasanuddin University, Makassar, Indonesia 2Department of Animal Science, Faculty of Agriculture, Lambung Mangkurat University, Banjarmasin, Indonesia 3Research Center for Applied Zoology, National Research and Innovation Agency, Cibinong, Indonesia 4Faculty of Animal Science, Gadjah Mada University, Yogyakarta, Indonesia 5Faculty of Mathematics and Natural Science, Tadulako University, Palu City, Indonesia *Corresponding Author: Ismah Ulfiyah Azis. Faculty of Animal Science, Hasanuddin University, Makassar, Indonesia. Email: ismahulfiyahazis [at] unhas.ac.id Submitted: 22/10/2025 Revised: 25/12/2025 Accepted: 05/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
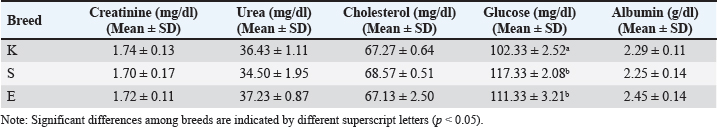
AbstractBackground: The productivity of goats in the tropics is largely hampered by low feeding efficiency and inefficient nutrient utilization. Understanding the breed-specific metabolic response is essential for developing a feeding strategy under low environmental conditions. Metabolomics provides a systemic approach to identifying metabolic changes that would not be detected by conventional biochemical markers. Aim: This study aimed to identify breed-specific metabolic differences among tropical goat breeds and identify potential nutritional biomarkers for precision feeding. Methods: An integrated biochemical and Gas Chromatography–Mass Spectrometry (GC-MS)-based metabolomic profiling approach was used in Kacang, Etawa crossbred, and Saanen goats (n=9). Three animals from each breed were included for comparative analysis. An analysis investigation involving analysis was carried out using biochemical, statistical methods ANOVA and multivariate statistics (principal component analysis, and partial least squares–discriminant analysis) in plasma. Results: A distinct breed-specific clustering and metabolite patterns characterized the two biotic groups of Saanen goats, which were characterized by enriched carbohydrate-related metabolites, including sorbitol and D-xylose, potentially reflecting increased energy turnover associated with high milk production potential. Compared with the Etawa crossbred goat, the kacang goat (L-alanine), L-isoleucine, L-methionine etc were higher suggesting that glucoseand alanine -aspartate- glutamate metabolism pathway was a predominant pathway supporting energy regulation and protein turnover. Conclusion: This study revealed clear breed-specific plasma metabolomic profiles among Kacang, Etawa crossbred, and Saanen goats, highlighting distinct carbohydrate- and amino acid–related pathways that underlie differences in energy and protein metabolism. These findings yield preliminary candidate plasma metabolites linked to feed efficiency and environmental adaptation, providing a basis for precision nutrition strategies to improve productivity and sustainability in tropical goat farming. Keywords: Goat breeds, Plasma metabolomics, Nutritional biomarkers, Pathway enrichment, Precision nutrition. IntroductionGoat production contributes substantially to food security, particularly in tropical regions where these animals provide affordable sources of meat and milk while sustaining rural livelihoods (Michalk et al., 2019; Cooke et al., 2024). Goats are often raised under a smallholder system that relies on locally available feed resources, allowing them to thrive in environments where other livestock species struggle to survive. Despite their remarkable adaptability to marginal conditions, the productivity of many goat production systems remains suboptimal. This limitation is largely driven by nutritional inefficiency, inconsistent feed quality, and limited understanding of the physiological mechanisms that underlie nutrient utilization (Godber et al., 2020). Therefore, improving feed efficiency and metabolic adaptability is a crucial step toward strengthening the role of goats in achieving sustainable livestock production and food security in tropical regions. Conventional biochemical indicators, such as blood glucose, urea, creatinine, and cholesterol levels, have long been used to assess the nutritional and physiological status of animals (Kaniamuthan et al., 2025). These markers provide useful diagnostic information for evaluating animal health and nutritional balance. However, they only capture isolated aspects of metabolism and fail to represent the organism. In practical terms, relying solely on these parameters limits our understanding of how nutrients are processed, distributed, and transformed across multiple pathways in response to environmental or dietary variations. This shortcoming highlights the need for a more holistic, systems-level approach that can integrate multiple metabolism dimensions to evaluate adaptive nutritional responses in livestock. Here, metabolomics comes into play to circumvent these challenges as a potential source of lead. It allows for the profiling of small molecules (metabolites) that are intermediates and products of metabolism, reflecting the physiological state of an organism (Rinschen et al., 2019). In contrast to traditional biochemical analyses of just a few metabolites, metabolomics enables the simultaneous determination of hundreds of endogenous metabolites related to energy/glucose, amino acids, and lipid metabolism. Such a global approach provides a complete overview of animal reactions to nutrition, physiological challenges, or environmental modifications. Through its framework, biomarkers associated with feed efficiency and metabolic resilience/health state can be identified. Recent data in cattle and sheep have demonstrated the possibility of predicting metabolic efficiency (metabolic profiling) with a given accuracy when combining molecular information percentage precision feeding programs. This suggests the potential to cross-bridge molecular data to those phenotypes for which we are actually measuring. However, the use of metabolomics in goats is very limited. Goats are an important component of the tropical livestock system; however, little research has been done in this regard compared with their counterparts, namely, cattle and sheep. This knowledge gap is especially concerning with the extreme variability existing among goat breeds at the level of genotypic, production objectives and adaptability. Moreover, local goat breeds are generally more adaptable to low-quality feed and unfavorable conditions than high-producing dairy breeds, which have higher nutrient requirements and may become more sensitive and vulnerable to metabolic disturbance (Kapusniaková, 2023). These differences demonstrate the importance of species-specific studies on physiological adaptation for productivity and metabolic health. However, the literature on breed-specific goat metabolic processes is scarce. Nutritional strategies tend to be of a general nature and may compromise feeding efficacy and possibly even animal welfare when not considered (Sonea et al., 2023). To circumvent this bottleneck, an untargeted approach that combined traditional biochemical assays with untargeted metabolomic profiling was necessary. Not only classical physiological modulators but also detailed fingerprints of metabolites were considered at the same time, providing a global view of metabolic adaptation. Furthermore, the comparison of different genetically diverse breeds is an excellent source of valuable information for understanding how biochemically reflected phenotypic differences in production type, metabolism, and adaptation. Three goat breeds have opposite physiological and production characteristics. A promising model for nutrient efficiency and environmental adaptation is the Kacang goat: a native Indonesian breed that is phenotypically meat-type goats that are well adapted to low-quality feed and limited resource environments. Open-access novel crossbreeding of Etawa goats and crossing between local and Jamnapari goats reared for dual purposes (milk and meat) with intermediate productivity ability and balance adaptability. Saanen goats, originally from Europe, are a dairy breed with high milk production that has been shown to have higher metabolic turnover due to their high milk-producing capacity and require a nutrient-rich diet. The fact that these three breeds could be compared directly, whereas local and crossbred dairy types cannot easily be managed similarly, provides an opportunity to identify metabolic markers relevant for feed efficiency, production potential, and adaptability. In addition to conventional biochemical indices, pathway-level interpretation of metabolomic data is essential for understanding how sets of metabolites are linked to underlying physiological processes related to energy metabolism, protein turnover, and environmental adaptation. Kyoto Encyclopedia of Genes and Genomes (KEGG)–based pathway analysis enables the mapping of identified plasma metabolites onto canonical metabolic pathways, thereby revealing the biochemical routes that are differentially represented among breeds (Adetunji et al., 2025). KEGG pathway enrichment was used to contextualize breed-specific metabolomic signatures in terms of carbohydrate and amino acid metabolism, providing mechanistic insight into the physiological basis of feed efficiency and adaptation in tropical goats. Materials and MethodsExperimental designThe study utilized three goat breeds of the tropical production system: Kacang (K), Saanen (S), and their crossbreed Etawa (E). Nine adult clinically healthy goats (3 heads per breed) were obtained from the experimental herd of the Faculty of Animal Science, Hasanuddin University in Makassar, Indonesia. All the goats were 2–3 years of age, adult females who were both non-pregnant and non-lactating and similar in body condition to minimize physiological variability. The animals were managed under normal husbandry and welfare conditions with ad libitum water access and a basal diet appropriate to their nutritional needs. Elephant grass and concentrate (60:40 ratio) as basal diet, which contained approximately 14.5% of crude protein and 65% of total digestible nutrients, were offered to the animals. Biochemical analysis of blood metabolitesBlood sampling and metabolomic analyses were conducted to capture the metabolic responses of the breed under common environmental feeding conditions. Samples were collected in the morning (9–11 a.m.) to avoid the influence of circadian rhythms on metabolite plasma concentrations. Blood (approximately 3 ml) was aseptically withdrawn from the jugular vein and put into vacutainer tubes with Ethylenediaminetetraacetic Acid (EDTA) as an anticoagulant. Plasma was collected after centrifugation at 3,000 rpm for 15 minutes and stored at −20ºC until subsequent biochemical and metabolomic assays (Nurlatifah et al., 2022). Biochemical measures of blood plasma were assessed to investigate basic energy and protein metabolism indicators. Blood biochemical analyses were performed based on the method of Nurlatifah et al. (2022) and blood metabolite was used according to the method of Aziz et al. (2023). The following parameters were estimated: creatinine (mg/dl), cholesterol (mg/dl), glucose (mg/dl), and albumin (g/dl). The plasma samples collected after centrifugation were analyzed using an enzymatic colorimetric method and commercial diagnostic kits (Sigma-Aldrich, St. Louis, MO) according to the manufacturer’s instructions. Metabolomic analysis (GC–MS)Metabolomic profiling of plasma samples was performed as described in Huang et al. (2023) with slight modifications. GC-MS Analysis GC-MS analysis was performed using a SH-Rxi-5 MS capillary column (Agilent, 30 m × 0.25 mm ID × 0.25 μm film thickness) in a Shimadzu GCMS-QP2010 system. Pooled quality control (QC) samples were used to control quality, and blank injections during the same analytical run were used as both a measure of consistency and to monitor for contamination. Sample injection was randomized to avoid equipment drift and batch effects. The compounds were tentatively identified by matching the mass spectra to that of NIST 20 and confirmed by libratory Gas Chromatography (GC) retention indices (RI) available in the NIST Chemistry WebBook. After comparison with the in-house library, only those metabolites with a match factor above 80% and RI tolerance (±10–20) were considered for putative identification. The validation of the threshold and RI was performed similarly to other methods in GC-MS-based matabolomics to increase specificity and reduce false positives. Statistical analysisThe raw GC-MS chromatograms were analyzed with Shimadzu GC-MS solution for peak-picking Xsignal (signal-to-noise ratio=5; minimum peak width=5 seconds) and aligned, allowing a retention time tolerance of ±0.2 minutes to compensate for slight drifts in the retention time. Ribitol was chosen as an internal standard to estimate the concentration because it is chemically stable and none of the endogenous plasma metabolites interfered with its use. The peak intensity was normalized against the internal standard and log-converted before statistical analysis. To avoid bias, missing values were filled with half of the minimum positive value observed in all means. To ensure reproducibility and transparency, all processed peak data and raw chromatograms will be reported in the metabolites repository. Comparisons among breeds regarding biochemical parameters and selected metabolite intensities were performed using one-way ANOVA followed by Tukey’s HSD post hoc test, with p < 0.05 considered statistically significant. Principal component analysis (PCA) was used as an unsupervised approach for an overview of global metabolic differences between breeds. Partial least squares–discriminant analysis (PLS–DA) was employed to identify discriminant metabolites, and variables with VIP > 1.0 were significant contributors. In the supervised PLS–DA models, R²Y and Q² values ranged from 0.982 to 0.999 and 0.930 to 0.945, respectively, indicating good model fit and predictive performance. A 1,000 time permutation test (p < 0.01) was used to validate the model and exclude overfitting, and 10fold cross validation was also applied to ensure model robustness. Identified plasma metabolites were then annotated and mapped to metabolic pathways using the KEGG database. Compound identifiers were first matched to KEGG compound IDs, and pathway enrichment analysis was performed to determine which metabolic pathways were overrepresented among the discriminant metabolites identified by the PCA and PLS–DA multivariate models. MetaboAnalyst 6.0 (www.metaboanalyst.ca) and SPSS v26.0 (IBM Corp., Armonk, NY) were used for all statistical analyses. Ethical approvalThe Animal Ethics Committee, Faculty of Animal Science, Hasanuddin University (approval no. 009/UN4.12/EC/V/2025) reviewed and approved the experiment protocol, which adhered to recommendations provided in the national guidelines for experimental animal care and use. Efforts were made to alleviate the suffering of animals, and the guidelines for animal welfare were strictly followed during the experiment. ResultsBiochemical parameters of blood plasmaTable 1 presents the serum biochemical parameters of different blood plasma goat breeds. In generals, the levels of creatinine, ureas, and albumin did not vary between the breedss, indicating that the biochemical profiles of these parameters were quite consistent. On the other hand, glucose concentration had a highly significant breed effect, with K having lower levels than S and E (p ˂ 0.05). Table 1. Biochemical parameters of plasma blood in different goat breeds.
Metabolomic profilingFigure 1 illustrates the distribution of detected plasma metabolites per breed. There were 26 (34.7%) metabolites common to K, E, and S goats, each of which also showed unique sets of metabolites. Kacang harbored more breed-specific metabolites, followed by Etawa and Saanen, with a smaller subset shared by both breeds. Breed-dependent physiological adaptation to energy metabolism and nutrient use is evident in breed-specific hormone response patterns.
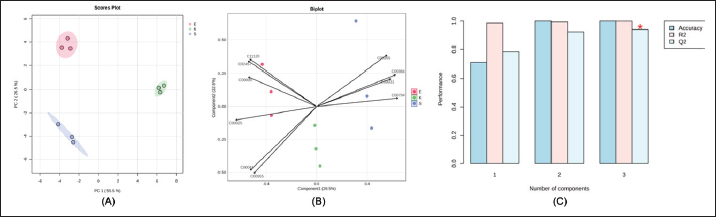
Fig. 1. Venn diagram showing shared and unique plasma metabolites identified across Kacang (K), Etawa (E), and Saanen (S) goat breeds after QC filtering. Multivariate analysis also suggested breed-dependent formation of metabolomic profiles. PCA analysis (Fig. 2A) revealed a clear clustering of the three breeds with the top two principal components accounting for most variance. The corresponding biplot (Fig. 2B) demonstrated that a unique discriminant marker set caused the observed differentiation. The supervised PLS-DA model also showed great interbreed classification and robustness with high accuracy, R2, and Q2 values (Fig. 2C), confirming the results’ reliability. The evident separation of the breeds in the PCA and PLS-DA plots indicated that they possess individual-specific metabolic fingerprints. These differences may be associated with genetic diversity and breed-specific adaptation to feed efficiency, production type, and environmental settings. There is a clear-cut clustering, which supports the purpose of the study, which was to identify goat metabolomic biomarkers for precision nutrition.
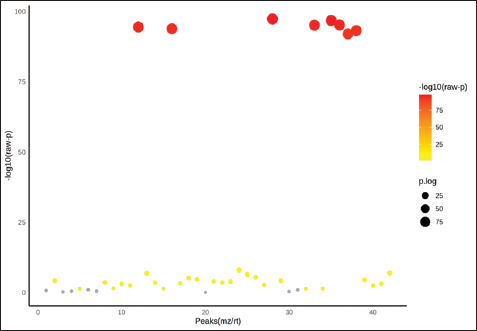
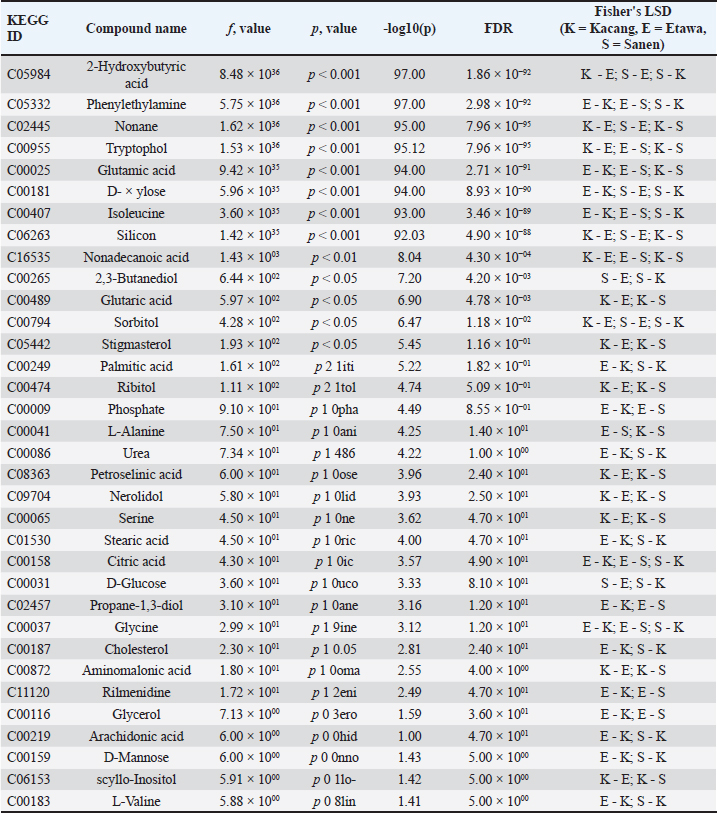
Fig. 2. Multivariate analysis of plasma metabolomic profiles in different goat breeds. (A) Principal component analysis score plot showing separation among breeds. (B) PCA biplot indicating discriminant metabolites contributing to breed clustering. (C) PLS–DA cross-validation plot showing model performance across components. Differential metabolitesThe mean values of the plasma metabolites in goat breeds are presented in Figure 3, with some metabolites having significant differences (FDR-adjusted p ˂ 0.05). Table 2 shows a complete list of identified differential metabolites, and most of the metabolites (e.g., C05984, C02445, C00955, C00025, and C00181) were both statistically significant at strong levels as well as having very low FDRs across the two breeds. Other metabolites, such as C16535, C00265, and C00794, also had moderate associations, and a few exhibited weaker correlations that were not significant (for FDR) after being corrected. Differential metabolites are mainly involved in carbohydrate, amino acid, and lipid metabolism. For instance, D-glucose, sorbitol, and 2,3-butanediol participate in carbohydrate metabolism, from which energy is derived, while L-alanine and glutamic acid act as crucial intermediates in amino acid metabolism and nitrogen circulation. The differences in these profiles indicate that the breeds likely have different mechanisms of energy and nitrogen metabolism modulation, which is consistent with their physiological roles as dairy or meat goats.
Fig. 3. Plasma metabolite distribution based on one-way ANOVA. The y-axis represents –log10 (p-value), and the x-axis corresponds to metabolite peaks (m/z/rt). Red to yellow circles indicate metabolites with significant differences between goat breeds (p < 0.05). Table 2. Differential metabolites across the goat breeds.
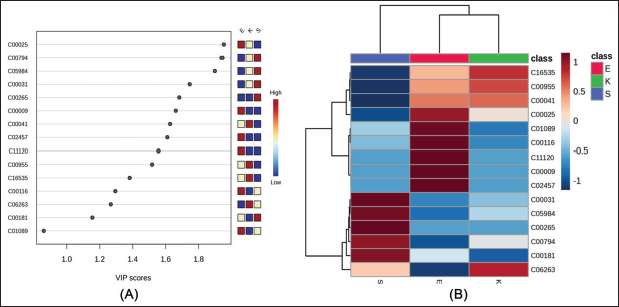
Additional examination of discriminant metabolites by PLS-DA VIP analysis (Fig. 4A) identified several major compounds (e.g., C00025, C00794, C05984, and C00031 with scores > 1.0) that significantly contributed to breed division. Hierarchical clustering of these metabolites (Fig. 4B) showed separation among breeds, and clear breed enrichment or depletion was observed. Significantly, the enriched abundance of carbohydrate-derived metabolites in Saanen goats elevated glycolytic activity for milk production, and higher enrichment levels of amino acid and lipid intermediates in Etawa and Kacang goats demonstrated metabolic flexibility and adaption to feed quality variation. Collectively, these results define a set of metabolites whose response underlies the noted interbreed metabolic adaptation, which is directly fitting to our study aim of identifying nutritional biomarkers for precision feeding interventions.
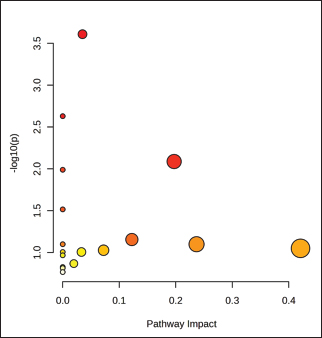
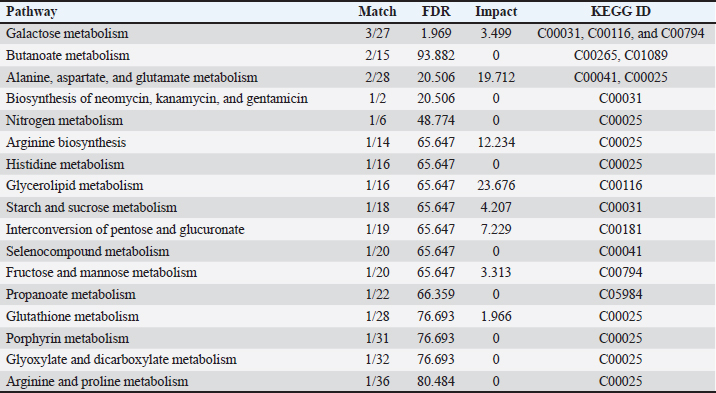
Fig. 4. Discriminant metabolites. (A) Variable importance in projection (VIP) scores from PLS–DA, with higher values indicating a stronger contribution to breed separation. (B) Hierarchical clustering heatmap of significant metabolites showing distinct abundance patterns across goat breeds. Pathway enrichmentPathway enrichment analysis for significantly altered plasma metabolites is shown in Figure 5 and detailed in Table 3. Galactose metabolism, which was supported by three matched metabolites (C00031, C00116, and C00794), remained the most significant pathway with a substantial impact. Furthermore, alanine, aspartate, and glutamate metabolism was enriched with C00041 and C00025, which possess a high pathway impact value. Other pathways that were highlighted included butanoate metabolism, glycerolipid metabolism, starch and sucrose metabolism, and pentose and glucuronate interconversions (although the latter three by having fewer metabolites involved with lower statistical significance). Other pathways, including arginine biosynthesis, nitrogen metabolism, and glutathione metabolism, were assigned based on single metabolite hits but did not meet the levels of significant enrichment. Genetically, galactose metabolism prevails as a metabolic advantage in carbohydrate utilization and lactose biosynthesis, which is very suitable for dairy-type breeds such as Saanen. The enrichment of alanine, aspartate, and glutamate metabolism indicates increased amino acid turnover and nitrogen cycling/adaptive protein utilization in indigenous versus local goats in drought. The pathways of butanoate and glycero lipid metabolism, for example, suggest the participation of lipid oxidation and energy balance mechanisms that can engender metabolic versatility under different nutritional states.
Fig. 5. Pathway enrichment analysis of significant identified plasma metabolites among goat breeds Each bubble represents a metabolic pathway, where the bubble size indicates the pathway impact and the color intensity represents statistical significance (p-value). Galactose and alanine–aspartate–glutamate metabolism emerged as dominant pathways linked to energy regulation and amino acid turnover. Table 3. Pathway enrichment analysis of significant metabolites.
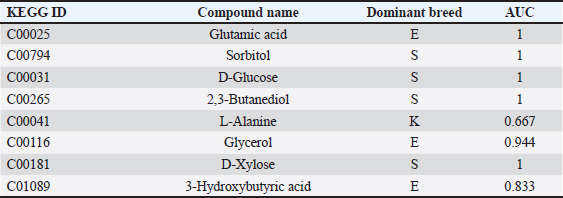
Taken together, these enriched pathways indicate that each goat breed uses different metabolic strategies for energy and nutrient homeostasis. This functional divergence is consistent with the study’s goal of connecting metabolomic patterns with feeding efficiency and designing breed-specific biomarker candidates for sustainable animal production. Potential biomarkersThe sensitivity and specificity of candidate biomarker metabolites were evaluated using the receiver operating characteristic (ROC) curve analysis, as presented in Table 4. A few metabolites with Area Under the Curve (AUC)=1.0 were identified, such as glutamic acid (C00025), sorbitol (C00794), 2,3-butanediol (C00265), and D-xylose (C00181), suggesting that they are effective breed-specific biomarkers. Other metabolites, such as glycerol (C00116; AUC=0.944) and third-hydroxybutyric acid (C01089; AUC=0.833), had robust diagnostic values, while L-alanine (C00041; AUC=0.667) had low predictive ability. These metabolites represent pivotal components of nutrient metabolism that differentiate breeds. Glutamic acid and L-alanine are the precursors of amino acid metabolism, which reflects the rate of nitrogen turnover between breeds. Sorbitol, D-glucose, and D-xylose are intermediates of dairy-type goats. Glycerol and 3-hydroxybutyric acid, on the other hand, indicate lipid mobilization and ketone body formation, which are accustomed measures of energy state and metabolic capacity. Table 4. Potential biomarker metabolites identified from ROC analysis.
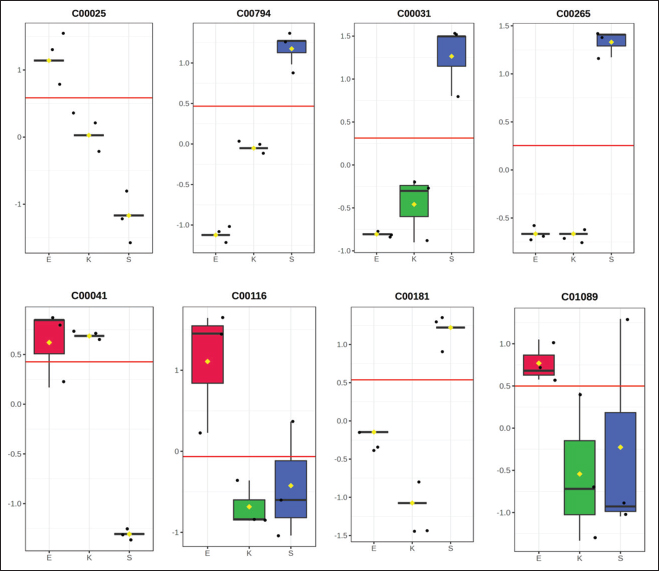
Figure 6 shows representative boxplots of the selected discriminant metabolites. There were mixed breed-specific patterns, with glutamic acid (C00025) enriched in Etawa, sorbitol (C00794) and D-xylose (C00181) enhanced in Saanen and L-alanine higher relatively higher abundance in Kacang. Some metabolites (e.g., glycerol (C00116) and 3-hydroxybutyric acid (C01089)) showed mid-range but consistent breed-specific profiles. These distributions provide an obvious picture of metabolic specialization throughout the breeds, with Saanen goats favoring carbohydrate metabolism to lactation energy, Etawa preferring amino acid pathways for dual-purpose metabolism, and Kacang goats emphasizing protein-derived substrates to survive in limited diets.
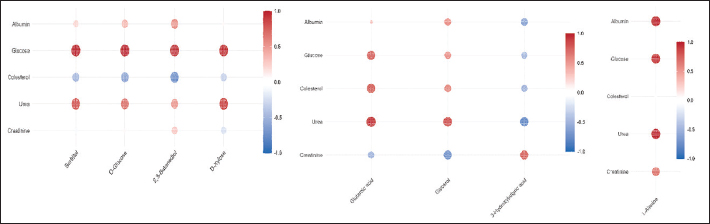
Fig. 6. Boxplots showing the relative abundance of selected discriminant metabolites across breeds. Glutamic acid and L-alanine are elevated in Etawa and Kacang, respectively, while sorbitol and D-xylose are enriched in Saanen. These profiles illustrate the breed-specific metabolic adaptations supporting the utilization of carbohydrates and proteins. Collectively, these results confirm the discovery of a panel of discriminative plasma metabolites, which are potential candidate biomarkers for discrimination between goat breeds. The robust expression of Monoacylglycerols (MGs) increases confidence in their potential use as markers for precision nutrition and gastric health status in goats. Correlation between plasma biochemical and DMsMultiple significant correlations were observed between plasma biochemistry and discriminant metabolites (Fig. 7). In the breeds, significant positive correlations were found between plasma glucose and carbohydrate related metabolites including sorbitol (r=0.81, p ˂ 0.01), D-glucose (r=0.87, p ˂ 0.01), 2,3-butanediol (r=0.68, p < 0.05) and D-xylose (r=.74, p < 0.01) while sorbitol also had a negative correlated with albumin herein; r=−46). amino acid_fmvs2 A log (L-valine/linked leucine) was negatively correlated with current albumin levels (r=−0.88, p < 0.01), and L-alanine was both positively associated with albumin levels (r=0.72, p < 0.01) and negatively correlated with cholesterol levels (r=−0.41, p < 0.05). In Etawa goats showed a positive correlation between glutamic acid with creatinine (r=0.66, p ˂ 0.05), also for energy metabolites i.e glycerol with albumin (r=0.79, p < 1%) and 3-hydroxybutiric acid (r=−71 p <5) but the value of glycerol also has negative relation with cholesterol (r=−52, p < 5). These correlations suggest synchronized regulation of biochemical markers and metabolomic pathways indicative of energy and nutrient metabolism. Strong relationships between glucose and carbohydrate-derived metabolites (sorbitol, D-glucose, and D-xylose) indicate a direct reflection of plasma glucose status on systemic carbohydrate turnover and energy supply, particularly in high-yielding genotypes. The positive association between L-alanine and albumin indicates an adaptive connection between amino acid metabolism and protein synthesis, whereas the negative association between L-alanine and cholesterol represents a metabolic trade-off between protein turnover and lipid metabolism. The correlation of glutamic acid with creatinine in Etawa goats shows improved amino-acid mobilization related to muscle metabolism and that of glycerol and 3-hydroxybutyric acid with albumin displays synchronization between lipid mobilization and protein balance. Overall, these trends establish the connected nature of metabolic networks and confirm that metabolomic-biochemical relationships can reveal physiological processes beyond those provided by single biochemical measures.
Fig. 7. Heatmap of Pearson correlation coefficients between plasma biochemical parameters and metabolites discriminant across breeds. Positive (red) and negative (blue) correlations illustrate integrative metabolic relationships linking carbohydrate, amino acid, and lipid metabolism. Strong associations between glucose and carbohydrate metabolites confirm consistency between biochemical and metabolomic indicators. DiscussionBiochemical assays combined with matabolomic profiling provide a larger window to assess the nutritional physiology of goats. Traditionally, established markers such as glucose, urea, cretinin, and cholesterol have been studied as metrics of metabolic and health well-being based on functional information (Kuru et al., 2022); however, their outlook is by nature circumscribed on discrete processes. The present results show that the metabolomics substrates of this model can be extended to detect global metabolic variation and breed-specific adaptation, as illustrated by multivariate clustering (Fig. 2) and metabolite dispersion (Fig. 1). This complements earlier conclusions in cattle and sheep that conventional indices underestimate the complexity of partitioning (Byrne and Murphy, 2022). Therefore, this study places metabolomics as an essential support to conventional methodologies in improving goat precision nutrigenomics. The plasma biochemical parameters (Table 1) indicated that the levels of creatinine, urea, cholesterol, and albumin remained constant among the different breeds, but the glucose concentrations varied significantly. The higher serum glucose levels in Saanen and Etawa goats may be attributed to their potential for high milk production and metabolic demands compared to indigenous breeds, given that Zamuner et al. (2020); Danso et al. (2024) revealed that high-yielding breeds have a significantly higher blood glucose level than their indigenous counterparts due to lactose synthesis requirements. In contrast, the lower glucose levels in Kacang goats could be attributed to their native adaptation under low-input feeding conditions and metabolism efficiency under restricted resources (Irawan et al., 2024). These findings indicate that energy metabolism, in terms of plasma glucose level, is a sensitive trait that reveals breed differences in physiological demands. Metabolomic profiling added richness to the characterization of breed differences with a good separation in PCA and PLS-DA scores plots (Fig. 2A–C), underlying clear sets of discriminant metabolites. The Venn diagram (Fig. 1) also reflected the above common and specific metabolite sets. Saanen goats are enriched in carbohydrate-derived metabolites such as sorbitol, D-glucose, 2,3-butanediol, and D-xylose, which is consistent with their preference for fast carbohydrate turnover to meet high energy requirements (Ghavipanje et al., 2021). The high levels of glutamic acid, glycerol, and 3-hydroxybutyric acid are metabolites related to nitrogen metabolism and ketone body production, respectively, which is in agreement with previous studies that reported crossbred goats mobilize both protein and lipid pools for milk production (Ediriweera, 2025). However, Kacang goats showed higher levels of L-alanine, which is in agreement with the suggestion that alanine has a critical importance in protein energy adaptation to nutritional stress (Teixeira et al., 2024). As a whole, those metabolite signatures indicated (Table 2; Fig. 3) breed specific dependence on carbohydrate, amino acid, or lipid metabolism, which is congruent with the fact that in other ruminants genetic and production type: metabolic fingerprints reflect genetic and production type differences (Zhang et al., 2020). Pathway analysis (Fig. 5; Table 3) put the metabolite differences in a biological network. In dairy animals, galactose metabolism was the most significant pathway, which includes sorbitol, glycerol, and D-glucose essential for energy regulation and lactose synthesis (Liu et al., 2024). This enrichment indicates a more pronounced carbohydrate-dependent metabolism in dairy-type goats, such as the Saanen, and is congruent with their higher glucose exploitation for milk production and energy maintenance. The metabolism of alanine, aspartate, and glutamate also appears to be important, which is consistent with amino acid turnover and the need for nitrogen recycling, which has been demonstrated to be critical for the protein utilization efficiency of ruminants (Daddam et al., 2025). This implies that Etawa goats efficiently recycle amino acids to maintain energy and protein balance, which is a physiological adaptation of the crossbreed. Other pathways were butanoate metabolism, starch and sucrose metabolism, pentose and glucuronate interconversions, fructose and mannose metabolism, propanoate metabolism, glycerolipid metabolism, and glutathione metabolism. Taken together, these pathways suggest a flexible relationship between carbohydrate and lipid metabolism, which enables various goat breeds to adapt to their nutritional or environmental conditions. Althought represented by fewer metabolites, these pathways show lipid mobilization, carboydrate interconversion, and oxidantive balance, which is consistent with evidence that ruminant metabolism can adaptively renallocate its substrates in response to dietary manipulation (Nunes et al., 2024). One-metabolite pathways, including nitrogen metabolism, arginine biosynthesis, arginin and proline metabolism, porphyrin merabolism, and glyoxylate/dicarboxylate metabolism, were also found. Although not as enriched, its presence highlights the metabolic versatility of cross breeds. Taken together, pathways-level results generally support that the between-breeds metabolic differences are polygenic and multifactorial, which is consistent with the findings of a multibreed analysis from a cattle study (Wu et al., 2024). Taken together, these data indicate that every breed uses a different biochemical strategy—carbohydrate utilization in Saanen, integration of amino acid and lipid metabilism in Etawa, and energy conservation in Kacang—of specific physiological adaptive mechanisms. The discriminating performance of eight metabolites as candidate biomarkers was rated by the whole ROC analysis (Table 4; Fig. 6). The AUC values for glutamic acid (C00025), sorbitol (C00794), D-glucose (C00031), 2,3-butanediol (C00265), and D-xylose (C00181) were all perfect at 1.0, indicating that they can be used as biomarkers for breed differentiation. Glycerol (C00116; AUC=0.944) and 3-hydroxybutyric acid (C01089; AUC=0.833) also exhibited strong discriminatory ability, which was in line with their involvement in ruminant lipid and energy metabolism (Deda et al., 2024). L-alanine (C00041) was also moderately predictive (AUC=0.667), and its relationship with albumin may implicate it as an indicator of protein turnover, a pattern previously documented in stress-adapted sheep (Ginting et al., 2020). These biomarker patterns reflect the physiological diversity of goat breeds: carbohydrate-related metabolites dominate in high-yielding breeds, whereas amino acid or ketone-body–related metabolites predominate in adaptive local types. Previous studies have reported sorbitol and glucose derivaties as energy indicators (Islamiyati et al., 2025), glutamic acid as a protein catabolic process marker (Stolarz and Hanaka, 2025), and ketone bodies such as 3-hydroxybutyric acid as lipid mobilization and metabolic imbalance markers (Zhou et al., 2025). Thus, the dual metabolic and biochemical evidence here mechanistically explains how different breeds affect energy homeostasis and production efficiency under different nutritional challenges. Although some metabolites showed a perfect AUC (1.0), we should interpret these results cautiously because of the small sample size (n=3/bree). The limited number of animals in the present study might have caused an overestimation for predictive accuracy; therefore, evaluation of these biomarker patterns on a larger independent population is needed to confirm their diagnostic strength and general relevance. Correlation (Fig. 7; Table 4) also proved the clinical significance of these metabolites by correlation with routine biochemistry. Plasma glucose also showed significant positive associations with sorbitol, D-glucose, 2,3-butanediol, and D-xylose, supporting the theory between systemic glucose status and carbohydrate metabolism, as previously reported in bovine metabolomic analysis. This connection provides an additional biological validation of the metabolic pathways found, especially in terms of the relationship between energy metabolism and carbohydrate metabolism: with a report on such relatedness. Furthermore, the positive associations between glucose and these metabolites are congruent with glucose providing energy requirements, especially in dairy-type breeds. L-alanine was positively correlated with albumin and negatively associated with cholesterol, indicating a trade-off between protein turnover and lipid metabolism and agreeing with the metabolic flexibility of dietary stress (Zhao X Wang et al., 2024). This finding narrows amino acid metabolism to an important physiological theme, i.e., the adaptation of PROT to fluctuating nutritional conditions. Glutamic acid was positively related to creatinine in Etawa goats, reflecting the existence of interrelationships between amino acid metabolism and muscle turnover, while glycerol and 3-hydroxybutyric acid were positively correlated with albumin but negatively correlated with cholesterol; these were further explained by lipid mobilization and protein availability. These relationships offer further evidence that amino acids and lipids are metabolically regulated to optimize energy balance (or minimize energy deficits) PoPt]!n syntheSiS. The results may also replicate previous observations that metabolomic-biochemical correlations could reveal integrative metabolic relations that single parameter analyses cannot (Wu et al., 2024; Daddam et al., 2025). They showed that the discovered metabolic pathways are of biological significance and have direct relations with traditional levels of protein (glucose, albumin, and cholesterol). Metabolomics analyses illuminate the potential to unlock complex nutrition-related and metabolic areas. These findings have important implications for future precision goat nutrition practices. By defining breed-specific metabolic phenotypes (Figs. 1–6, Tables 1–4) and correlations with blood biochemical markers (Fig. 7 and Table 4), the current study contributes to the design of feeding strategies that are adapted to genetic background to achieve optimal feed efficiency by reducing nutrient losses and reducing the risk of metabolic disorders. Such approaches are particularly important for tropical production systems because feed resources in these areas are constrained and environmental challenges are relatively intense. These observations are in concordance with previous metabolomic studies revealing that carbohydrate and amino acid metabolic pathways contribute significantly to nutrient utilization and energy homeostasis in ruminants (Xue et al., 2020; Polizel et al., 2025), implying that these metabolites can be used as potential targets for precision feeding. Conversely, the modest sample size and single-institution nature of this study may limit generalizability. Future research should confirm these biomarkers in larger, diverse cohorts, assess their responsiveness to different feeding strategies, and combine metabolomic data with genomic or proteomic strategies. Knowledge of integrating information gaps will improve confidence in identifying biomarkers and promote the development of metabolomics-based management approaches for sustainable goat farming. Nevertheless, the small sample size and single institution of the current study were important limitations that reduce the generalizability of these findings. The identified biomarkers need to be validated in further studies with larger and more mixed cohorts to confirm their stability under different nutritional and management regimens. ConclusionIn summary, the findings show that metabolomic fingerprinting displays clear breed-specific signatures, although traditional biochemical markers provide a modest degree of discrimination between goat breeds. In Saanen goats, carbohydrate-related metabolites were dominant amino acid and lipid intermediates in Etawa goats and L-alanine in Kacang goats. Galactose and amino acid metabolism were the most significant related pathways, and several metabolites stood out as promising candidate biomarkers. These metabolites were further validated by relationships with traditional biochemical indices, suggesting that the highlighted metabolites have functional importance and might be exploited to develop precision nutrition strategies for improving goat production sustainability. However, these findings are preliminary and derived from a small experimental cohort, implying that further validation is crucial. Future research should focus on validating the identified biomarkers in larger, more diverse goat populations, testing their responsiveness to varying diets, and exploring the genetic basis of these biomarkers to strengthen these results. Such steps are essential for translating these findings into practical applications that can improve livestock management and feed efficiency in sustainable goat production systems. AcknowledgmentsThe authors gratefully acknowledge funding from the Institute for Research and Community Service, Hasanuddin University (LPPM Unhas), the Thematic Research Group on Local Livestock Genetic Resources and the Faculty of Animal Sciences, Hasanuddin University, and the collaboration of the National Research and Innovation Agency (BRIN) (Credit Number: 01260/UN4.22/PT.01.03/2025). Conflict of interestThe authors declare no conflict of interest. FundingThis research was funded by the Institute for Research and Community Service, Hasanuddin University (LPPM Unhas), under Contract Number 01260/UN4.22/PT.01.03/2025. Authors’ contributionsI.U.A., A.M.D., A.N.C., and M.I.A.D. supervised the experiment, conception, and design of the study. I.U.A., A.M.D., A.N.C., F.I., and E.D. contributed to conception and design of the study and improvement of the manuscript. I.U.A., A.M.D., F.I., M.I.A.D., and M.M. collected data and drafted the manuscript. R.F., T.M., A.N., and S.N. drafted the manuscript and performed data analysis. Data availabilityAll data supporting this study’s findings are available within the manuscript. ReferencesAdetunji, A.O., Price, J., Owusu, H., Adewale, E.F., Adesina, P.A., Saliu, T.P., Zhu, Z., Xedzro, C., Asiamah, E. and Islam, S. 2025. Mechanisms by which phytogenic extracts enhance livestock reproductive health: current insights and future directions. Front. Vet. Sci. 12, 1568577; doi:10.3389/fvets.2025.1568577 Aziz, I.U., Astuti, A., Agus, A., Bintara, S. and Zainuddin, A. 2023. Effect of nutrient improvement and mineral premix supplementation on weight, average daily gain, and metabolite profile of repeat breeder cows. IOP Conf. Series: Earth Environ. Sci. 1183(1), 12–013; doi:10.1088/1755-1315/1183/1/012013 Byrne, L. and Murphy, R.A. 2022. Relative bioavailability of trace minerals in production animal nutrition: a review. Animals 12(15), 1981; doi:10.3390/ani12151981 Cooke, A.S., Machekano, H., Ventura-Cordero, J., Louro-Lopez, A., Joseph, V., Gwiriri, L.C., Takahashi, T., Morgan, E.R., Lee, M.R.F. and Nyamukondiwa, C. 2024. Opportunities to improve goat production and food security in Botswana through forage nutrition and the use of supplemental feeds. Food Secur. 16(3), 607–622; doi:10.1007/s12571-024-01452-1 Daddam, J.R., Sura, M., Sarmikasoglou, E., Ahmad, G., Naughton, S., Mills, M., White, H.M., Vandehaar, M. and Zhou, Z. 2025. Differences in amino acid and fatty acid metabolism contribute to variability in dairy cattle feed efficiency. J. Dairy. Sci. 108(8), 8367–8379; doi:10.3168/jds.2025-26468 Danso, F., Iddrisu, L., Lungu, S.E., Zhou, G. and Ju, X. 2024. Effects of heat stress on goat production and mitigating strategies: a review. Animals 14(12), 1793; doi:10.3390/ani14121793 Deda, O., Armitage, E.G., Mouskeftara, T., Kachrimanidou, M., Zervos, I., Malousi, A., Loftus, N.J., Taitzoglou, I. and Gika, H. 2024. Unraveling cecal alterations in Clostridioides difficile colonized mice through comprehensive metabolic profiling. J. Proteome. Res. 23(12), 5462–5475; doi:10.1021/acs.jproteome.4c00578 Ediriweera, M.K. 2025. Analyzing the fatty acid composition of goat milk and its HDAC inhibitory effects: a rational approach towards understanding the epigenetic impacts of goat milk. Nat. Prod. Res. 39(6), 1714–1717; doi:10.1080/14786419.2023.2298379 Ghavipanje, N., Fathi Nasri, M.H., Farhangfar, S.H., Ghiasi, S.E. and Vargas-Bello-Pérez, E. 2021. Regulation of nutritional metabolism in transition dairy goats: energy balance, liver activity, and insulin resistance in response to berberine supplementation. Animals 11(8), 2236; doi:10.3390/ani11082236 Ginting, S.P., Tarigan, A. and Simanuhuruk, K.A.S. 2020. Effects of two different energy sources in total mixed diets on the performances and blood metabolites of lactating Boerka goats. J. Ilmu. Ternak. Vet. 25(1), 26–33; doi: 10.14334/jitv.v25i1.2196 Godber, O.F., Chentouf, M. and Wall, R. 2020. Sustainable goat production: modelling optimal performance in extensive systems. Anim. Prod. Sci. 60(6), 843–851; doi:10.1071/AN18481 Gómez, E., Salvetti, P., Gatien, J., Muñoz, M., Martín-González, D., Carrocera, S. and Goyache, F. 2020. Metabolomic profiling of Bos taurus beef, dairy, and crossbred cattle: a between-breeds meta-analysis. J. Agric. Food. Chem. 68(32), 8732–8743; doi:10.1021/acs.jafc.0c02129 Huang, Y., Kong, Y., Shen, B., Li, B., Loor, J.J., Tan, P., Wei, B., Mei, L., Zhang, Z., Zhao, C., Zhu, X., Qi, S. and Wang, J. 2023. Untargeted metabolomics and lipidomics to assess plasma metabolite changes in dairy goats with subclinical hyperketonemia. J. Dairy. Sci. 106(5), 3692–3705; doi:10.3168/jds.2022-22812 Islamiyati, R., Azis, I. U., Amal, I., Bahar, M. R., Sabil, S., Santoso, S., Khan, F. A., Nurlatifah, A., Diansyah, A. M., Irawan, F., and Damayanti, E. 2025. Integrative metabolomics and hormonal profiling reveal biomarkers of milk yield efficiency in Sapera dairy goats under tropical conditions. Vet. World, 18(11), 3594–3606; doi: 10.14202/vetworld.2025.3594-3606 Irawan, A., Hartatik, T., Bintara, S., Astuti, A. and Kustantinah, K. 2024. Nutrient digestibility, N balance, performance, and blood parameters of Kacang goats differing in GDF9 genotype fed different sources of dietary fiber. Trop. Anim. Sci. J. 47(1), 33–41; doi:10.5398/tasj.2024.47.1.33 Kaniamuthan, S., Manimaran, A., Kumaresan, A., Wankhade, P.R., Karuthadurai, T., Sivaram, M. and Rajendran, D. 2025. Biochemical indicators of energy balance in blood and other secretions of dairy cattle: a review. Agric. Rev. 46(2), 247-255. Kapusniaková, M. 2023. Alimentary metabolic disorders in high-producing dairy cows: a review. Acta Fytotech. Zootech. 26(4), 354–359; doi:10.15414/afz.2023.26.04.354-359. Kuru, M., Akyüz, E. and Makav, M. 2022. Some metabolic profile markers in goats. Turk. J. Vet. Intern. Med. 1(2), 32–39; doi:10.5281/zenodo.7486065 Liu, Z., Jiang, A., Lv, X., Fan, D., Chen, Q., Wu, Y., Zhou, C. and Tan, Z. 2024. Combined metabolomics and biochemical analyses of serum and milk revealed parity-related metabolic differences in Sanhe dairy cattle. Metabolites 14(4), 227; doi:10.3390/metabo14040227 Michalk, D.L., Kemp, D.R., Badgery, W.B., Wu, J., Zhang, Y. and Thomassin, P.J. 2019. Sustainability and future food security—a global perspective for livestock production. Land. Degrad. Dev. 30(5), 561–573; doi:10.1002/ldr.3217 Nunes, A.T., Faleiros, C.A., Poleti, M.D., Novais, F.J., López-Hernández, Y., Mandal, R., Wishart, D.S. and Fukumasu, H. 2024. Unraveling ruminant feed efficiency through metabolomics: a systematic review. Metabolites 14(12), 675; doi:10.3390/metabo14120675 Nurlatifah, A., Khotijah, L., Arifiantini, R.I., Maidin, M.S. and Astuti, D.A. 2022. Change in hematology prepartum and postpartum of Garut ewe fed with flushing diet containing lemuru oil. IOP Conf. Ser. Earth Environ. Sci. 1020(1), 12–005; doi:10.1088/1755-1315/1020/1/012005 Polizel, G.H.G., Diniz, W.J.S., Cesar, A.S.M., Ramírez-Zamudio, G.D., Cánovas, A., Dias, E.F.F., Fernandes, A.C., Prati, B.C.T., Furlan, E., Pombo, G.D.V. and Santana, M.H.D.A. 2025. Impacts of prenatal nutrition on metabolic pathways in beef cattle: an integrative approach using metabolomics and metagenomics. BMC. Genomics. 26(1), 359; doi:10.1186/s12864-025-11545-6 Rinschen, M.M., Ivanisevic, J., Giera, M. and Siuzdak, G. 2019. Identification of bioactive metabolites using activity metabolomics. Nat. Rev. Mol. Cell. Biol. 20(6), 353–367; doi:10.1038/s41580-019-0108-4 Sonea, C., Gheorghe-Irimia, R.A., Tapaloaga, D., Gurau, M.R., Udrea, L. and Tapaloaga, P.R. 2023. Optimizing animal nutrition and sustainability through precision feeding: a mini review of emerging strategies and technologies. Ann. Valahia Univ. Târgovişte Agric. 15(2), 7-11. doi: 10.2478/AGR-2023-0011 Stolarz, M., and Hanaka, A. 2025. Glutamate and its role in the metabolism of plants and animals. Processes, 13(7), 2084; doi: 10.3390/pr13072084 Teixeira, I.A.M.A., Härter, C.J., Vargas, J.A.C., Souza, A.P. and Fernandes, M.H.M.R. 2024. Review: update of nutritional requirements of goats for growth and pregnancy in hot environments. Animal 18(Suppl 2), 101219; doi:10.1016/j.animal.2024.101219 Wu, G., Bazer, F.W., Johnson, G.A., Satterfield, M.C. and Washburn, S.E. 2024. Metabolism and nutrition of L-glutamate and L-glutamine in ruminants. Animals 14(12), 1788; doi:10.3390/ani14121788 Xue, M.Y., Sun, H.Z., Wu, X.H., Liu, J.X. and Guan, L.L. 2020. Multi-omics reveals that the rumen microbiome and its metabolome together with the host metabolome contribute to individualized dairy cow performance. Microbiome 8(1), 64; doi:10.1186/s40168-020-00819-8 Zamuner, F., Cameron, A.W.N., Carpenter, E.K., Leury, B.J. and Digiacomo, K. 2020. Endocrine and metabolic responses to glucose, insulin, and adrenocorticotropin infusions in early-lactation dairy goats of high and low milk yield. J. Dairy. Sci. 103(12), 12045–12058; doi:10.3168/jds.2020-18625 Zhang, H., Tong, J., Zhang, Y., Xiong, B. and Jiang, L. 2020. Metabolomics reveals potential biomarkers in the rumen fluid of dairy cows with different levels of milk production. Asian-Aust. J. Anim. Sci. 33(1), 79–90; doi:10.5713/ajas.19.0214 Zhao, X., Wang, Y., Wang, L., Sun, S., Li, C., Zhang, X., Chen, L. and Tian, Y. 2024. Differences in serum glucose and lipid metabolism, immune parameters, and blood metabolomics in transition cows during the antepartum and postpartum periods. Front. Vet. Sci. 11, 1347585; doi:10.3389/fvets.2024.1347585 Zhou, Z., Yong, K., Luo, Z., Du, Z., Zhou, T., Li, X., Yao, X., Shen, L., Yu, S., Huang, Y. and Cao, S. 2025. Positive regulatory effect of DBT on lipid metabolism in postpartum dairy cows. Metabolites 15(1), 58; doi:10.3390/metabo15010058.a | ||
| How to Cite this Article |
| Pubmed Style Azis IU, Diansyah AM, Churriyah AN, Dagong MIA, Irawan F, Damayanti E, Masturi M, Fidriyanto R, Maulana T, Nurlatifah A, Nurhaliza S. Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Vet. J.. 2026; 16(2): 847-860. doi:10.5455/OVJ.2026.v16.i2.9 Web Style Azis IU, Diansyah AM, Churriyah AN, Dagong MIA, Irawan F, Damayanti E, Masturi M, Fidriyanto R, Maulana T, Nurlatifah A, Nurhaliza S. Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. https://www.openveterinaryjournal.com/?mno=291936 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.9 AMA (American Medical Association) Style Azis IU, Diansyah AM, Churriyah AN, Dagong MIA, Irawan F, Damayanti E, Masturi M, Fidriyanto R, Maulana T, Nurlatifah A, Nurhaliza S. Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Vet. J.. 2026; 16(2): 847-860. doi:10.5455/OVJ.2026.v16.i2.9 Vancouver/ICMJE Style Azis IU, Diansyah AM, Churriyah AN, Dagong MIA, Irawan F, Damayanti E, Masturi M, Fidriyanto R, Maulana T, Nurlatifah A, Nurhaliza S. Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 847-860. doi:10.5455/OVJ.2026.v16.i2.9 Harvard Style Azis, I. U., Diansyah, . A. M., Churriyah, . A. N., Dagong, . M. I. A., Irawan, . F., Damayanti, . E., Masturi, . M., Fidriyanto, . R., Maulana, . T., Nurlatifah, . A. & Nurhaliza, . S. (2026) Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Vet. J., 16 (2), 847-860. doi:10.5455/OVJ.2026.v16.i2.9 Turabian Style Azis, Ismah Ulfiyah, Athhar Manabi Diansyah, Andi Nimahtul Churriyah, Muhammad Ihsan Andi Dagong, Fahrul Irawan, Erni Damayanti, Masturi Masturi, Rusli Fidriyanto, Tulus Maulana, Aeni Nurlatifah, and Sitti Nurhaliza. 2026. Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Veterinary Journal, 16 (2), 847-860. doi:10.5455/OVJ.2026.v16.i2.9 Chicago Style Azis, Ismah Ulfiyah, Athhar Manabi Diansyah, Andi Nimahtul Churriyah, Muhammad Ihsan Andi Dagong, Fahrul Irawan, Erni Damayanti, Masturi Masturi, Rusli Fidriyanto, Tulus Maulana, Aeni Nurlatifah, and Sitti Nurhaliza. "Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition." Open Veterinary Journal 16 (2026), 847-860. doi:10.5455/OVJ.2026.v16.i2.9 MLA (The Modern Language Association) Style Azis, Ismah Ulfiyah, Athhar Manabi Diansyah, Andi Nimahtul Churriyah, Muhammad Ihsan Andi Dagong, Fahrul Irawan, Erni Damayanti, Masturi Masturi, Rusli Fidriyanto, Tulus Maulana, Aeni Nurlatifah, and Sitti Nurhaliza. "Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition." Open Veterinary Journal 16.2 (2026), 847-860. Print. doi:10.5455/OVJ.2026.v16.i2.9 APA (American Psychological Association) Style Azis, I. U., Diansyah, . A. M., Churriyah, . A. N., Dagong, . M. I. A., Irawan, . F., Damayanti, . E., Masturi, . M., Fidriyanto, . R., Maulana, . T., Nurlatifah, . A. & Nurhaliza, . S. (2026) Breed-specific plasma metabolomic profiles of tropical goats as a basis for precision nutrition. Open Veterinary Journal, 16 (2), 847-860. doi:10.5455/OVJ.2026.v16.i2.9 |