| Research Article | ||
Open Vet. J.. 2026; 16(1): 478-491 Open Veterinary Journal, (2026), Vol. 16(1): 478-491 Research Article Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic productsRumiyati Rumiyati1, Miftiana Nugraha Sari1, Hazza’ Hammam Nawwaruddin1, Marlyn Dian Laksitorini1 and Abdul Rohman1,2*1Faculty of Pharmacy, Universitas Gadjah Mada, Yogyakarta, Indonesia 2Center of Excellence, Institute for Halal Industry and System, Universitas Gadjah Mada, Yogyakarta, Indonesia *Corresponding Author: Abdul Rohman. Faculty of Pharmacy and Center of Excellence, Institute for Halal Industry and System, Universitas Gadjah Mada, Yogyakarta, Indonesia. Email: abdul_kimfar [at] ugm.ac.id Submitted: 26/05/2025 Revised: 11/11/2025 Accepted: 11/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
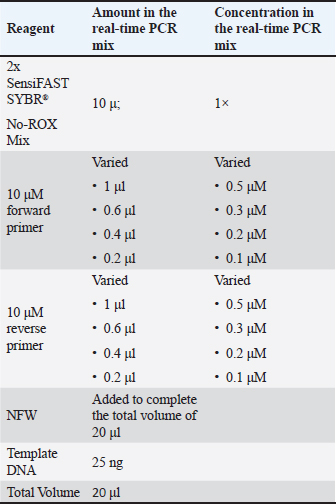
AbstractBackground: Processed food and cosmetic products are integral to modern life, and technological advances have enabled the creation of increasingly complex formulations. Many of these products contain animal-derived ingredients, such as gelatin, whose source must be verified due to religious, health, and ethical concerns. Porcine is one of the primary sources of gelatin production, which has become a major concern. Its detection in complex matrices, such as processed food and cosmetics, is challenging, as gelatin itself undergoes extensive production processes. Real-time polymerase chain reaction (PCR) is widely recognized as one of the analytical methods of choice for species determination; however, its application to gelatin source identification in complex samples remains limited. Aim: This study aimed to develop and optimize a real-time PCR protocol for identifying porcine gelatin contamination in highly processed matrix samples (processed foods and cosmetics) for halal authentication. Methods: This study was established through the following steps: DNA extraction, post-isolation DNA analysis, annealing temperature and primer concentration optimization, specificity assay, amplification efficiency trial, sensitivity test, repeatability examination, and marketed sample analysis. Results: The developed method demonstrated good specificity under optimized conditions (annealing temperature of 58.4°C and primer concentration of 0.2 µM). It achieved a good amplification efficiency of 101.2% with an R² of 0.994. Further validation results showed that the real-time PCR technique had a limit of detection of 1,316 pg in the sensitivity examination and a satisfying coefficient of variation of 0.81% in the repeatability testing. The marketed sample analysis (utilizing five cosmetic samples in the form of facial masks and five food samples in the form of one food additive gelatin powder, two marshmallow products, and two gummy candy products) showed that all of the samples displayed no amplification and were thus considered not to contain porcine DNA, consistent with the manufacturers’ labels. Conclusion: The real-time PCR method meets the validation criteria for qualitative analysis, including specificity, amplification efficiency, sensitivity, and repeatability. This method provides a valuable foundation for advancing halal authentication, particularly for detecting porcine gelatin contamination in processed food and cosmetic products. Keywords: Cosmetic, Halal, Processed food, Real-time PCR. IntroductionIn modern society, processed food and cosmetic products are essential parts of human life. The importance of processed foods and cosmetics in social life is reflected in the industry’s recent growth and is expected to increase even further in the future (Berg et al., 2023). The advancement of technology plays an important role in this growth, resulting in a greater variety of products with increasingly complex compositions (Martínez Steele et al., 2016; Rico et al., 2023). Alongside this advancement, consumers’ demand for transparency in product composition has also increased. One of these transparency demands concerns the use of animal-derived components, as certain consumer populations are restricted from using products containing materials derived from specific animals due to religious beliefs, health considerations, or ethical preferences (Karim and Bhat, 2008; Karim and Bhat, 2009; Alves et al., 2017; Briliana and Mursito, 2017). One of the animal-derived components commonly utilized in the cosmetic and food industries is gelatin, due to its unique properties, which result in a wide range of applications, such as foaming, stabilizing, thickening, gelling, emulsifying, binding, and moisture retention (Uddin et al., 2021). This popular biopolymer is mainly produced from porcine skin, contributing approximately 46% of global gelatin production, followed by bovine hides at 29.4%, as well as porcine and bovine bones at 23.1% (Duconseille et al., 2015). The extensive application of gelatin in the cosmetic and food industries, along with the predominance of porcine as the main source of gelatin production, makes the determination of gelatin sources in cosmetic and food products critical, since Muslim and Jewish dietary laws prohibit the use of products containing porcine derivatives (Akbarzadegan et al., 2020; Hassan et al., 2024). Therefore, developing an analytical method protocol with good sensitivity and specificity to analyze the pork gelatin content in these samples is necessary. The determination of gelatin sources in cosmetic and processed food matrices is quite challenging because these matrices are highly complex due to their variety of forms and the presence of many distinct compounds with specific characteristics (Elgadir et al., 2013; Naharros-Molinero et al., 2024). Moreover, research on the development of analytical methods’ protocols for gelatin source authentication in cosmetic and processed food matrices is still limited and has not been extensively explored. Nonetheless, several analytical methods have been used to determine gelatin sources in various other highly complex matrix products, namely liquid chromatography–mass spectrometry (LC-MS) and liquid chromatography–tandem mass spectrometry (LC-MS/MS) (Guo et al., 2018; Chia et al., 2020; Sha et al., 2020; Zhu et al., 2023), enzyme-linked immunosorbent assay (ELISA) (Venien and Levieux, 2005; Doi et al., 2009; Tukiran et al., 2016a), and polymerase chain reaction (PCR) (Cai et al., 2012; Mohamad et al., 2018). Despite having high resolution that allows for precise identification and making them appropriate for analyzing extremely complex matrices, the LC-MS and LC-MS/MS methods have several disadvantages, including high execution costs, time-consuming procedures, the requirement for specialized knowledge, and limited accessibility in laboratories (Deng et al., 2020; Rohman et al., 2020; Rico et al., 2023). Consequently, these methods are unsuitable for routine analysis in the authentication of gelatin sources. ELISA is a protein-based method well known for its low cost, specificity, sensitivity, and accuracy in identifying antigenic proteins (Asensio et al., 2008; El Sheikha et al., 2017; Nhari et al., 2019; Chia et al., 2020). Unfortunately, this approach has limitations in identifying complex matrices, as accompanying components may interfere with antigen detection, reduce sensitivity, and cause false antibody cross-reactions (Venien and Levieux, 2005; Doi et al., 2009; Tukiran et al., 2016a). These factors demonstrate that using this method to identify gelatin sources in various matrix samples for halal authentication is quite challenging. Nucleic acid-based techniques, such as PCR, are renowned for their selectivity and sensitivity. DNA is distinct and species-specific; therefore, this technique is commonly used to identify species by detecting animal DNA present in samples (Rohman et al., 2020). Moreover, this procedure is fast and easy to perform because PCR supporting kits are readily available. Several studies have utilized various PCR approaches for gelatin analysis in various matrix samples, such as commercial gelatin, gelatin mixtures, and capsules (Cai et al., 2012; Shabani et al., 2015; Sudjadi et al., 2016). However, there has not been much research done on using this technique for halal verification to identify the sources of gelatin in processed food and cosmetic matrices. Since these assurances follow zero-tolerance policies, halal and kosher are included as qualitative studies, meaning that the presence of porcine gelatin is prohibited in samples, even at trace levels (Ab Latiff, 2020; Akbarzadegan et al., 2020; Herdiana et al., 2024). For such a qualitative study, real-time PCR is preferred over conventional PCR because it does not require a post-amplification procedure, making the analysis less time-consuming and labor-intensive. Moreover, real-time PCR has superior sensitivity compared with conventional PCR (Ramamurthy et al., 2011; Kokkinos et al., 2014; Xia et al., 2018). This advantage is highly beneficial, especially for identifying small amounts of DNA present in low concentrations of gelatin excipient within a highly complex matrix, as the DNA in gelatin excipients has undergone extensive degradation due to the high pressure and temperature involved in gelatin production, which can destroy most of the DNA (Khayyira et al., 2018; Chia et al., 2020; Yang et al., 2020; Zhu et al., 2023). Furthermore, using SYBR Green as the fluorescence signal reporter offers several advantages over the TaqMan probe in real-time PCR analysis, as SYBR Green is inexpensive and is therefore more suitable for routine analysis. Additionally, several studies have shown that SYBR Green-based real-time PCR systems can provide comparable specificity and performance to probe-based real-time PCR systems (Maeda et al., 2003; Tajadini et al., 2014). In this study, we aim to optimize and validate a simple and affordable real-time PCR analytical protocol using SYBR Green fluorescent dye for detecting pig-derived gelatin in processed food and cosmetic products. Materials and MethodsThe porcine and bovine gelatin standards were purchased from Sigma AldrichTM (Product numbers: G2500 and G9382, respectively). Raw meat samples, namely pork, beef, goat, chicken, tuna, and catfish (Clarias sp. and Pangasius sp.), were obtained from local markets in Yogyakarta, Indonesia. These species were selected because they are widely consumed and used in the local community, and porcine, bovine, fish, and poultry are among the major commercial gelatin sources. The primers used in this study refer to a porcine-specific primer that was developed by Wardani et al. (2015) and is routinely used for the analysis of pork contamination in food, targeting the D-Loop 108 gene (forward: 5’-CGT ATG CAA ACC AAA ACG CCA-3’; reverse: 5’-GCT TAT ATG CAT GGG GAC TAG C-3’). Five cosmetic samples (in the form of facial masks) and five food samples (one food additive gelatin powder, two marshmallow products, and two gummy candy products) containing gelatin were purchased from e-commerce platforms and local markets in Yogyakarta, Indonesia. Commercial samples were selected based on their declared gelatin content on the labels and their easy accessibility to the public. These samples were used to confirm the gelatin source in widely circulated products within the community. DNA isolationDNA was isolated from gelatin standards, raw meat samples, and commercially available cosmetic and food samples using the FavorPrep™ Tissue Genomic DNA Extraction Mini Kit (Favorgen, Taiwan) according to the manufacturer’s instructions. The kit employs a silica matrix-based procedure involving lysis, binding, washing, and elution. Approximately 25 mg of the raw meat was transferred to a microcentrifuge tube and ground. For gelatin and commercial samples, 50 mg of material was used to obtain a higher DNA yield, as preliminary experiments showed that 25 mg produced insufficient DNA concentrations for these matrices (data not shown). The samples were mixed with 200 µl FATG1 buffer and 20 µl proteinase K, then incubated at 60°C for 1–3 hours. Next, 200 µl FATG2 buffer was added, followed by incubation at 70°C for 10 minutes. Ethanol (200 µl, 96%–100%) was added, and the mixture was pulse-vortexed. Then, after briefly centrifuging to remove droplets from the tube lid, the mixture (including any precipitate) was transferred to a FATG Mini Column placed in a collection tube. The column was centrifuged at 12,500 rpm for 1 minute, transferred to a new collection tube, washed with 400 µl W1 buffer, centrifuged (12,500 rpm, 1 minute), and the flow-through discarded. A second wash was performed using 750 µl wash buffer under the same conditions. Then, the column was centrifuged at full speed for 3 minutes to dry the membrane. DNA was eluted with 100 µl preheated elution buffer, left to stand for 3 minutes, and centrifuged at 12,500 rpm for 2 minutes. DNA concentration and purity were determined at 260 and 280 nm using a NANO-Quant SPARK TECANTM spectrophotometer (Switzerland). Real-time PCR optimization and analysisReal-time PCR conditions were optimized by applying a primer annealing temperature gradient and varying primer concentrations in the reaction mix. In this study, a temperature gradient ranging from 52.0°C to 62.0°C (62.0°C, 61.4°C, 60.3°C, 58.4°C, 55.9°C, 54.0°C, 52.7°C, and 52.0°C) was used to amplify porcine gelatin standards in order to identify the optimal annealing temperature for the real-time PCR protocol. This eight-point temperature gradient was determined based on the calculated annealing temperature (Ta) from the “Tm Calculator for Primers” available at www.thermofisher.com. The gradient starts at a temperature 6°C–10°C lower than the Ta predicted by the calculator—in this case, 58.9°C—and is increased stepwise. The gradient approach allowed further empirical optimization of the annealing temperature for each combination of template–primer pairs. The optimum annealing temperature candidates obtained from this step were subsequently evaluated in a specificity test. Primer concentration optimization was performed using real-time PCR mixes as described in Table 1. According to the manufacturer’s protocol for the SensiFAST™ SYBR No-ROX Mix, the optimal concentration should be determined by titrating in the range of 0.1–1 μM, with the forward and reverse primers at equimolar concentrations. In this study, we tested primer concentrations of 0.5, 0.3, 0.2, and 0.1 µM for optimization. The temperature program was set based on the optimum annealing temperature determined from the preceding annealing temperature optimization and specificity tests. Table 1. Real-time PCR mixes for primer concentration optimization.
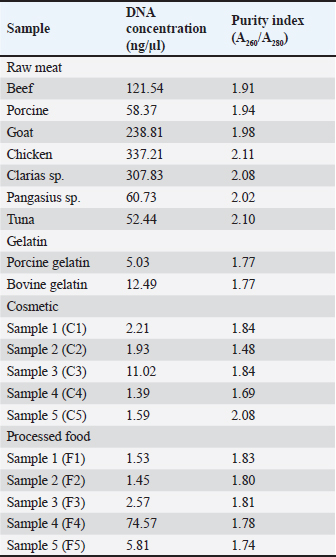
Real-time PCR was performed using a CFX96 system (Bio-Rad, USA). Reactions were prepared according to the SensiFAST™ SYBR No-ROX Mix (Meridian Bioscience, USA) manufacturer’s guidelines, containing 10 µl of 2× mix, 25 ng of template DNA, equimolar forward and reverse primers at concentrations of 0.5, 0.3, 0.2, or 0.1 µM (for optimization), and nuclease-free water to a final volume of 20 µl (Table 1). The cycling conditions followed the SensiFAST™ manufacturer’s guidelines: a polymerase activation step at 95°C for 3 minutes, a denaturation step at 95°C for 5 seconds, an annealing step for 10 seconds at the optimal annealing temperature (varied from 52°C to 62°C to determine the optimum annealing temperature), and an extension step at 72°C for 20 seconds. The melting curve analysis was performed using the default program provided by the instrument: the melt curve temperature was set from 65°C to 95°C, increasing gradually by 0.5°C every 5 seconds. Real-time PCR validation and analysis of marketed samplesThe developed method underwent qualitative PCR validation, including specificity, amplification efficiency, sensitivity, and repeatability assessments (Rohman et al., 2022). Specificity was evaluated using DNA from two sources: raw meat and gelatin standards. For raw meat isolates, pork DNA served as the target, whereas beef, goat, chicken, Pangasius sp., tuna, Clarias sp., and bovine gelatin were non-targets. For gelatin isolates, porcine and bovine gelatin were the target and non-target, respectively. A no-template control (NTC) was used to monitor amplification. Amplification efficiency, coefficient of determination (R²), and y-intercept were determined via linear regression between the logarithm of serially diluted DNA concentrations (x-axis) and corresponding quantification cycle (Cq) values (y-axis). Five concentrations of porcine gelatin DNA were used as templates: 10,000, 6,666, 4,444, 1,975, and 1,316 pg/µl. Sensitivity was assessed using a serial dilution series (10,000; 6,666; 4,444; 2,962; 1,975; 1,316; 878 pg/µl) to determine the limit of detection (LOD), defined as the lowest DNA concentration consistently amplified and detected. The repeatability was evaluated by calculating the relative standard deviation of the Cq values from six replicates of the target DNA. Subsequently, the validated method was applied to detect porcine gelatin DNA in commercial cosmetic and processed food products containing gelatin. Ethical approvalNot applicable to this study. No human or animal subjects were involved. ResultsDNA isolationSignificant differences in DNA concentration were observed between isolates sourced from raw meat samples and gelatin powder (commercial gelatin) or gelatin-containing samples (cosmetic and processed food) (Table 2). For the isolates from raw meat samples, the DNA concentration ranges from 52.44 to 337.21 ng/µl, whereas the majority of the results for the gelatin powder or gelatin-containing samples show that the DNA concentration is below 10 ng/µl. However, the purity index assessment showed a broad variance among all samples, with ratios ranging from 1.48 to 2.11. Table 2. The DNA concentration and purity index of all samples, including raw meats, gelatins, cosmetics, and processed foods.
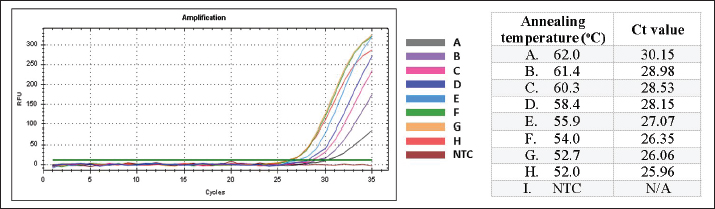
Real-time PCR optimization: annealing temperatureThe gradient-based annealing temperature optimization revealed variations in the Cq values (Fig. 1), with higher annealing temperatures generally producing higher Cq values. Among the eight temperatures tested, 52.0°C produced the lowest Cq value (25.96), while 62.0°C produced the highest value (30.15). The annealing temperature of 58.4°C, previously reported by Wardani et al. (2015) to provide good primer specificity, yielded a Cq value of 28.15 in our testing. Based on these results, 52.0°C and 58.4°C were selected as optimum annealing temperature candidates for subsequent specificity analysis to confirm the optimized program conditions.
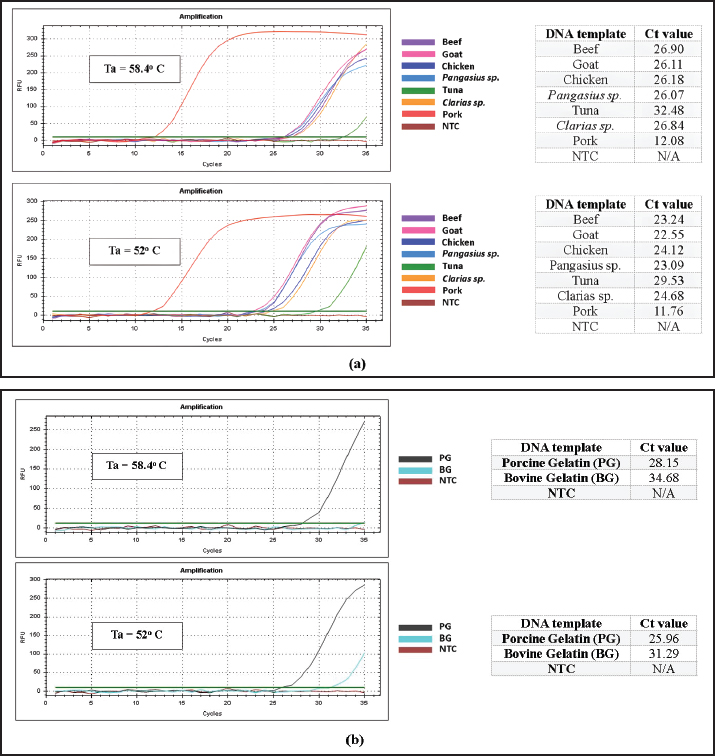
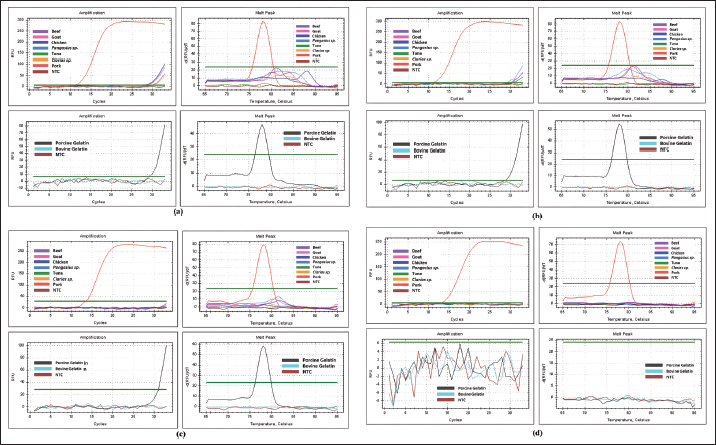
Fig. 1. Results of annealing temperature optimization. A–H represent the annealing temperature levels used in the experiment. The table shows the corresponding Cq values for each annealing temperature. Real-time PCR optimization: specificity test using optimum annealing temperature candidatesThe specificity test using optimum annealing temperature candidates showed that primers with annealing temperatures of 58.4°C and 52.0°C still amplified both target and non-target DNA (Fig. 2). However, the annealing temperature of 58.4°C provided better specificity. In the specificity test using raw meat samples, 58.4°C allowed better discrimination between pork DNA and DNA from other species, given the greater difference in Cq values between target and non-target DNA (pork DNA at a Cq value of 12.08 and chicken, the closest non-target, at a Cq value of 26.18) compared to 52.0°C (pork DNA at a Cq value of 11.76 and goat, the closest non-target, at a Cq value of 22.55) (Fig. 2a). Moreover, the specificity test utilizing porcine gelatin (target) and bovine gelatin (nontarget) also demonstrated that an annealing temperature of 58.4°C offered superior specificity over 52.0°C, as 52.0°C produced a stronger amplification for bovine gelatin, with a lower Cq value of 31.29 compared to 34.68 at 58.4°C (Fig. 2b). Additionally, the target DNA from porcine gelatin exhibited a higher Cq value (28.15) than target DNA from raw pork (12.08).
Fig. 2. Results of the specificity test using the two optimal annealing temperature candidates, 58.4°C and 52°C: (a) raw meat DNA isolates and (b) gelatin DNA isolates as templates. Real-time PCR optimization: primer concentration optimizationFigure 3 presents the primer concentration optimization results. Based on the corresponding amplification curves and melt peak graphs, 0.2 μM exhibited the highest specificity among all tested concentrations, showing strong amplification for target DNAs (DNA from pork and porcine gelatin, at Cq values of 13.66 and 31.66, respectively) with no non-specific amplification (Fig. 3c). At higher concentrations of 0.5 and 0.3 μM (Fig. 3a and b), non-specific responses were still observed, particularly in raw meat DNA isolates, where several non-specific amplifications occurred in later cycles (around Cq=28 or higher). Conversely, at the lowest concentration of 0.1 μM (Fig. 3d), the reaction failed to amplify target DNA from porcine gelatin, as both the amplification and melt peak curves fell below the threshold. Moreover, all successfully amplified target DNAs (from pork or gelatin) exhibited the same melt peak temperature of 78.0°C (Fig. 3a–d). The primer concentration of 0.2 μM was selected as the optimal primer concentration for subsequent analyses.
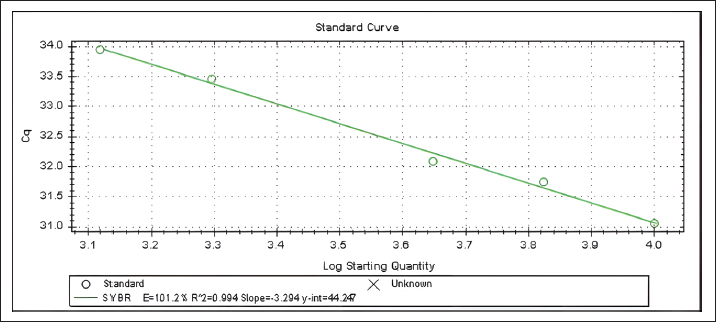
Fig. 3. The specificity test results (amplification curve and melt peak on the left and right sides, respectively) using DNA isolates from raw meat and gelatin DNA isolates (upper and lower graphs, respectively) for primer concentrations of 0.5 µM (a), 0.3 µM (b), 0.2 µM (c), and 0.1 µM (d). Amplification efficiencyFigure 4 shows the standard curve result of the testing (the logarithmic DNA quantity on the x-axis against the Cq value of the respective concentration on the y-axis). The obtained slope is −3.294, with an R² value of 0.994 and a y-intercept of 44.247. Then, the slope was used to calculate the amplification efficiency, which was determined automatically by the real-time PCR instrument, resulting in an efficiency of 101.2%.
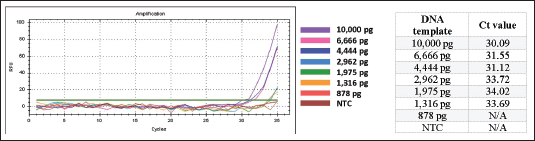
Fig. 4. The linear regression correlating the logarithmic of the DNA quantity (x-axis) and the respective Cq values (y-axis). Sensitivity and repeatability testsSensitivity testing was performed to determine the smallest amount of template DNA that could still be amplified and detected. The findings showed that the established technique achieved a LOD value of 1,316 pg (Fig. 5).
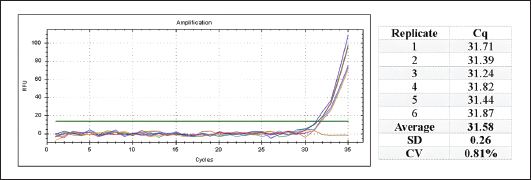
Fig. 5. Amplification curves and Cq values from the sensitivity assay using serially diluted porcine gelatin DNA as the reaction template. The repeatability test was assessed based on the coefficient of variation (CV) of Cq values from six replicates using porcine gelatin-derived DNA as the template. The resulting Cq values are presented in Figure 6. The developed method achieved a low CV value of 0.81% (Fig. 6).
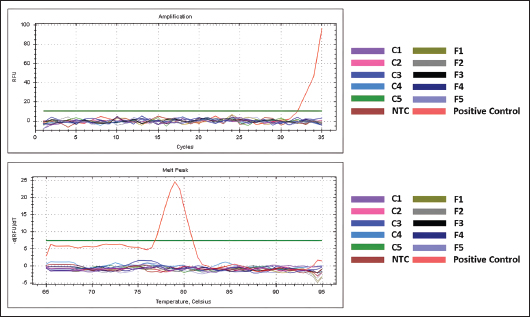
Fig. 6. The repeatability assay result: the amplification curve (left), amplification response parameter (Cq) and the CV value calculation (table). Analysis of the marketed samplesMarketed sample analysis showed amplification only in the positive control, with a Cq value of 32.03, whereas the marketed cosmetic (C) samples, processed food (F) samples, and the NTC exhibited no amplification. The melt-peak curve further confirmed the presence of a single peak corresponding to the positive control (Fig. 7).
Fig. 7. Result of the application of the developed method for identifying commercially available cosmetic (C) and processed food samples (F). The amplification curve (above) and melt peak (below) are shown. DiscussionDNA isolationThe DNA concentrations obtained from gelatin standards, as well as from cosmetic and processed food samples containing gelatin, are significantly lower than those of raw meat samples (Table 2). This result may be due to the high temperature and pressure used in the gelatin production process from the parent protein, collagen, which breaks down DNA into tiny fragments and destroys most of the DNA contained in the sample. Further cosmetic and food production processes, as well as numerous contaminant components present in marketed samples, may contribute to the low yield of DNA extracts in such samples (Khayyira et al., 2018; Chia et al., 2020; Yang et al., 2020; Zhu et al., 2023). Some samples have a purity index greater than 2.0 or lower than 1.8, as shown in Table 2, indicating that several DNA extracts contain RNA or protein contaminants and do not meet the criteria for good purity. Contaminants within DNA isolates must be minimized, especially in quantitative studies, since they may inhibit the PCR reaction (Shen, 2019; Jue et al., 2020; Weng et al., 2020). Nonetheless, the main objective of the post-isolation test in qualitative studies is to ensure that DNA is successfully extracted; therefore, the purity index is not a major concern. The DNA isolates can still be used for further analysis. Real-time PCR optimization: annealing temperatureThe primer annealing temperature optimization aims to determine the optimal binding temperature at which template DNA and primers bond with high efficiency and specificity. Determining the primer binding temperature is crucial because an excessively high annealing temperature may result in insufficient or unsuccessful primer-DNA template hybridization, lowering the PCR output or causing amplification failures. However, if the annealing temperature is not high enough, the primer may amplify the DNA sequence at the wrong site (mispriming), resulting in a non-specific product (Clark et al., 2019). One consideration for determining the optimum binding temperature is the temperature that gives the minimal Cq value (Bustin and Huggett, 2017; Zhao et al., 2021). Hence, a setting of 52.0°C was selected as a potential optimal condition for annealing in the subsequent specificity analysis, with a Cq value of 25.96 (Fig. 1). This early Cq value indicates that the primer binds to the template DNA more efficiently at 52.0°C than at other temperatures. The results (Fig. 1) also show that the Cq value increases as the annealing temperature rises, indicating that a higher annealing temperature makes it more difficult for the primer to bind to the template DNA, resulting in less efficient binding. Furthermore, although 52.0°C provides superior efficiency among the tested temperatures, further specificity analysis is necessary because the optimum annealing temperature must also ensure excellent specificity. The annealing temperature of 58.4°C was also selected as a candidate for the optimum annealing temperature, as the optimization by Wardani et al. (2015) showed that the primer provided good specificity for raw meat samples at this temperature (data not shown). However, in this optimization test using a porcine gelatin standard, this temperature did not exhibit a superior Cq value compared to the others, with a Cq value of 28.15 (Fig. 1). Further specificity tests were performed on the selected annealing temperatures to compare the specificity of these two conditions. Real-time PCR optimization: specificity test using optimum annealing temperature candidatesAlthough the Cq value from the annealing temperature optimization indicated that 52.0°C provided better efficiency (Fig. 1), the specificity examination showed that it did not outperform 58.4°C. At 52.0°C, the Cq discrimination between target and nontarget in raw meat isolates was lower, and nonspecific amplification in gelatin isolates was higher (Fig. 2a, b). We considered 52.0°C too low, as it allowed the primer to bind to incorrect DNA sequences in addition to the target (mispriming), increasing nonspecific amplification. At lower annealing temperatures, partial matching between the primer and the template DNA—despite gaps or mismatches—can remain stable, allowing the amplification of non-target sequences (Malhotra, 1998). On the other hand, an annealing temperature of 58.4°C provided better specificity, as it produced greater differences in Cq values between target and nontarget DNA compared to 52.0°C, along with lower nonspecific DNA amplification (Fig. 2a and b). Increasing the annealing temperature enhances specificity by improving primer annealing stringency, promoting more specific amplification (Malhotra, 1998; He et al., 2009). A higher annealing temperature ensures stronger and more stable primer–template binding, thereby minimizing partial or incorrect attachments. Therefore, between these two annealing temperature candidates, 58.4°C is preferred over 52.0°C, as the optimal temperature should balance both primer binding efficiency and specificity. However, nonspecific amplifications were still observed at 58.4°C, indicating the need for further optimization of real-time PCR. Although higher annealing temperatures can enhance specificity, we did not explore values above 58.4°C. As shown in Figure 1, each increment above this point caused a gradual rise in Cq values and a decline in Relative Fluorescence Unit reflecting reduced amplification. This occurs because excessively high temperatures hinder primer hybridization, lowering attachment efficiency and leading to inadequate amplification (Clark et al., 2019). Consequently, we opted to optimize primer concentration to achieve high specificity while maintaining the annealing temperature at 58.4°C rather than further increasing the annealing temperature. The findings also revealed that target DNA derived from porcine gelatin had a higher Cq value (28.15) than target DNA from raw pork (12.08). This phenomenon could be attributed to the harsh production processes that gelatin undergoes, which involve high temperatures, pressures, or the use of hydrolysis enzymes. These factors contribute to diminished DNA yield, DNA degradation into small fragments, and impurity accumulation during processing, ultimately leading to elevated Cq values (Khayyira et al., 2018; Yang et al., 2020). Yang et al. (2020) confirmed that the Cq values of gelatin increase when the temperature or pressure is increased or prolonged during production. In addition, gelatin products consist of lipids, water, and hydrolyzed collagen combined with saccharides. Components such as polysaccharides can act as PCR inhibitors, leading to higher Cq values (King et al., 2009; Schrader et al., 2012). Real-time PCR optimization: primer concentration optimizationFurther optimization using a range of primer concentrations in the real-time PCR mix was performed to improve specificity, as the preceding annealing temperature optimization alone was insufficient to achieve the required specificity. Primer concentration optimization is essential for each real-time PCR assay, with optimal values determined empirically (Green and Sambrook, 2018). Figure 3 shows the amplification curves and melt peak results for each primer concentration tested (0.5, 0.3, 0.2, and 0.1 μM). These concentration levels were selected based on the SensiFAST SYBR® kit protocol, which recommends titrating within the range of 0.1–1 μM with equimolar forward and reverse primers to determine the optimal concentration. In the previous annealing temperature optimization, a primer concentration of 0.5 μM was used; therefore, in this assay, lower concentrations were evaluated. This approach was adopted because higher primer concentrations can promote mispriming, increasing the likelihood of primers binding to non-specific sites and producing non-specific amplification (Lorenz, 2012). A similar approach to primer concentration optimization for enhancing PCR performance has also been reported by Zhao et al. (2021) and Fatoni et al. (2025). The results indicated that the primer concentration of 0.2 μM provided the highest specificity among the tested levels. The amplification curves showed that only target DNA (derived from pork and porcine gelatin) was amplified, with no evidence of non-specific amplification (Fig. 3c). Correspondingly, melting peak analysis revealed a single peak above the threshold at 78°C for both pork DNA and porcine gelatin DNA, confirming the amplification of the same porcine sequence region (Farrel, 2023). In contrast, higher primer concentrations of 0.5 and 0.3 μM produced non-specific amplification, particularly in raw meat samples, where non-target DNA amplification increased in later cycles (around Cq=28 or higher) (Fig. 3a and b). At the lowest concentration, 0.1 μM, target DNA from porcine gelatin failed to amplify, as indicated by amplification curves and melt peaks falling below the threshold (Fig. 3d). This outcome shows that excessively low primer concentrations can result in minimal or absent target DNA amplification. Consequently, PCR conditions using an annealing temperature of 58.4°C and a primer concentration of 0.2 μM were selected for subsequent validation. Amplification efficiencyFigure 4 shows that the proposed method yielded a slope value of −3.294, an R² value of 0.994, and a y-intercept of 44.247. The amplification efficiency calculation resulted in a value of 101.2%. The amplification efficiency exceeding 100% may be attributed to inaccuracies in the sample and reagent pipetting during preparation. Another cause is the presence of PCR inhibitors in the template DNA, as the previous DNA purity assessment showed that the purity of standard gelatin was below 1.8, indicating the presence of contaminants in the gelatin isolates (Nybo, 2011). However, the obtained efficiency and R² values remained within the acceptable range for real-time PCR qualitative purposes, which require R² ≥ 0.98 and efficiency (E) between 80% and 120% (Broeders et al., 2014). Sensitivity and repeatability testsThe sensitivity test was conducted using seven serially diluted levels of porcine gelatin DNA template (10,000; 6,666; 4,444; 2,962; 1,975; 1,316; and 878 pg/µl). The results demonstrated that the developed real-time PCR technique successfully amplified the porcine gelatin DNA template down to 1,316 pg, with a Cq value of 33.69 (Fig. 5). The LOD for the method was 1,316 pg, as the DNA template at a lower concentration (878 pg) showed no amplification response and did not exceed the threshold limit. This LOD is notably higher than that reported by Kim et al. (2023), who achieved a detection limit of 5 pg for porcine DNA in processed food. However, their sensitivity was determined using raw porcine DNA, whereas our study employed standard gelatin. In raw meat samples, DNA is typically more abundant and less degraded, whereas in gelatin samples, it is often highly degraded, and the presence of an interfering substance or inhibitor makes detection more challenging, and the LOD value tends to be higher. Moreover, the repeatability test results are shown in Figure 6. The obtained CV value was 0.81%, which meets the real-time PCR validation criteria requirement (<25%) (Broeders et al., 2014). The real-time PCR protocol developed in this study met the validation criteria and demonstrated strong potential as an alternative analytical approach for gelatin determination in complex matrices. Unlike LC-MS/MS, which is costly, time-intensive, and requires specialized expertise, factors that pose challenges for routine analysis (Deng et al., 2020; Rohman et al., 2020; Rico et al., 2023), this method offers a more accessible option. ELISA-based detection has been explored in only a limited number of studies for complex matrices such as processed foods and capsule products. Moreover, several reports indicate that ELISA methods often struggle to differentiate gelatin sources due to cross-reactivity with non-target species (Doi et al., 2009; Tukiran et al., 2016a, 2016b; Nawwaruddin et al., 2024). The developed real-time PCR protocol can serve as a foundation for further advancements in halal food authentication, particularly for gelatin determination in cosmetic and processed food samples. Analysis of the marketed samplesThe analysis of marketed samples showed that only the positive control was amplified (Cq=32.03), whereas five cosmetic samples (facial masks), five food samples (one gelatin powder, two marshmallow products, and two gummy candies), and the NTC showed no amplification (Fig. 7). Melting peak analysis further indicated that only the positive control exhibited a peak, whereas all samples and the NTC did not (Fig. 7). These results shows that none of the marketed samples contained porcine DNA, which is consistent with the manufacturers’ labels indicating the use of bovine gelatin. Nevertheless, the absence of matrix-specific validation remains a key limitation of this study. Such validation is crucial for assessing method performance and understanding matrix effects, thereby ensuring accurate positive or negative results in marketed sample analysis (Hedman et al., 2018; Sun et al., 2023). Future studies should incorporate spiking commercial samples with known amounts of porcine gelatin DNA to simulate real contamination scenarios, which would strengthen confidence in the method’s reliability and detection limits (Goris et al., 2009; Kim et al., 2018). This approach would help mitigate the risk of false-negative results due to inefficient DNA extraction, poor DNA quality or quantity, and matrix complexity, including the presence of inhibitors (Al-Shaibany et al., 2022). Moreover, this study had a limited sample size; therefore, future studies should include a greater number and variety of cosmetic and processed food samples to more comprehensively assess the applicability of the method. ConclusionThe optimized annealing temperature (58.4°C) and primer concentration (0.2 µM) provided excellent specificity. The optimized real-time PCR conditions demonstrated good amplification efficiency, an LOD value of 1,316 pg, and excellent results in the repeatability testing. Based on these results, the developed real-time PCR technique meets the real-time PCR validation criteria (specificity, amplification efficiency, sensitivity, and repeatability) for qualitative purposes. The method was then applied to the analysis of gelatin-containing commercially available cosmetics and processed food samples. None of the samples exhibited amplification, indicating the absence of porcine DNA. This finding aligns with the manufacturers’ labeling, which states that the products contain bovine gelatin. This method provides a valuable foundation for advancing halal authentication, particularly for detecting porcine gelatin contamination in processed food and cosmetic products. AcknowledgementsThanks to the Academic Excellence Improvement Program year 2024 (with contract number 6526/UN1.P1/PT.01.03/2024) Universitas Gadjah Mada, Yogyakarta, Indonesia, which has funded this research. The research grant was awarded to Dr. Rumiyati. Conflict of interestThe authors declare that there is no conflict of interest. FundingThis project was funded by the Academic Excellence Improvement Program year 2024 (with contract number 6526/UN1.P1/PT.01.03/2024) Universitas Gadjah Mada, Yogyakarta, Indonesia, and awarded to Dr. Rumiyati. Author’s contributionsRR: designing the research method, conducting the experiments and collecting data, conducting data analysis, and drafting and reviewing the manuscript; MNS: designing the research method, conducting the experiments and collecting data, conducting data analysis, and drafting manuscripts; HHN: conducting the experiments and collecting data, conducting data analysis, and drafting manuscripts; MDL: designing the research method, drafting, and reviewing the manuscript. AR: designing the research method, supervising the experiments and collecting data, performing data analysis, and reviewing the manuscript. Data availabilityAll data supporting this study’s findings are available within the manuscript. ReferencesAb Latiff, J. 2020. Halal certification procedure in Malaysia and Indonesia. Petita. Jurnal. Kajian. Ilmu. Hukum. Dan. Syariah. 5(2), 121–132. doi:10.22373/petita.v5i2.102 Akbarzadegan, R., Ahari, H., Sharifan, A. and Anvar, A.A. 2020. Overview of the studies on authentication of gelatin using Fourier Transform Infrared spectroscopy coupled with chemometrics. Hum. Health Halal Metrics 1(2), 86–93. doi:10.30502/jhhhm.2021.244697.1021 Al-Shaibany, A., Gargouri, H. and Hadj Kacem, H. 2022. Evaluation of DNA extraction protocols and real-time PCR-based methods for efficient investigation of pig traces in foods. Int. Food Res. J. 29(4), 828–842; doi:10.47836/ifrj.29.4.10 Alves, A., Marques, A., Martins, E., Silva, T. and Reis, R. 2017. Cosmetic potential of marine fish skin collagen. Cosmetics 4(4), 4; doi:10.3390/cosmetics4040039 Asensio, L., González, I., García, T. and Martín, R. 2008. Determination of food authenticity by enzyme-linked immunosorbent assay (ELISA). Food Control. 19(1), 1–8; doi: 10.1016/j.foodcont.2007.02.010 Berg, A., Hudson, S., Klitsch Weaver, K., Lesko Pacchia, M. and Amed, I. 2023. The beauty market in 2023: A special State of Fashion report. McKinsey. Available via https://www.mckinsey.com/industries/retail/our-insights/the-beauty-market-in-2023-a-special-state-of-fashion-report. Briliana, V. and Mursito, N. 2017. Exploring antecedents and consequences of Indonesian Muslim youths’ attitude towards halal cosmetic products: a case study in Jakarta. Asia. Pacific. Manage. Rev. 22(4), 176–184; doi:10.1016/j.apmrv.2017.07.012 Broeders, S., Huber, I., Grohmann, L., Berben, G., Taverniers, I., Mazzara, M., Roosens, N. and Morisset, D. 2014. Guidelines for validation of qualitative real-time PCR methods. Trends Food Sci. Technol. 37(2), 115–126; doi:10.1016/j.tifs.2014.03.008 Bustin, S. and Huggett, J. 2017. QPCR primer design revisited. Biomolecular. Detection. Quantification. 14, 19–28; doi:10.1016/j.bdq.2017.11.001 Cai, H., Gu, X., Scanlan, M.S., Ramatlapeng, D.H. and Lively, C.R. 2012. Real-time PCR assays for detection and quantitation of porcine and bovine DNA in gelatin mixtures and gelatin capsules. J. Food. Composition. Anal. 25(1), 83–87; doi:10.1016/j.jfca.2011.06.008 Chia, T., Boon, F.T.H., Cheah, C., Yang, C.T. and Ghosh, D. 2020. Porcine gelatin peptide detection in cosmetic products and food confectionery using a TSQ Altis Triple Quadrupole Mass Spectrometer. Thermo Fisher. https://assets.thermofisher.com/TFS-Assets/CMD/Application-Notes/an-90606-porcine-gelatin-peptide-detection-halal-testing-an90606-en.pdf. Clark, D.P., Pazdernik, N.J. and McGehee, M.R. 2019. Molecular biology, 3rd ed. Amsterdam, Netherlands: Academic Press, an imprint of Elsevier. Deng, G., Guo, S., Zaman, F., Li, T. and Huang, Y. 2020. Recent advances in animal origin identification of gelatin-based products using liquid chromatography-mass spectrometry methods: a mini review. Rev. Anal. Chem. 39(1), 260–271; doi:10.1515/revac-2020-0121 Doi, H., Watanabe, E., Shibata, H. and Tanabe, S. 2009. A reliable enzyme linked immunosorbent assay for the determination of bovine and porcine gelatin in processed foods. J. Agricult. Food. Chem. 57(5), 1721–1726; doi:10.1021/jf802733y Duconseille, A., Astruc, T., Quintana, N., Meersman, F. and Sante-Lhoutellier, V. 2015. Gelatin structure and composition linked to hard capsule dissolution: a review. Food Hydrocolloids 43, 360–376; doi:10.1016/j.foodhyd.2014.06.006 Elgadir, M.A., Mirghani, M.E.S. and Adam, A. 2013. Fish gelatin and its applications in selected pharmaceutical aspects as alternative source to pork gelatin. Available via https://www.researchgate.net/publication/287311456_Fish_gelatin_and_its_applications_in_selected_pharmaceutical_aspects_as_alternative_source_to_pork_gelatin. El Sheikha, A.F., Mokhtar, N.F., Amie, C., Lamasudin, D.U., Isa, N.M. and Mustafa, S. 2017. Authentication technologies using DNA-based approaches for meats and halal meats determination. Food Biotechnology. 31(4), 281–315; doi: 10.1080/08905436.2017.1369886 Farrel, R.E. 2023. RNA methodologies: quantitative PCR techniques. Amsterdam, Netherlands: Elsevier. pp: 301–49. Fatoni NA, Djuminar A, Hardiana AT, Merdekawati F. 2025. Optimization of annealing temperature and primer concentration of cytochrome B (CYT B) gene for pig DNA detection with real-time PCR method. J. Vocational. Health. Stud. 8(3), 164–170. Goris, N., Vandenbussche, F., Herr, C., Villers, J., Van Der Stede, Y. and De Clercq, K. 2009. Validation of two real-time RT-PCR methods for foot-and-mouth disease diagnosis: rNA-extraction, matrix effect, uncertainty of measurement and precision. J. Virological Methods 160(1–2), 157–162; doi:10.1016/j.jviromet.2009.05.005 Green, M.R. and Sambrook, J. 2018. Optimizing primer and probe concentrations for use in real-time polymerase chain reaction (PCR) assays. Cold. Spring. Harbor. Protocols. 10(10), 825–835. doi:10.1101/pdb.prot095018 Guo, S., Xu, X., Zhou, X. and Huang, Y. 2018. A rapid and simple UPLC-MS/MS method using collagen marker peptides for identification of porcine gelatin. RSC. Adv. 8(7), 3768–3773; doi:10.1039/c7ra12539a Hassan, M., Hussain, D., Kanwal, T., Xiao, H.M. and Ghulam Musharraf, S. 2024. Methods for detection and quantification of gelatin from different sources. Food Chem. 438, 137970; doi:10.1016/j.foodchem.2023.137970 He, X.J., Zhang, Q., Liu, Y.J. and Pan, X.Y. 2009. Increasing specificity of real time PCR to detect microRNA through primer design and annealing temperature increase. Beijing. Da. Xue. Xue. Bao. Yi. Xue. Ban.=. J. Peking. Univ. Health. Sci. 41(6), 691–698. Hedman, J., Lavander, M., Salomonsson, E.N., Jinnerot, T., Boiso, L., Magnusson, B. and Rådström, P. 2018. Validation guidelines for PCR workflows in bioterrorism preparedness, food safety and forensics. Accreditation Qual. Assurance 23(3), 133–144; doi:10.1007/s00769-018-1319-7 Herdiana, Y., Sofian, F.F., Shamsuddin, S. and Rusdiana, T. 2024. Towards halal pharmaceutical: exploring alternatives to animal-based ingredients. Heliyon 10(1), e23624; doi:10.1016/j.heliyon.2023.e23624 Jue, E., Witters, D. and Ismagilov, R.F. 2020. Two-phase wash to solve the ubiquitous contaminant-carryover problem in commercial nucleic-acid extraction kits. Scientific. Rep. 10(1), 1940; doi:10.1038/s41598-020-58586-3 Karim, A.A. and Bhat, R. 2008. Gelatin alternatives for the food industry: recent developments, challenges and prospects. Trends Food Sci. & Technol. 19(12), 644–656; doi:10.1016/j.tifs.2008.08.001 Karim, A.A. and Bhat, R. 2009. Fish gelatin: properties, challenges, and prospects as an alternative to mammalian gelatins. Food. Hydrocolloids. 23(3), 563–576; doi:10.1016/j.foodhyd.2008.07.002 Khayyira, A.S., Estepane, V.M. and Malik, A. 2018. Rapid PCR–based detection optimization of porcine DNA in gelatin capsule shell. Int. J. Appl. Pharmaceutics. 10(6), 217–223; doi:10.22159/ijap.2018v10i6.29346 Kim, Y., Lee, H.S. and Lee, K.G. 2023. Detection of porcine DNA in Korean processed foods by real-time PCR. Food. Sci. Biotechnol. 32(1), 21–26; doi:10.1007/s10068-022-01169-x Kim, Y.S., Yu, H.K., Lee, B.Z. and Hong, K.W. 2018. Effect of DNA extraction methods on the detection of porcine ingredients in halal cosmetics using real-time PCR. Appl. Biol. Chem. 61(5), 549–555; doi:10.1007/s13765-018-0389-x King, C.E., Debruyne, R., Kuch, M., Schwarz, C. and Poinar, H.N. 2009. A quantitative approach to detect and overcome PCR inhibition in ancient DNA extracts. BioTechniques 47(5), 941–949; doi:10.2144/000113244 Kokkinos, P.A., Ziros, P.G., Bellou, M. and Vantarakis, A. 2014. Loop-mediated isothermal amplification (LAMP) for the detection of Salmonella in food. Food Anal. Methods 7(2), 512–526; doi:10.1007/s12161-013-9748-8 Lorenz, T.C. 2012. Polymerase chain reaction: basic protocol plus troubleshooting and otimization strategies. J. Visualized Experiments 63(3998), 10.3791/3998. Maeda, H., Fujimoto, C., Haruki, Y., Maeda, T., Kokeguchi, S., Petelin, M., Arai, H., Tanimoto, I., Nishimura, F. and Takashiba, S. 2003. Quantitative real-time PCR using TaqMan and SYBR Green for Actinobacillus actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, tetQ gene and total bacteria. FEMS. Immunol. Med. Microbiol. 39(1), 81–86; doi:10.1016/S0928-8244(03)00224-4 Malhotra, K. 1998. Interaction and effect of annealing temperature on primers used in differential display RT-PCR. Nucleic. Acids. Res. 26(3), 854–856; doi:10.1093/nar/26.3.854 Martínez Steele, E., Baraldi, L.G., Louzada, M.L.D.C., Moubarac, J.C., Mozaffarian, D. and Monteiro, C.A. 2016. Ultra-processed foods and added sugars in the US diet: evidence from a nationally representative cross-sectional study. BMJ. Open 6(3), 9892; doi:10.1136/bmjopen-2015-009892 Mohamad, N.A., Mustafa, S., Khairil Mokhtar, N.F. and El Sheikha, A.F. 2018. Molecular beacon-based real-time PCR method for detection of porcine DNA in gelatin and gelatin capsules. J. Sci. Food Agriculture 98(12), 4570–4577; doi:10.1002/jsfa.8985 Naharros-Molinero, A., Caballo-González, M.A., De La Mata, F.J. and García-Gallego, S. 2024. Shell formulation in soft gelatin capsules: design and characterization. Adv. Healthcare Mater. 13(1), 2302250; doi:10.1002/adhm.202302250 Nawwaruddin, H.H., Rohman, A. and Laksitorini, M.D. 2024. Porcine derived ingredients in cosmetic products and its halal authentication method within complex matrices. J. Food Pharm. Sci. 12(3), 241–265; doi:10.22146/jfps.17717 Nhari, R.M.H.R., Hanish, I., Mokhtar, N.F.K., Hamid, M. and El Sheikha, A.F. 2019. Authentication approach using enzyme-linked immunosorbent assay for detection of porcine substances. Qual Assur Safety Crops Foods. 11(5), 449–457; doi: 10.3920/QAS2018.1415 Nybo, K. 2011. QPCR efficiency calculations. BioTechniques 51(6), 401–402. Ramamurthy, M., Alexander, M., Aaron, S., Kannangai, R., Ravi, V., Sridharan, G. and Abraham, A. 2011. Comparison of a conventional polymerase chain reaction with real-time polymerase chain reaction for the detection of neurotropic viruses in cerebrospinal fluid samples. Indian. J. Med. Microbiol. 29(2), 102–109; doi:10.4103/0255-0857.81777 Rico, F., Mazabel, A., Egurrola, G., Pulido, J., Barrios, N., Marquez, R. and García, J. 2023. Meta-analysis and analytical methods in cosmetics formulation: a review. Cosmetics 11(1), 1; doi:10.3390/cosmetics11010001 Rohman, A., Orbayinah, S., Hermawan, A., Sudjadi, S., Windarsih, A. and Handayani, S. 2022. The development of real-time polymerase chain reaction for identification of beef meatball. Appl. Food Res. 2(2), 100148; doi:10.1016/j.afres.2022.100148 Rohman, A., Windarsih, A., Erwanto, Y. and Zakaria, Z. 2020. Review on analytical methods for analysis of porcine gelatine in food and pharmaceutical products for halal authentication. Trends. Food. Sci. Technol. 101, 122–132; doi:10.1016/j.tifs.2020.05.008 Schrader, C., Schielke, A., Ellerbroek, L. and Johne, R. 2012. PCR inhibitors—occurrence, properties and removal. J. Appl. Microbiol. 113(5), 1014–1026; doi:10.1111/j.1365-2672.2012.05384.x Sha, X.M., Wang, G.Y., Li, X., Zhang, L.Z. and Tu, Z.C. 2020. Identification and quantification of gelatin by a high-resolution mass spectrometry-based label-free method. Food Hydrocolloids 101, 105476; doi:10.1016/j.foodhyd.2019.105476 Shabani, H., Mehdizadeh, M., Mousavi, S.M., Dezfouli, E.A., Solgi, T., Khodaverdi, M., Rabiei, M., Rastegar, H. and Alebouyeh, M. 2015. Halal authenticity of gelatin using species-specific PCR. Food. Chem. 184, 203–206; doi:10.1016/j.foodchem.2015.02.140 Shen, C.H. 2019. Diagnostic molecular biology: detection and analysis of nucleic acids. Amsterdam, Netherlands: Academic Press, pp: 167–85. Sudjadi., Wardani, H.S., Sepminarti, T. and Rohman, A. 2016. Analysis of porcine gelatin DNA in a commercial capsule shell using real-time polymerase chain reaction for halal authentication. Int. J. Food. Properties. 19(9), 2127–2134; doi:10.1080/10942912.2015.1110164 Sun, Y., Nakamura, T., Ohtsu, Y., Kakehi, M., Danno, N., Shimizu, H., Tanaka, Y., Serelli-Lee, V., Tanaka, S., Okayama, T., Suda, Y., Moriya, Y., Hanada, T. and Saito, Y. 2023. Development and validation of qPCR methods for nucleic acid biomarkers as a drug development tool: points to consider. Bioanalysis 15(17), 1069–1081; doi:10.4155/bio-2023-0071 Tajadini, M., Panjehpour, M. and Javanmard, S. 2014. Comparison of SYBR Green and TaqMan methods in quantitative real-time polymerase chain reaction analysis of four adenosine receptor subtypes. Adv. Biomed. Res. 3, 85; doi:10.4103/2277-9175.127998 Tukiran, N.A., Ismail, A., Mustafa, S. and Hamid, M. 2016a. Determination of porcine gelatin in edible bird’s nest by competitive indirect ELISA based on anti-peptide polyclonal antibody. Food. Control. 59, 561–566; doi:10.1016/j.foodcont.2015.06.039 Tukiran, N.A., Ismail, A., Mustafa, S. and Hamid, M. 2016b. Development of antipeptide enzyme-linked immunosorbent assay for determination of gelatin in confectionery products. Int. J. Food. Sci. Technol. 51(1), 54–60; doi:10.1111/ijfs.12971 Uddin, S.M.K., Hossain, M.A.M., Sagadevan, S., Al Amin, M. and Johan, M.R. 2021. Halal and Kosher gelatin: applications as well as detection approaches with challenges and prospects. Food. Bioscience. 44, 101422; doi:10.1016/j.fbio.2021.101422 Venien, A. and Levieux, D. 2005. Differentiation of bovine from porcine gelatines using polyclonal anti-peptide antibodies in indirect and competitive indirect ELISA. J. Pharm. Biomed. Anal. 39(3–4), 418–424; doi:10.1016/j.jpba.2005.04.013 Wardani, H.S., Sudjadi and Rohman, A. 2015. Analysis of porcine DNA in capsule shells using porcine mitochondrial DNA D-loop pimers with real-time polymerase chain reaction method. Master Thesis, Faculty of Pharmacy, Gadjah Mada University, Yogyakarta, Indonesia. Weng, C.L., Yazid, H., Appalasamy, S., Geng, B.J., Nasir, W.M.N.W.M., Muhammad, N.M.N., Iman, A.H.M. and Kumaran, J.V. 2020. Optimization of binding, washing and elution buffer for development of DNA isolation kit. In Proceeding of the 2020 International Conference on Science and Technology Series: Earth and Environmental Science. 596(1), 012008; doi:10.1088/1755-1315/596/1/012008 Xia, Z., Johansson, M.L., Gao, Y., Zhang, L., Haffner, G.D., Macisaac, H.J. and Zhan, A. 2018. Conventional versus real-time quantitative PCR for rare species detection. Ecol. Evol. 8(23), 11799–11807; doi:10.1002/ece3.4636 Yang, Y., Li, L., Wang, H., Liu, M. and Wu, Y. 2020. Development and verification of a quantitative real-time PCR method to identify and quantify gelatin derived from animal hide. J. Food. Sci. 85(9), 2762–2772; doi:10.1111/1750-3841.15362 Zhao, F., Maren, N.A., Kosentka, P.Z., Liao, Y.Y., Lu, H., Duduit, J.R., Huang, D., Ashrafi, H., Zhao, T., Huerta, A.I., Ranney, T.G. and Liu, W. 2021. An optimized protocol for stepwise optimization of real-time RT-PCR analysis. Horticulture Res. 8(1), 179; doi:10.1038/s41438-021-00616-w Zhu, X., Gu, S., Guo, D., Huang, X., Chen, N., Niu, B. and Deng, X. 2023. Determination of porcine derived components in gelatin and gelatin-containing foods by high performance liquid chromatography-tandem mass spectrometry. Food. Hydrocolloids. 134, 107978; doi:10.1016/j.foodhyd.2022.107978 | ||
| How to Cite this Article |
| Pubmed Style Rumiyati R, Sari MN, Nawwaruddin HH, Laksitorini MD, Rohman A. Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Vet. J.. 2026; 16(1): 478-491. doi:10.5455/OVJ.2026.v16.i1.45 Web Style Rumiyati R, Sari MN, Nawwaruddin HH, Laksitorini MD, Rohman A. Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. https://www.openveterinaryjournal.com/?mno=260701 [Access: January 31, 2026]. doi:10.5455/OVJ.2026.v16.i1.45 AMA (American Medical Association) Style Rumiyati R, Sari MN, Nawwaruddin HH, Laksitorini MD, Rohman A. Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Vet. J.. 2026; 16(1): 478-491. doi:10.5455/OVJ.2026.v16.i1.45 Vancouver/ICMJE Style Rumiyati R, Sari MN, Nawwaruddin HH, Laksitorini MD, Rohman A. Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Vet. J.. (2026), [cited January 31, 2026]; 16(1): 478-491. doi:10.5455/OVJ.2026.v16.i1.45 Harvard Style Rumiyati, R., Sari, . M. N., Nawwaruddin, . H. H., Laksitorini, . M. D. & Rohman, . A. (2026) Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Vet. J., 16 (1), 478-491. doi:10.5455/OVJ.2026.v16.i1.45 Turabian Style Rumiyati, Rumiyati, Miftiana Nugraha Sari, Hazza' Hammam Nawwaruddin, Marlyn Dian Laksitorini, and Abdul Rohman. 2026. Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Veterinary Journal, 16 (1), 478-491. doi:10.5455/OVJ.2026.v16.i1.45 Chicago Style Rumiyati, Rumiyati, Miftiana Nugraha Sari, Hazza' Hammam Nawwaruddin, Marlyn Dian Laksitorini, and Abdul Rohman. "Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products." Open Veterinary Journal 16 (2026), 478-491. doi:10.5455/OVJ.2026.v16.i1.45 MLA (The Modern Language Association) Style Rumiyati, Rumiyati, Miftiana Nugraha Sari, Hazza' Hammam Nawwaruddin, Marlyn Dian Laksitorini, and Abdul Rohman. "Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products." Open Veterinary Journal 16.1 (2026), 478-491. Print. doi:10.5455/OVJ.2026.v16.i1.45 APA (American Psychological Association) Style Rumiyati, R., Sari, . M. N., Nawwaruddin, . H. H., Laksitorini, . M. D. & Rohman, . A. (2026) Optimization and validation of the real-time polymerase chain reaction protocol for porcine gelatin detection in processed food and cosmetic products. Open Veterinary Journal, 16 (1), 478-491. doi:10.5455/OVJ.2026.v16.i1.45 |