| Research Article | ||
Open Vet. J.. 2026; 16(1): 616-625 Open Veterinary Journal, (2026), Vol. 16(1): 616-625 Research Article Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, BrazilCarla Maria do Carmo Resende Martins1*, Daniele Rayssa Cavalcanti de Sá2, Marlos Gomes Martins2, Maria Antônia Sampaio de Sousa2, Amanda Daiane da Silva Bezerra2, Matheus Rodrigues de Souza3, Beatriz Nascimento Araújo2, Danillo Sales Rosa2, Gisele Veneroni Gouveia2, Mateus Matiuzzi da Costa2, and Rafael Torres de Souza Rodrigues21Department of Veterinary Medicine, Federal University of the San Francisco Valley, Petrolina, Brazil 2Federal University of the San Francisco Valley, Petrolina, Brazil 3Brazilian Agricultural Research Corporation, Embrapa Semiárido , Petrolina, Brazil *Corresponding Author: Carla Maria do Carmo Resende Martins. Department of Veterinary Medicine, Federal University of the San Francisco Valley, Petrolina, Brazil. Email: cresende.veterinaria [at] gmail.com Submitted: 05/08/2025 Revised: 30/11/2025 Accepted: 14/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
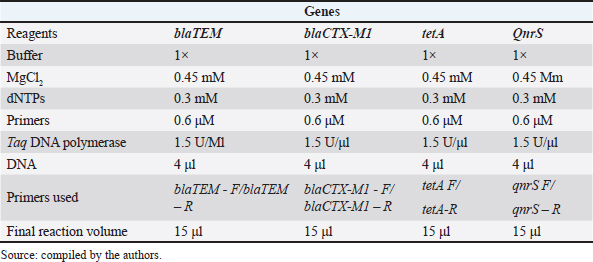
AbstractBackground: Although free-range chicken meat is widely consumed in Brazil, it may pose a public health risk due to contamination with pathogenic and antimicrobial-resistant bacteria, such as Escherichia coli (EC). Aim: This study aimed to investigate the occurrence of EC, antimicrobial resistance, and biofilm formation in free-range chicken meat samples collected from restaurants in Petrolina, Pernambuco, Brazil. Methods: Microbiological and molecular methods were used to analyze 40 samples collected from eight restaurants to detect EC, evaluate antimicrobial resistance profiles, assess biofilm formation, and identify the presence of resistance genes. Results: The overall frequency of EC infection was 57.5%. All bacterial strains (100%) were resistant to at least one antimicrobial agent, and 78.2% (18/23) were classified as MDR, showing resistance to three or more classes of antibiotics, with the highest rates observed for ciprofloxacin and tetracycline. Among the detected resistance genes, the blaTEM gene had the highest prevalence (17.4%), followed by blaCTX-M1 (4.3%), while tetA and qnrS were not detected. A discrepancy was observed between high phenotypic resistance and low detection of resistance genes, suggesting the involvement of other mechanisms. In addition, 95.5% of the strains were classified as biofilm producers, although most exhibited weak biofilm formation. Conclusion: This study confirms the presence of multidrug-resistant EC strains in food served at restaurants, highlighting the risk of transmission of resistant bacteria to humans through the consumption of contaminated meat and reinforcing the need for continuous monitoring of food safety. Keywords: Biofilm, Escherichia coli, Poultry meat, Resistance genes. IntroductionFoodborne diseases (FBDs) represent one of the major public health problems worldwide, being responsible for high morbidity and mortality as well as significant economic losses (World Health Organization, 2020; CDC, 2022). Globally, approximately 600 million people fall ill each year after consuming contaminated food, resulting in approximately 420,000 deaths (World Health Organization, 2020). Between 2013 and 2022, 6,523 FBD outbreaks were reported in Brazil, involving 107,513 individuals, 12,722 hospitalizations, and 112 confirmed deaths. The main etiological agents identified were Escherichia coli (EC), Salmonella spp., and Staphylococcus spp., with restaurants highlighted as one of the primary locations where outbreaks occur (Brazil, Ministry of Health, 2023). However, the actual number of cases may be even higher because many affected individuals do not seek medical care, and some episodes are not reported to official surveillance systems (Scallan et al., 2015; World Health Organization, 2020). This gap underscores the need for continuous monitoring of foods of animal origin, especially poultry meat, due to its widespread consumption and epidemiological relevance. Free-range chicken, traditionally consumed in Brazil, is an important source of animal protein and holds strong cultural and socioeconomic significance, particularly for family farmers and small-scale producers (Embrapa, 2019; Grossi et al., 2023a,b). Nevertheless, the artisanal nature of its production can increase the risk of microbiological contamination and, consequently, FBDs (Batista et al., 2024a,b). Birds raised in alternative systems can act as reservoirs of bacterial pathogens, including strains resistant to clinically important antimicrobials (Grossi et al., 2023a,b; Yu et al., 2025a,b; Almeida et al., 2025). Although these systems use fewer antibiotics than intensive production systems, resistance still occurs, possibly due to cross-contamination, inappropriate therapeutic use, or the environmental circulation of resistance genes (Singh et al., 2025a,b). In the Brazilian context, free-range chicken is widely offered in traditional food restaurants and in the gastronomic sector, often considered a high-value dish with prices that can exceed those of premium beef cuts (Embrapa, 2019). Restaurants are recognized as recurrent sites of foodborne outbreaks, increasing consumer concerns regarding food safety despite this added value. Food handlers play a central role in the transmission of FBD-associated pathogens in these settings (Mylius et al., 2024a,b). Given the economic and cultural importance of free-range chicken, its potential contribution to the spread of resistant pathogens, and the scarcity of studies on the microbiological safety of this product in collective consumption settings, this study aimed to investigate the occurrence of EC in free-range chicken meat sold in restaurants in Petrolina, Pernambuco, Brazil, and to assess its antimicrobial susceptibility profile, biofilm-forming ability, and the presence of resistance genes. Materials and MethodsSampling and data collectionThe sampling plan consisted of surveying restaurants that serve free-range chicken meat in the municipality of Petrolina, Pernambuco, based on information provided by the municipality’s official sanitary surveillance service. All restaurants were located within the urban perimeter of Petrolina, characterized by the sale of typical foods, and all samples originated from artisanal husbandry (small-scale producers not inspected by regulatory authorities). Fifty samples of fresh free-range chicken meat (breast, thigh, and wing) were collected from eight restaurants, totaling 40 samples. This number was established on the basis of the requirements of Normative Instruction No. 313/2024 issued by the Ministry of Health/ANVISA (Brazil, 2024). Samples were collected upon arrival at the restaurants to evaluate the products of the suppliers. Samples were placed in sterile plastic bags and stored in isothermal boxes with ice after collection. The samples were then sent to the Microbiology Laboratory at the Federal University of San Francisco Valley (UNIVASF) within a maximum period of 2 hours for EC analysis. This bacterium is a recognized indicator of hygienic quality (Niyonzima et al., 2015; World Health Organization, 2020; Brazil, 2023). Isolation and confirmation of ECThe identification and quantification of EC were carried out using the most probable number method, with the following steps: a presumptive test, in which three aliquots from three dilutions of the sample were inoculated into a series of three tubes containing lactose broth (KASVI, Paraná, Brazil) per dilution. Growth with gas production from lactose was observed in lactose broth at 35°C after 24–48 hours of incubation and was considered presumptive for the presence of coliforms. One loopful from each presumptive tube was transferred to EC broth tubes (KASVI, Paraná, Brazil) for EC identification. Growth with gas production in EC broth after 48 hours of incubation at 42.5°C was considered presumptive for the presence of EC. One loopful from each EC tube was streaked onto Levine Eosin Methylene Blue (EMB) Agar (KASVI, Paraná, Brazil) for confirmation. Plates were incubated at 35°C (Nova Técnica, NT 245, São Paulo, Brazil) for 24 hours and observed for the development of typical EC colonies. EMB plates with typical EC colonies, corresponding to EC-positive tubes with gas, were used for the calculation of EC per gram of food. Colonies with a greenish metallic sheen were observed, reflecting vigorous lactose fermentation and acid production. Colonies with typical characteristics were subjected to biochemical tests: oxidase, Indole Sulfide Motility Agar, Triple Sugar Iron, and Simmons Citrate (KASVI, Paraná, Brazil) (Quinn et al., 1993; ISO, 2005). After biochemical confirmation, each isolate was inoculated into sterilized microtubes containing tryptone soy agar (KASVI, Paraná, Brazil) medium and glycerol. The samples were then incubated at 37°C (SOLAB – SL 102, São Paulo, Brazil) for 24 hours and subsequently stored at −20°C until further analysis. Antimicrobial susceptibility test resultsAfter the microbiological analysis, the isolates were tested against different antimicrobials using the disk diffusion technique to determine the antimicrobial susceptibility profile (Bauer et al., 1966; CLSI, 2024). Disks of ampicillin (AMP, 10 µg), cefotaxime (CTX, 30 µg), chloramphenicol (CLO, 30 µg), streptomycin (EST, 10 µg), ciprofloxacin (CIP, 5 µg), tetracycline (TET, 30 µg), and azithromycin (AZI, 15 µg) were used (DME, São Paulo, Brazil). For the tests, the strains were taken from plates containing NA and incubated at 35°C for 24 hours. Colonies of each strain were inoculated into 5 ml of 0.85% saline solution using a sterile loop from the nutrient agar plates in sufficient quantity to reach the 0.5 McFarland turbidity standard. All materials used were previously sterilized. Then, using a sterile swab, the inoculum was uniformly spread onto a Petri dish containing Mueller–Hinton agar. Antimicrobial discs were then evenly placed on the agar surface. Plates were incubated at 35°C for 24 hours, and the diameter of the inhibition halo, including the disc diameter, was measured using a caliper. The diameters were recorded and compared with those of drug-specific reference standards. Strains were classified as susceptible, intermediate, or resistant based on the halo diameter (CLSI, 2024). EC American Type Culture Collection (ATCC) 25922 was used as a quality control strain for antimicrobial susceptibility testing. Quantification of biofilm productionThe gentian violet test (adapted from Stepanović et al., 2007; Merino, 2009) was used to quantify biofilm formation. The ATCC strain EC ATCC 25922 was used in addition to the isolates. A volume of 5 µl of a bacterial suspension at 6 × 106 CFU/ml was added to a 96-well microplate (OLEN, São José dos Pinhais, Brazil) containing 195 µl of Luria-Bertani Broth (Sigma-Aldrich, St. Louis, MO). Negative controls were prepared using only the culture medium, and the microplates were incubated at 37°C for 24 hours. Subsequently, the wells were washed thrice with 200 µl of sterile distilled water, the contents were discarded, and 150 µl of methyl alcohol P.A. (ISOFAR, Trenthorst, Germany) was added for 20 minutes. The contents were discarded, dried overnight, and addition of 100 µl of 0.25% gentian violet (Proquímios, Rio de Janeiro, Brazil) for 5 minutes. Three additional washes were performed as previously described, and 200 µl of ethanol-acetone solution [99.8% absolute ethyl alcohol (NEON, São Paulo, Brazil) with 99.5% acetone (Fmaia, Belo Horizonte, Brazil) in an 80:20 ratio] was added. The absorbance was read at 620 nm using an EXPERT PLUS-UV model microplate reader (ASYS, Cambridge, UK) at 620 nm. The samples were analyzed in triplicate. Qualitative classification of biofilm formation was performed according to Stepanović and S (2000). DNA extraction and polymerase chain reaction for detection of resistance genes in ECThe DNA of each EC strain was extracted by adapting the methodology proposed by Regitano (2001). Initially, 500 µl of sodium dodecyl sulfate/Proteinase K lysis solution (10 mM Tris-HCl, pH 8.0, 100 mM NaCl, 10 mM EDTA, pH 8.0, 0.5% SDS, and 2 µg of proteinase K; autoclaved Milli-Q water added to a final volume of 500 µl) was added to the tubes containing the pellet. The tubes were then vortexed until the pellet completely detached from the bottom. Subsequently, the samples were incubated in a dry bath at 55°C for 4–6 hours and left overnight when necessary. After incubation, 210 µl of Tris-EDTA buffer (pH 7.6) and 240 µl of 5 M NaCl were added to each tube. The tubes were gently inverted until small clots formed. The samples were then incubated on ice for 10 minutes and centrifuged for 19 minutes at 12,740 × g. Next, 750 µl of the supernatant was collected and transferred to a new microtube. To this volume, 750 µl of 100% isopropanol (at room temperature) was added, followed by gentle inversion to ensure homogeneous mixing. The samples were centrifuged again for 19 minutes at 12,740 × g, and the resulting supernatant was discarded. The pellet was left to dry inverted on absorbent paper for 10 minutes, after which 500 µl of chilled 70% ethanol was added to the pellet. The samples were maintained at room temperature for 5 minutes and then centrifuged for 6 minutes at 12,740 × g. After this step, the supernatant was discarded, and the pellet was left to air-dry inverted on absorbent paper until complete alcohol evaporation (approximately 10 minutes). A TE solution containing RNase was prepared by adding 0.1 µl of RNase (10 µg/µl) for every 100 µl of TE. The pellet was resuspended in up to 100 µl of this solution. The samples were incubated for 1 hour at 37°C for 1 hour and then stored overnight in a refrigerator to ensure complete RNA digestion. DNA was quantified and stored at −20 °C for later use. For EC isolates, PCR was performed to detect genes encoding resistance to β-lactams, TETs, and quinolones. All isolates were genotyped for blaTEM, blaCTX-M1, tetA, and qnrS. Table 1 lists the conditions used in the PCR mix for each gene. Table 1. PCR mix conditions for blaTEM, blaCTX-M1, tetA, and qnrS genes.
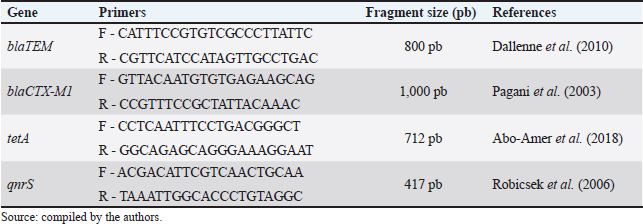
The amplification reactions consisted of an initial denaturation at 94°C for 10 minutes, followed by 30 cycles of denaturation at 94°C for 40 seconds, annealing at 60°C for 40 seconds, and extension at 72°C for 60 seconds, with a final extension at 72°C for 7 minutes. Table 2 lists the primers used in the amplifications. Each gene was amplified separately, and the PCR products were analyzed by electrophoresis in 2% (w/v) agarose gel prepared in 1× TBE buffer containing ethidium bromide (0.5 µg/ml) for visualization of DNA fragments. The gels were run at 80 V, with an approximate current of 120 mA and a power of 20 W, for 1 hour 30 minutes to 2 hours, until the molecular weight marker reached adequate migration. Subsequently, the gels were documented under ultraviolet light using a transilluminator. Table 2. Primers used for target gene amplification.
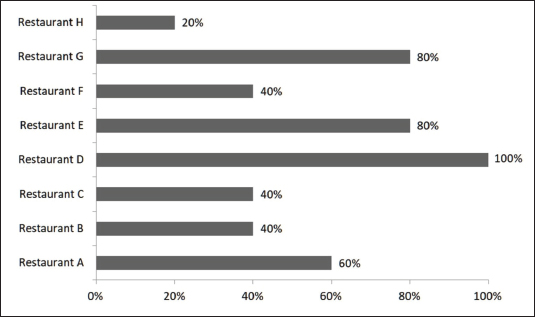
Statistical analysisStatistical analyses were performed using SAS® 9.4 (SAS Institute Inc., Cary, NC). The procedures PROC FREQ (for chi-square tests and effect size estimation), PROC MEANS (for descriptive statistics), and PROC SGPLOT (for graphical visualization) were employed. All tests were two-tailed, and the significance level was set at α=0.05. The dataset comprised 40 observations (eight restaurants, labeled A–H; five samples per restaurant). Associations between restaurant identity and conformity were evaluated using Pearson’s chi-square test. When the expected cell frequencies were <5, exact chi-square and Fisher’s exact tests were computed to verify the robustness of the results. The strength of association was quantified using Cramér’s V and the contingency coefficient, while standardized residuals (|z| > 2.0) were examined to identify the cells that contributed the most to the overall chi-square statistic. The variable conformity was recoded into a binary outcome to estimate the rate of non-conformity per restaurant (1=No, 0=Yes). The mean of this binary variable was used to calculate the proportion of non-conforming samples, expressed as a percentage with exact 95% binomial confidence intervals (Clopper–Pearson method, two-sided). These percentages were visually represented as horizontal bar plots (0%–100% scale) and ordered from restaurant H to A, with percentage values displayed above each bar. Ethical approvalNot needed for this study. ResultsOccurrence of EC infection in free-range chicken meat samplesOverall, 57.5% of the samples (23/40) were classified as nonconforming, whereas 42.5% (17/40) met the established conformity criteria. The proportion of nonconforming samples varied among restaurants, ranging from 20% in Restaurant H to 100% in Restaurant D (Fig. 1). The effect size (Cramér’s V=0.51) indicated a moderate practical association between restaurant identity and conformity status.
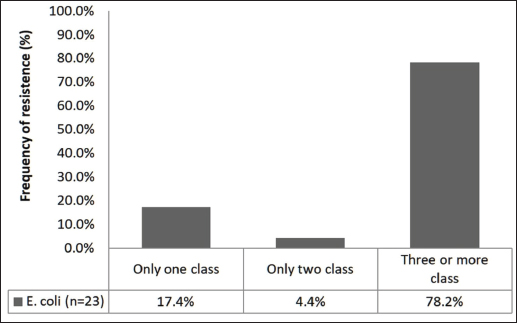
Fig. 1. EC in fresh free-range chicken meat samples collected from restaurants in Petrolina, Pernambuco, Brazil. However, the Pearson chi-square test revealed no statistically significant association between restaurant and conformity (χ²(7)=10.54, p=0.1601). Similarly, the likelihood-ratio chi-square test supported this result (χ²(7)=12.62, p=0.0820). Although the latter p-value approached the conventional threshold of significance, the test assumptions were not fully satisfied because several expected cell frequencies were below five. Therefore, Fisher’s exact test was applied as a more robust alternative, confirming the absence of a significant relationship (p=0.2003). These findings indicate that the observed variations in conformity rates across restaurants were not statistically significant. Bacterial resistance profileAntimicrobial susceptibility testing revealed that all EC isolates (23/23) were resistant to at least one antimicrobial agent. Among them, 78.2% (18/23) were classified as multidrug resistance, exhibiting resistance to three or more classes of antibiotics (Fig. 2).
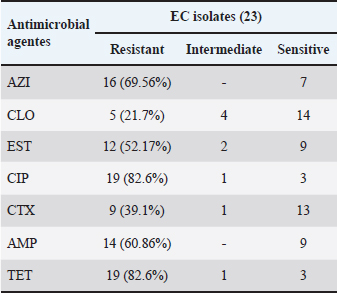
Fig. 2. Antimicrobial susceptibility profiles of EC isolates obtained from fresh free-range chicken meat samples collected from restaurants in Petrolina, Pernambuco, Brazil. The highest resistance rates among EC isolates were observed against CIP and TET (both 82.6%), followed by AZI (69.5%) and AMP (60.8%). The lowest rate was observed for CLO (21.7%). The complete data are presented in Table 3. Table 3. Antimicrobial susceptibility profiles of EC isolates obtained from fresh free-range chicken meat samples collected from restaurants in Petrolina, Pernambuco, Brazil.
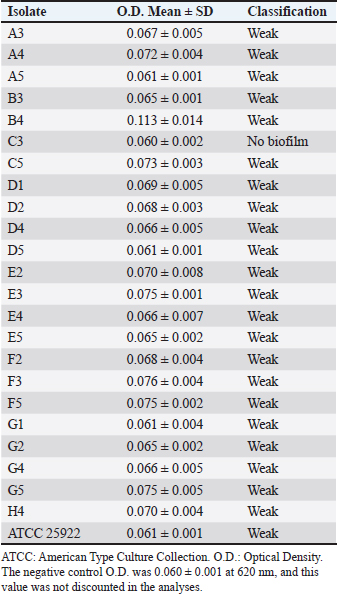
Biofilm formationAmong the 23 EC isolates tested, 22 (95.65%) were classified as weak biofilm formers, whereas one isolate (C3) showed no biofilm formation ability. The ATCC 25922 strain also exhibited weak biofilm formation. The quantification results are presented in Table 4. Table 4. Quantification and classification of biofilm formation by EC isolates obtained from fresh free-range chicken meat samples from restaurants in Petrolina, Pernambuco, Brazil.
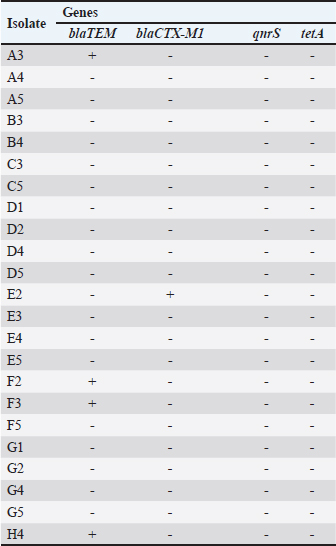
Detection of resistance genes using PCRAll EC strains were tested for the presence of the genes blaTEM and blaCTX-M1, which are associated with resistance to β-lactams, as well as the tetA and qnrS, which are associated with resistance to TETs and quinolones, respectively. The blaTEM gene showed the highest frequency (17.4%; 4/23) (Fig. 3), followed by the blaCTX-M1 gene (4.3%; 1/23). All EC isolates tested were negative for the presence of tetA and qnrS (Table 5).
Fig . 3. Detection of the blaTEM gene in EC isolates by gel electrophoresis. L – base- pair marker (bp); N, negative control; P, positive control. Source: The author. Table 5. PCR results for the presence of antibiotic resistance genes in EC obtained from fresh free-range chicken meat samples collected from restaurants in Petrolina, Pernambuco, Brazil.
DiscussionThe 57.5% prevalence of EC in this study confirms the microbiological vulnerability of free-range chicken meat. This rate is higher than that reported in other countries, ranging from 33% to 58% (Saud et al., 2019; Li et al., 2019; Guo et al., 2019). These differences reflect variations in husbandry systems, processing practices, and sanitary oversight. In the case of Petrolina, the uniformity of small-scale, locally managed, and uninspected production systems likely contributed to the observed elevated prevalence. Most available literature on poultry contamination in Brazil generally focuses on samples from markets and retail outlets (Crecencio et al., 2020; Grossi et al., 2023a,b). To the best of our knowledge, no previous studies have specifically evaluated free-range chicken served in restaurants, a context in which consumption is immediate and the risk of dissemination to consumers is greater. This distinctive focus makes this study particularly valuable for sanitary monitoring efforts. Although cooking reduces bacterial load, pathogens have been reported to persist in processed meats has been reported (Niyonzima et al., 2015; Anihouvi et al., 2024). Inadequate hygiene practices may lead to cross-contamination after preparation (Sharma et al., 2024). Thus, free-range chicken meat served in restaurants poses a risk during raw handling, subsequent processing steps, and environmental exposure. All isolates in this study exhibited resistance to at least one antimicrobial agent, and 78.2% were classified as multidrug-resistant, a notably high rate even for poultry meat. Resistance to CIP and TET was the most prevalent, with additional concern raised by resistance to AMP and AZI, which are drugs relevant to both veterinary and human medicine (Abreu et al., 2023). This finding is consistent with studies from different countries (Zhang et al., 2017; Yu et al., 2025a,b) but is noteworthy given that antibiotics are less used in free-range systems than in intensive production systems. The observed resistance may be associated with multiple factors, including environmental exposure, contact with animal carriers, contaminated water, inappropriate therapeutic use, and hygiene failures during processing. The relatively lower resistance to CLO (21.7%) and CTX (39.1%) suggests limited exposure to these drugs. However, the presence of CTX resistance remains a concern, as it is frequently associated with HAIs (Callens et al., 2018). An important finding was the discrepancy between the high phenotypic resistance and the low frequency of detected genes. The panel included only blaTEM, blaCTX-M1, tetA, and qnrS; therefore, other relevant determinants, such as tetB or qnrA/qnrB variants, were not captured. In addition, chromosomal mechanisms, such as mutations in gyrA and parC, which confer fluoroquinolone resistance, or the overexpression of efflux pumps, may explain some of the observed phenotypes. These results highlight the need to expand molecular analysis in future studies to better understand the genetic basis of resistance (Thomrongsuwannakij and T, 2020; Roy and R, 2021; Haque and S, 2022). Regarding biofilm formation, 95.5% of the isolates were classified as biofilm formers, although most showed a weak production profile. However, even low-density biofilms pose a risk on food-contact surfaces because they increase bacterial persistence and promote the transfer of resistance genes (Scherrer and Marcon, 2016; Maheshwari et al., 2016; Sekoai et al., 2020). Therefore, even at low levels, the high frequency of strains capable of forming biofilms should be considered a critical factor in maintaining contamination in restaurant kitchens. Taken together, these findings reinforce the role of free-range chicken served in restaurants as an important vehicle for the dissemination of multidrug-resistant EC. Beyond highlighting a poorly explored risk in the literature, this study helps fill knowledge gaps regarding the microbiological safety of this product and points to the need for specific monitoring and control strategies in collective consumption environments. An important limitation of this study was the difficulty in obtaining participation from some restaurants in the sample collection, possibly due to concerns about the microbiological analysis results. In addition, the number of establishments selling free-range chicken meat in Petrolina is relatively limited, which reduced sampling coverage. These factors may have limited the results’ representativeness relative to other contexts or regions, reinforcing the need for further investigations with a larger number of establishments across different locations. The panel of resistance genes analyzed was limited to a few determinants (blaTEM, blaCTX-M1, tetA, and qnrS), which may have contributed to the discrepancy between high phenotypic resistance and low genotypic detection. Nevertheless, the findings provide an important baseline, and future analyses using broader panels may better elucidate the mechanisms. Despite these limitations, this study represents the first Brazilian investigation to comprehensively assess the occurrence of MDR and biofilm-producing EC in free-range chicken meat served in restaurants, providing novel baseline data to support sanitary surveillance and future research. The combined use of microbiological and molecular methods strengthened isolate detection and characterization of antimicrobial resistance profiles, thereby increasing the reliability of the findings. Moreover, the focus on the food service environment provides information directly applicable to public health and food safety interventions. From a practical standpoint, these descriptive trends reinforce the importance of continuous staff training, implementation of standardized sanitation protocols, and periodic internal audits even in the absence of statistically significant differences. Altogether, such measures are essential to ensure sustained compliance with microbiological safety standards and to minimize the potential public health risks associated with foodborne pathogens in food service settings. ConclusionTo the best of our knowledge, this is the first Brazilian study to report that free-range chicken meat served in restaurants can harbor multidrug-resistant and biofilm-forming EC. Despite the prohibition of antibiotic use in free-range chicken production, the detection of resistant strains suggests possible illicit use of antimicrobials or cross-contamination during processing. These findings raise concerns regarding the potential occurrence of FBDs and therapeutic failures. Furthermore, biofilm formation may increase bacterial persistence along the food chain. This study provides important baseline data to support future investigations on food handling practices and highlights the need for production and supply chain surveillance to mitigate public health risks. AcknowledgmentsThe authors would like to thank the Foundation for the Support of Science and Technology of the State of Pernambuco (FACEPE) and the Federal University of the São Francisco Valley (UNIVASF) for their institutional support and for providing the equipment and materials necessary to conduct this study. Conflict of interestThe authors declare no conflicts of interest related to this study. FundingThis study was financially supported by the Foundation for the Support of Science and Technology of the State of Pernambuco (FACEPE), which provided full funding for the execution of the experiments. Authors’ contributionsAll authors contributed significantly to the study’s development. Their contributions included research conception and design, experiment execution, data analysis, and manuscript preparation. All authors have reviewed and approved the final version of the manuscript. Carla Maria do Carmo Resende Martins: Performed all laboratory experiments, including microbiological and molecular analyses; conducted biofilm assays; organized and interpreted the data; and was responsible for the initial drafting of the manuscript. Daniele Rayssa Cavalcanti de Sá: As an undergraduate research fellow, assisted in microbiological experiments and sample processing. Marlos Gomes Martins: Coordinated the microbiology laboratory, provided technical training to the team, and ensured laboratory resources for conducting the experiments. Maria Antônia Sampaio de Sousa: Assisted in microbiological experiments, culture handling, and antimicrobial susceptibility testing. Amanda Daiane da Silva Bezerra: Contributed to microbiological assays and supported routine laboratory activities. Matheus Rodrigues de Souza: Performed statistical analysis and revised the manuscript. Beatriz Nascimento Araújo: Assisted in molecular genetics experiments, including DNA extraction and polymerase chain reaction (PCR) assays. Danillo Sales Rosa: Collaborated in biofilm formation experiments and data collection related to biofilm assays. Gisele Veneroni Gouveia: Coordinated the molecular genetics laboratory, supervised genetic analyses, and provided molecular methodology guidance. Mateus Matiuzzi da Costa: Coordinated the microbiology laboratory, supervised biofilm evaluation experiments, and provided scientific advice throughout the project. Rafael Torres de Souza Rodrigues: Supervised the study, guided the experimental design and data interpretation, critically reviewed and edited the manuscript, and provided continuous scientific supervision throughout the research. Data availabilityThe data supporting the findings of this study are available upon reasonable request from the corresponding author. ReferencesAbo-Amer, A.E., Shobrak, M.Y. and Altalhi, A.D. 2018. Isolation and antimicrobial resistance among Escherichia coli isolated from farm chickens in Taif province, Saudi Arabia. J. Glob. Antimicrob. Resist. 15, 65–68. Abreu, J.M.P., Silva, L.M. and Lopes, V.C. 2023. Antimicrobial resistance in poultry production: a One Health perspective. Microorganisms 11, 953. Almeida, F., Barros, T. and Silva, M. 2025. Occurrence of antimicrobial resistance genes in free-range chicken meat. J. Appl. Microbiol. 138, 50–62. Anihouvi, D.G.H., Koné, K.M., Anihouvi, V.B. and Mahillon, J. 2024. Sanitary quality and bacteriological antibiotic-resistance pattern of frozen raw chicken meat sold in retail market in Benin. J. Agric. Food Res. 9, 100573. Batista, R., Oliveira, J. and Souza, C. 2024. Foodborne pathogens in free-range poultry meat: microbiological risks and control strategies. Food. Control. 156, 1–9. Batista, R., Valverde, M.M. and Cavalcante, A.K.S. 2024. Sanitary challenges in poultry farming. Rev. Meio. Ambient. Sustentab. 13, 92–105. Bauer, A.W., Kirby, W.M.M., Sherris, J.C. and Turck, M. 1966. Antibiotic susceptibility testing by a standardized single-disk method. Am. J. Clin. Pathol. 45, 493–496. Brazil, Ministry of Health. 2023. Outbreaks of waterborne and foodborne diseases in Brazil: 2023 Report. Brasília: Ministry of Health. BRAZIL. 2024. National Health Surveillance Agency (ANVISA). Normative Instruction N°. 313, of September 4, 2024. Amends Normative Instruction N°. 161, of July 1, 2022, which establishes the microbiological standards for food. Official Gazette of the Union: Section 1, Brasília, DF. (Accessed 25 June 2025) Callens, B., Cargnel, M., Sarrazin, S., Dewulf, J., Hoet, B., Vermeersch, K., Wattiau, P. and Welby, S. 2018. Associations between a decreased veterinary antimicrobial use and resistance in commensal Escherichia coli from Belgian livestock species (2011–2015). Prev. Vet. Med. 157, 50–58. CDC. 2022. Foodborne germs and illnesses. Atlanta, Georgia: CDC. Available via https://www.cdc.gov/foodborne (Accessed 16 September 2025) CLSI. 2024. Performance Standards for Antimicrobial Disk Susceptibility Tests; M02 – 14th ed. Wayne, PA: Clinical and Laboratory Standards Institute. Crecencio, R.B., Brisola, M.C., Bitner, D., Frigo, A., Rampazzo, L., Borges, K.A., Furian, T.Q., Salle, C.T.P., Moraes, H.L.S., Faria, G.A., Da Silva, A.S. and Stefani, L.M. 2020. Antimicrobial susceptibility, biofilm formation and genetic profiles of Escherichia coli isolated from retail chicken meat. Infect. Genet. Evol. 84, 104355. Dallenne, C., Da Costa, A., Decré, D., Favier, C. and Arlet, G. 2010. Development of a set of multiplex PCR assays for the detection of genes encoding important β-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 65, 490–495. Embrapa. 2019. Manual of the sustainable production system for free-range chickens. Brasília, Brazil: Brazilian Agricultural Research. Grossi, E., Silva, T.M., Menezes, L.G. and Campos, C.A. 2023. High prevalence of intermediate resistance to ciprofloxacin and detection of qnrS gene in Escherichia coli isolated from chicken meat in Brazil. Int. J. Food. Microbiol. 397, 110195. Grossi, M., Almeida, L. and Santos, P. 2023. Antimicrobial resistance in poultry production: implications for food safety. Poult. Sci. 102(1), 1–12. Guo, S., Tay, M.Y.F., Aung, K.T., Seow, K.L.G., Ng, L.C., Purbojati, R.W., Drautz-Moses, D.I., Schuster, S.C. and Schlundt, J. 2019. Phenotypic and genotypic characterization of antimicrobial resistant Escherichia coli isolated from ready-to-eat food in Singapore using disk diffusion, broth microdilution and whole genome sequencing methods. Food Control 99, 89–97. Haque, T. A., Urmi, U. L., Islam, A. B. M. M. K., Ara, B., Nahar, S., Mosaddek, A. S. M., Lugova, H., Kumar, S., Jahan, D., Rahman, N. A. A., Haque, M., Islam, S. and Godman, B. 2022. Detection of qnr genes and gyrA mutation to quinolone phenotypic resistance of UTI pathogens in Bangladesh and the implications. J. Appl. Pharm. Sci. 12(4), 185–198. ISO. 2005. ISO 7251:2005 — Microbiology of food and animal feeding stuffs — Horizontal method for the detection and enumeration of presumptive Escherichia coli — Most probable number technique. Geneva, Switzerland: ISO. Li, Y., Pei, X., Zhang, X., Wu, L., Liu, Y., Zhou, H., Ma, G., Chen, Q., Liang, H. and Yang, D. 2019. A surveillance of microbiological contamination on raw poultry meat at retail markets in China. Food Control 104, 99–104. Maheshwari, M., Ahmad, I. and Althubiani, A.S. 2016. Multidrug resistance and transferability of blaCTX-M among extended-spectrum β-lactamase-producing enteric bacteria in biofilm. J. Glob. Antimicrob. Resist. 6, 142–149. Merino, N., Toledo-Arana, A., Vergara-Irigaray, M., Valle, J., Solano, C., Calvo, E., Lopez, J.A., Foster, T.J., Penadés, J.R. and Lasa, I. 2009. Protein A-mediated multicellular behavior in Staphylococcus aureus. J. Bacteriol. Feb;191(3), 832–843. Mylius, C. da C., Wermann, R. G., Augustin, P. C., Strada, L. L., Lohmann, P. M., Vogel, P., Bertani, J. P. B., Conde, S. R., Adami, F. S. and Cé, P. F. 2024. Evaluation of good practices among food handlers in two food and nutrition units. Int. J. Sci. Manag. Tour. 10(5), 1129. Mylius, S., Jacobs J. and Verhoef, L. 2024. Food handlers as vectors of foodborne disease: a risk assessment perspective. Epidemiol. Infect. 152(1), 1–9. Niyonzima, E., Ongol, M.P., Kimonyo, A. and Sindic, M. 2015. Risk factors and control measures for bacterial contamination in the bovine meat chain: a review on Salmonella and pathogenic E. coli. J. Food Res. 4(5), 98. Pagani, L., Dell’Amico, E., Migliavacca, R., D’Andrea, M.M., Giacobone, E., Amicosante, G., Romero, E. and Rossolini, G.M. 2003. Multiple CTX-M-type extended-spectrum β-lactamases in nosocomial isolates of Enterobacteriaceae from a hospital in Northern Italy. J. Clin. Microbiol. 41(9), 4264–4269. Quinn, P.J., Carter, M.E., Markey B. and Carter, G.R. 1993. Clinical veterinary microbiology. London, UK: Mosby. Embrapa Informação Tecnológica (eds), Extração de DNA para aplicação em reação em cadeia da polimerase (PCR), Brasília, Brazil. Robicsek, A., Jacoby, G.A. and Hooper, D.C. 2006. The worldwide emergence of plasmid-mediated quinolone resistance. Lancet Infect. Dis. 6(10), 629–640. Roy, S., Chatterjee, S., Bhattacharjee, A., Chattopadhyay, P., Saha, B., Dutta, S. and Basu, S. 2021. Overexpression of efflux pumps, mutations in the gyrA and parC genes, and their correlation with fluoroquinolone resistance in Acinetobacter baumannii. Front. Microbiol. 12, 602724. Saud, B., Paudel, G., Khichaju, S., Bajracharya, D., Dhungana, G., Awasthi, M.S. and Shrestha, V. 2019. Multidrug-resistant bacteria from raw meat of buffalo and chicken, Nepal. Vet. Med. Int. 2019, 1–7. Scallan, E., Hoekstra, R.M., Angulo, F.J., Tauxe, R.V., Widdowson, M.A., Roy, S.L., Jones, J.L. and Griffin, P.M. 2015. Foodborne illness acquired in the United States—major pathogens. Emerg. Infect. Dis. 21(1), 119–128. Scherrer, J.V. and Marcon, L.N. 2016. Biofilm formation and food safety in food service establishments. RASBRAN. Rev. Assoc. Bras. Nutr. 7(2), 91–99. Sekoai, P. T., Feng, S., Zhou, W., Ngan, W. Y., Pu, Y., Yao, Y., Pan, J. and Habimana, O. 2020. Insights into the microbiological safety of wooden cutting boards used for meat processing in Hong Kong’s wet markets. Microorganisms 8(4), 579. Sharma, S., Chakraborty, R., Kumar, R. and Singh, N. 2024. Knowledge and practices related to antibiotics among poultry producers in South Asia: a One Health perspective. One. Health. 18, 100639. Singh, R., Kumar, A e. and Devi, S. 2025. Drivers of antimicrobial resistance in livestock and poultry production. Front. Vet. Sci. 12, 200–214. Singh, S., Kriti, M., K.S, A., Sharma, P., Pal, N., Sarma, D.K., Tiwari, R. and Kumar, M. 2025. A One Health approach addressing poultry-associated antimicrobial resistance: human, animal and environmental perspectives. Microbe 7, 100309. Stepanović, S., Vuković, D., Hola, V., Bonaventura, G.D., Djukić, S., Ćirković, I. and Ruzicka, F. 2007. Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. Apmis 115(8), 891–899. Stepanović, S., Vukovic, D., Dakic, I., Savic, B. and Svabic-Vlahovic, M. 2000. A modified microtiterplate test for quantification of staphylococcal biofilm formation. J. Microbiol. Methods. 40(2), 175–179. Thomrongsuwannakij, T., Blackall, P. J.; Chansiripornchai, N., Punyapornwithaya, V. and Singer, R. S. 2020. A comparison of virulence genes, antimicrobial resistance and molecular typing of avian pathogenic Escherichia coli (APEC) from Australia and Thailand. Avian Pathol. 49(5), 434–445. World Health Organization. 2020. Estimating the burden of foodborne diseases: a global perspective for public health action. Geneva, Switzerland: WHO. Yu, H., Zhang, L. and Li, Y. 2025. Multidrug-resistant bacteria in alternative poultry production systems. Int. J. Food Microbiol. 410(1), 1–10. Yu, H., Zhao, X., Wang, Y., Li, L. and Zhang, Q. 2025. Phylogenetic relationship, genomic heterogeneity, and population structure of Escherichia coli isolated from retail poultry meat in China. Food Res. Int. 178, 113305. Zhang, P., Shen, Z., Zhang, C., Song, L., Wang, B., Shang, J., Yue, X., Qu, Z., Li, X., Wu, L., Zheng, Y., Aditya, A., Wang, Y., Xu, S. and Wu, C. 2017. Surveillance of antimicrobial resistance among Escherichia coli from chicken and swine, China, 2008–2015. Vet. Microbiol. 203, 49–55. | ||
| How to Cite this Article |
| Pubmed Style Martins CMDCR, Sá DRCD, Martins MG, Sousa MASD, Bezerra ADDS, Souza MRD, Araújo BN, Rosa DS, Gouveia GV, Costa MMD, Rodrigues RTDS. Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 Web Style Martins CMDCR, Sá DRCD, Martins MG, Sousa MASD, Bezerra ADDS, Souza MRD, Araújo BN, Rosa DS, Gouveia GV, Costa MMD, Rodrigues RTDS. Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. https://www.openveterinaryjournal.com/?mno=275581 [Access: February 04, 2026]. doi:10.5455/OVJ.2026.v16.i1.57 AMA (American Medical Association) Style Martins CMDCR, Sá DRCD, Martins MG, Sousa MASD, Bezerra ADDS, Souza MRD, Araújo BN, Rosa DS, Gouveia GV, Costa MMD, Rodrigues RTDS. Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 Vancouver/ICMJE Style Martins CMDCR, Sá DRCD, Martins MG, Sousa MASD, Bezerra ADDS, Souza MRD, Araújo BN, Rosa DS, Gouveia GV, Costa MMD, Rodrigues RTDS. Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 Harvard Style Martins, C. M. D. C. R., Sá, . D. R. C. D., Martins, . M. G., Sousa, . M. A. S. D., Bezerra, . A. D. D. S., Souza, . M. R. D., Araújo, . B. N., Rosa, . D. S., Gouveia, . G. V., Costa, . M. M. D. & Rodrigues, . R. T. D. S. (2026) Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 Turabian Style Martins, Carla Maria Do Carmo Resende, Daniele Rayssa Cavalcanti De Sá, Marlos Gomes Martins, Maria Antônia Sampaio De Sousa, Amanda Daiane Da Silva Bezerra, Matheus Rodrigues De Souza, Beatriz Nascimento Araújo, Danillo Sales Rosa, Gisele Veneroni Gouveia, Mateus Matiuzzi Da Costa, and Rafael Torres De Souza Rodrigues. 2026. Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 Chicago Style Martins, Carla Maria Do Carmo Resende, Daniele Rayssa Cavalcanti De Sá, Marlos Gomes Martins, Maria Antônia Sampaio De Sousa, Amanda Daiane Da Silva Bezerra, Matheus Rodrigues De Souza, Beatriz Nascimento Araújo, Danillo Sales Rosa, Gisele Veneroni Gouveia, Mateus Matiuzzi Da Costa, and Rafael Torres De Souza Rodrigues. "Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil." doi:10.5455/OVJ.2026.v16.i1.57 MLA (The Modern Language Association) Style Martins, Carla Maria Do Carmo Resende, Daniele Rayssa Cavalcanti De Sá, Marlos Gomes Martins, Maria Antônia Sampaio De Sousa, Amanda Daiane Da Silva Bezerra, Matheus Rodrigues De Souza, Beatriz Nascimento Araújo, Danillo Sales Rosa, Gisele Veneroni Gouveia, Mateus Matiuzzi Da Costa, and Rafael Torres De Souza Rodrigues. "Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil." doi:10.5455/OVJ.2026.v16.i1.57 APA (American Psychological Association) Style Martins, C. M. D. C. R., Sá, . D. R. C. D., Martins, . M. G., Sousa, . M. A. S. D., Bezerra, . A. D. D. S., Souza, . M. R. D., Araújo, . B. N., Rosa, . D. S., Gouveia, . G. V., Costa, . M. M. D. & Rodrigues, . R. T. D. S. (2026) Multidrug resistance and biofilm-forming ability of Escherichia coli isolated from free-range chicken meat in restaurants in Petrolina, Pernambuco, Brazil. doi:10.5455/OVJ.2026.v16.i1.57 |