| Research Article | ||
Open Vet. J.. 2026; 16(1): 275-286 Open Veterinary Journal, (2026), Vol. 16(1): 275-286 Research Article Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencingMuhammad Shafique1, Waqas Ali1*, Syed Mohsin Bukhari1, and Shahid Mehmood21Department of Wildlife and Ecology, University of Veterinary and Animal Sciences, Lahore, Pakistan 2Department of Poultry Production, University of Veterinary and Animal Sciences, Lahore, Pakistan *Corresponding Author: Waqas Ali. Department of Wildlife and Ecology, University of Veterinary and Animal Sciences, Lahore, Pakistan. Email: waqas.ali [at] uvas.edu.pk Submitted: 23/08/2025 Revised: 25/11/2025 Accepted: 08/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
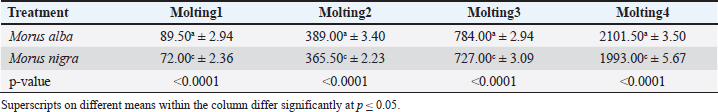
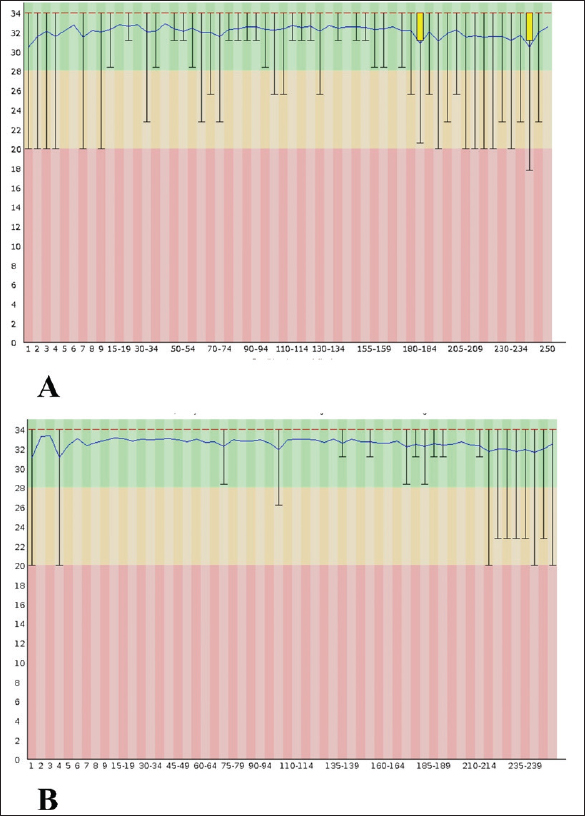
AbstractBackground: Bombyx mori is a key species in the sericulture industry and depends on mulberry leaves for growth and silk production. Aim: This study characterized the gut microbiota of B. mori fed on two mulberry species using 16S rRNA amplicon sequencing. Methods: Ten healthy fifth instar larvae were randomly selected from each treatment group. The larvae were dissected under sterile conditions to extract the gut tissues to obtain a pooled sample. DNA was extracted using a QIAamp DNA Microbiome kit, and the primer set 27F (5-AGAGTTTGATCCTGGCTCAG-3) and 1492-R (5-GGTTACCTTGTTACGACTT-3) was used to amplify variable regions V3–V4 of the 16S rRNA gene using Illumina sequencing. Using the EzBioCloud platform, Operational Taxonomic Unit data were used to estimate the bacterial species richness and evenness. The microbial communities’ phylum to genus level were grouped. Raw paired-end reads (FASTQ format) were imported into Qiime2 v2021.4 software, and a Krona plot was constructed. Results: The gut microbiota of B. mori larvae fed on Morus alba was predominantly composed of Proteobacteria (36.46%), Bacteroidota (22.98%), Firmicutes (20.31%), and Actinobacteriota (20.15%). All of these play critical roles in nitrogen fixation, carbohydrate breakdown, and immune modulation, thereby supporting optimal gut health and nutrient absorption. In contrast, silkworms fed on Morus nigra exhibited a microbiota dominated by Proteobacteria (31.74%), Verrucomicrobiota (23.35%), Bacteroidota (12.58%), and Bdellovibrionota (12.30%). Bdellovibrionota is a predatory bacterium that may disrupt the balance and potentially affect silkworm health and growth. Proximate analysis further supported the differences in microbiota composition of M. alba and M. nigra. M. alba had significantly higher levels of crude protein (26.16%) and crude fat (4.53%) than M. nigra (19.50% and 3.20%, respectively). Silkworms fed with M. alba exhibited superior growth across all molting stages compared with those fed with M. nigra. Conclusion: Future studies should focus on the effects of probiotic supplementation on microbial diversity, growth performance, and cocoon production. Increasing the sample size would provide more comprehensive results and individual-level variations in gut microbiota. These insights will contribute to the optimization of sericulture strategies for silkworm growth and silk yield. Keywords: Silkworms, NGS, Gut microbiota, Proteobacteria, Firmicutes. IntroductionSilkworms (Bombyx mori) have been vital to the textile industry for more than a thousand years. Silkworms are primarily used for silk production, which is a highly desired material due to its strength, luster, and special characteristics. Sericulture, the cultivation of silkworms for silk production, is believed to have originated in ancient China (Cappellozza et al., 2011). The domesticated silk moth (Bombyx mori) is one of the most specialized insects, subjected to rigorous artificial selection to increase silk yield. Silkworms are predominantly herbivorous and have a natural diet of mulberry leaves (Chen et al., 2018). Silkworms require mulberry leaves that are extremely enriched with vital ingredients, including proteins, carbohydrates, fats, and vitamins, to provide sufficient nutrition for their growth and development (Anand et al., 2010). Over the past few years, sericulture has developed as a cottage industry that provides support to the textile industry (Chen et al., 2020). However, the increasing demand for silk and silk products poses a significant challenge to sericulture (Dong et al., 2018). Many mulberry species and artificial diets have been used in sericulture to increase silk production efficiency. Morus alba leaves contain essential amino acids, carbohydrates, vitamins, and minerals (Shu et al., 2019). The use of M. alba leads to improved growth, metabolic functions, and high silk production (Feng et al., 2011, 2013; Gudda et al., 2020). Black mulberry (Morus nigra) is frequently used as an alternative to M. alba and has unique nutritional benefits (Fometu et al., 2022). Irrespective of these differences, little research has been conducted on the effects of both species on silkworm development and gut microbiota (Msango Soko et al., 2020; Heng et al., 2022). These variations can alter the microbial populations in the silkworm gut, potentially affecting digestion, growth rates, and silk production. Nonetheless, the specific effects of M. alba and M. nigra on the gut microbiota of silkworms remain unclear (Li et al., 2015). The gut microbiota is a diverse community of organisms, such as bacteria, fungi, and viruses, that inhabit the gastrointestinal tract of an organism. The gut microbiota plays a vital role in silkworm digestion, immune modulation, and nutrient absorption by directly affecting silkworm growth and silk production. Over the past few years, several studies have increased the evidence for the gut microbiota’s essential functions in food digestion, essential nutrient production, and immune system modulation (Li et al., 2022; Wei et al., 2024). Although research has shown that diet, particularly the type of mulberry leaves, influences the gut microbiota, the effects of M. nigra and M. alba on this microbiome remain unclear. A balanced gut microbiotais essential for a better growth rate, development, and production of high-quality silk (Li et al., 2020). Recent studies have reported that the composition and diversity of gut microbiota can be influenced by a number of factors, such as mulberry species, host physiological status, and environmental factors. Mulberry leaves provide more stable and high-quality nutrients to provide a balanced gut microbiota (Liang et al., 2014). Although the interaction between diet and gut microbiota in silkworms has been known for a long time, only recently have meta-genomic tools been used to properly reveal the composition and functionality of these microbial communities (Li et al., 2023). Metagenomics is the large-scale sequencing of genetic material directly from environmental samples that enables the identification and characterization of all possible microbes residing within the gut and can reveal their function within the host. Nevertheless, the impact of such varied diets on the gut microbiota and health of silkworms is mostly unexplored (Ma et al., 2020). Therefore, the present study aimed to characterize the gut microbiota of silkworms reared on two different mulberry species, e.g., M. alba and M. nigra, through meta-genomics and explore how the consumption of these two mulberry species may be reflected in the gut microbiota and how it could potentially be extended to sericulture. In addition, the findings of this study will help identify candidate microbial modulation agents, e.g., probiotics, aimed at enhancing the health status and productivity of silkworms. Materials and MethodsExperimental designThe experiment was conducted at the Sericulture Unit, Department of Wildlife and Ecology, UVAS, Ravi Campus, under semi-controlled environmental conditions. Fresh mulberry leaves of two different species, M. alba and M. nigra, were collected from Changa Manga Forest and visually inspected for pests and disease. Any leaves showing signs of contamination were excluded from the study. To ensure uniformity, only fresh leaves were used, and they were washed to remove any dirt or debris before being fed to the silkworms. Rearing of the silkwormEach treatment group consisted of 500 silkworm larvae. The first treatment group was provided with M. alba, and the second treatment group received M. nigra. The amount of food each larva received was uniform to control the diet intake by each treatment. Each treatment group received the same leaf weight. The proximate analysis of mulberry leaves used during the experiment was performed according to standard protocols (Sree and Vijayalakshmi, 2018). Sample’s collectionTo explore the gut microbiota of Bombyx mori under different dietary treatments, a total of 10 healthy fifth instar larvae were randomly selected from each treatment group. The larvae were surface sterilized using 70% ethanol and dissected under sterile conditions to extract the gut tissues to make a pooled sample (Schloss et al., 2011). The samples were stored at −80°C until DNA extraction for downstream next-generation sequencing (NGS) analysis was done (Martinson et al., 2017). DNA extraction, quantification, and sequencingDNA was extracted from the gut samples using the QIAamp DNA Microbiome kit following the manufacturer’s instructions and quantified using the Nano-Drop One (Thermo Fisher Scientific). The quality of DNA was also checked through 1% agarose gel using a UV transilluminator and Gel Doc system. The primer set 27F (5-AGAGTTTGATCCTGGCTCAG-3) and 1492-R (5-GGTTACCTTGTTACGACTT-3) was used to amplify variable regions V3–V4 of the 16S rRNA gene using Illumina sequencing (Nishida et al., 2016). The sequencing depth was maintained at approximately 20,000–30,000 reads per sample to ensure sufficient bacterial diversity coverage, as recommended for accurate taxonomic classification in microbiome studies (Li et al., 2020). Fast QC was used to evaluate read quality, and sequences were trimmed and filtered to remove low-quality sequences using Trimmomatic (Fig. 1). Sequences were then imported into Qiime2 v2021.4 for taxonomic classification, diversity estimation, and statistical testing (Ajmal et al., 2025).
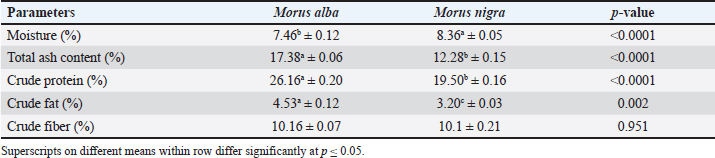
Fig. 1. Per-base sequence quality assessment of sequencing reads visualized using FastQC showing consistent high-quality scores (green) with some variability in the middle positions (yellow and red) A: Morus alba, B: Morus nigra. Clustering of Operational Taxonomic Units (OTUs) and species annotationThe PKSSU4.0 taxonomy databases of prokaryotic 16S rRNA gene sequences were used for preprocess analysis. Valid DNA reads were used for OTU picking (Qin et al., 2022). Estimation of the diversity indicesAlpha-diversity was estimated using metrics such as Shannon’s index and Simpson’s index to measure species richness and evenness within the microbial community. Diversity indices were estimated using the Bacterial and Viral Bioinformatics Resource Center at https://www.bv-brc.org (Ajmal et al., 2025; Hussain et al., 2025). Data analysisUsing OTU data, the microbial communities’ phylum to genus level were grouped. The subpopulations were compared using both ordination analysis and clustering. Raw paired-end reads (FASTQ format) from the original DNA fragments were imported into Qiime2 v2021.4 software, and a Krona plot was constructed (Pickard et al., 2017). The Sankey plot was constructed using the BV-BRC at https://www.bv-brc.org. The heatmap was constructed using Qiime2 v2021.4, with taxonomic assignment based on 16S rRNA gene sequencing data. The datawere presented as relative abundance for each genus. Silkworm growth data, including weight and length, were recorded for each treatment group. The obtained data were analysed through one-way ANOVA using PROC GLM in SAS software (version 9.1). For the comparison of significant treatment means, Fisher’s least significant difference test was applied (Steel et al., 1996), and the following model was applied: Yij=µ + τi + ϵij. Ethical approvalThe Ethics Review Committee (ERC), University of Veterinary and Animal Sciences, Lahore, Pakistan, approved this study vide letter No. DR/36. ResultsThe experiment was conducted under semi-controlled environmental conditions. During the experimental research period, the growth performance of B. mori was closely monitored to assess the effect of different mulberry species (M. alba and M. nigra) on growth performance. Proximate composition of the mulberry leavesThe proximate compositions of the different mulberry species are presented in Table 1. Significant differences (p ≤ 0.05) were observed in moisture, total ash, crude protein, and crude fat contents. Table 1. Proximate composition of M. alba and M. nigra (Mean ± SD, n=5)
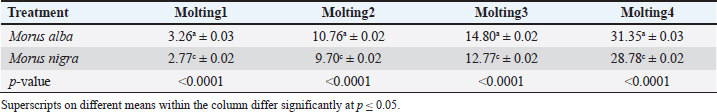
Growth performanceThe growth performance of B. mori larvae was assessed by measuring both body weight (BW) and body length at four successive molting stages under both dietary treatments. Randomly, 20 silkworm specimens from each treatment group were selected for morphometric measurements during each molting stage. Tables 2 and 3 present the statistical analysis of growth parameters among different treatment groups. Silkworms fed with M. alba leaves consistently exhibited the highest BW, reaching an average of 2101.50 ± 3.50 mg by the fourth molt. This was significantly higher than that in the M. nigra group (1993.00 ± 5.67 mg). Similarly, the body length increased significantly across the molting stages, with M. alba-fed larvae showing the maximum lengths at each stage. The silkworms fed with M. alba reached a mean length of 31.35 ± 0.03 mm at molting 4, which is significantly longer than those fed with M. nigra (28.78 ± 0.02 mm). Table 2. BW (mg) of silkworms at different molting stages during the present study (mean ± sd, n=20).
Table 3. Body length (mm) of silkworms at different molting stages during the present study (Mean ± SD, n=20).
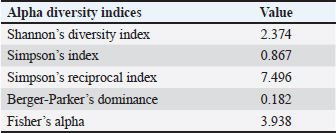
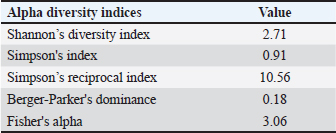
Alpha diversity of the gut microbiota of silkworms fed with M. albaSeveral alpha diversity indices were computed to assess the bacterial diversity within the gut microbiome of B. mori fed on M. alba (Table 4). The Shannon diversity index was 2.374, indicating moderate microbial diversity and an even distribution of species. This value indicated the presence of a community structure where no single species dominates overwhelmingly, supporting the coexistence of multiple functionally diverse taxa. Simpson’s diversity index, recorded at 0.867, further confirmed the sample’s moderate diversity. A value closer to 1 implies a balanced community with higher evenness, which complements the results of the Shannon index. Simpson’s reciprocal index was 7.496, indicating that the microbiome comprises several dominant taxa rather than a few highly abundant species. The Berger–Parker index was relatively low (0.182), indicating that the most abundant taxon did not disproportionately dominate the community, thereby reflecting taxonomic richness and uniformity. In addition, Fisher’s alpha index, a metric of species richness, was calculated to be 3.938, supporting the conclusion that the M. alba diet sustains a moderately diverse and complex gut microbiota in B. mori larvae. Collectively, these alpha diversity results indicate that a mulberry leaf-based diet maintains a stable gut environment with functional redundancy and taxonomic diversity, which may be advantageous for host digestion and immunity. Table 4. Alpha diversity indices of gut microbiota in Bombyx mori fed on M. alba
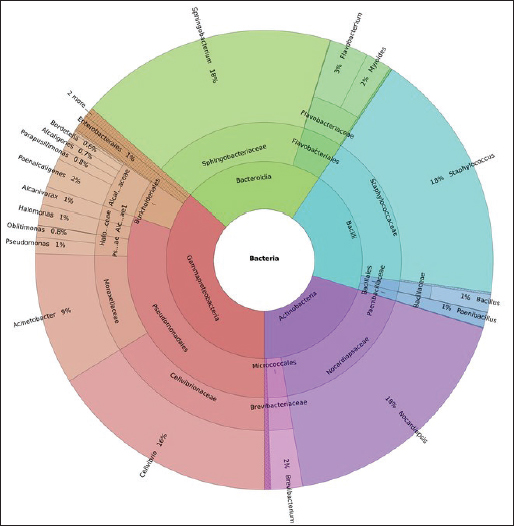
Gut microbiota composition of Bombyx mori fed with M. albaKRONA plot based on the relative abundance of different bacterial groups is shown in Figure 2. The gut microbiota of B. mori larvae reared on M. alba was dominated by four major bacterial phyla, including Proteobacteria (36.46%), Bacteroidota (22.98%), Firmicutes (20.31%), and Actinobacteriota (20.15%). The predominance of Proteobacteria is consistent with the findings of previous studies on lepidopteran gut microbiomes, highlighting its ecological versatility and role in nutrient acquisition. At the class level, the most abundant taxa were Gammaproteobacteria (36.45%) and Bacteroidia (22.97%), followed by Bacilli (20.36%) and Actinobacteria (20.22%). These classes include functionally diverse microbes involved in fermentation, nitrogen fixation, and immune modulation. Order-level classification revealed that Pseudomonadales (30.27%) and Sphingobacteriales (18.31%) were dominant, with notable contributions from Staphylococcales (18.05%), Streptosporangiales (17.58%), and Flavobacteriales (4.75%). These orders are often linked to complex plant-based carbohydrate degradation and microbial interactions important for host physiology. At the family level, Sphingobacteriaceae (19.514%) and Staphylococcaceae (19.159%) were the most abundant, followed by Nocardiopsaceae (18.627%), Cellvibrionaceae (17.244%), Moraxellaceae (9.917%), and Flavobacteriaceae (4.879%). Other families, such as Alcaligenaceae (4.648%), Brevibacteriaceae (2.395%), and Pseudomonadaceae (2.111%), were present at lower abundance, with Halomonadaceae (1.506%) contributing minimally. Genus-level analysis showed a relatively even distribution, with Sphingobacterium (20.33%), Staphylococcus (19.96%), and Nocardiopsis (19.59%) followed by Cellvibrio (18.13%) and Acinetobacter (10.43%). Flavobacterium (3.06%), Brevibacterium (2.52%), Paenalcaligenes (2.33%), Myroides (2.07%), and Halomonas (1.58%) also contributed to the microbial community. These genera are essential for cellulose degradation, antimicrobial compound production, and nitrogen fixation.
Fig. 2. KRONA plot illustrating the hierarchical taxonomic composition of the gut microbiota in Bombyx mori larvae reared on M. alba. Functional role of dominant bacterial groups in the gut of M. alba-fed silkwormsThe phylum proteobacteria, particularly gammaproteobacteria, was most abundant in the gut of silkworms fed with M. alba. These bacteria play critical roles in the fermentation of plant polysaccharides and in the breakdown of complex carbohydrates from mulberry leaves into simple sugars. In addition, they contribute to nitrogen fixation and the production of bioactive compounds, such as antimicrobial peptides, that help maintain a balanced gut environment. Bacteria belonging to the genus Sphingobacterium help to degrade cellulose and plant polysaccharides. In addition, these bacteria break down plant fiber and release simple sugars for energy. They play a critical role in fermentation, which can also contribute to gut health by regulating intestinal pH. The members of the order Bacilli and Staphylococcales belong to the phylum Firmicutes and help in the fermentation of undigested carbohydrates. They convert them into SCFAs, including acetate and butyrate, which in turn support gut health and reduce inflammation. Furthermore, these bacteria are involved in the synthesis of essential vitamins required for metabolism. The genus Nocardiopsis belongs to the phylum Actinobacteriota and plays a critical role in the breakdown of complex organic compounds, such as plant proteins, thus helping the silkworm to digest its diet efficiently. These bacteria also produce enzymes that help breakdown plant cell walls and aid in nutrient absorption. Alpha diversity of the gut microbiota of silkworms fed with M. nigraTable 5 presents the alpha diversity indices of the gut microbiota in Bombyx mori fed on Morus nigra. Shannon’s diversity index was calculated as 2.374, indicating a moderate level of bacterial diversity. Simpson’s index (0.867) indicated that the silkworm gut bacterial community is relatively diverse. Simpson’s reciprocal index (7.496) indicated higher diversity. The Berger––Parker’s dominance index value (0.182) revealed that the bacterial community is not overly dominated by a single group of bacteria. Finally, Fisher’s alpha (3.938) value indicated a moderate level of bacterial richness in the gut microbiota of silkworms fed on Morus nigra. Collectively, diversity indices values indicated a well-balanced bacterial community in the silkworm gut. Table 5. Alpha diversity indices of gut microbiota in Bombyx mori fed on M. nigra.
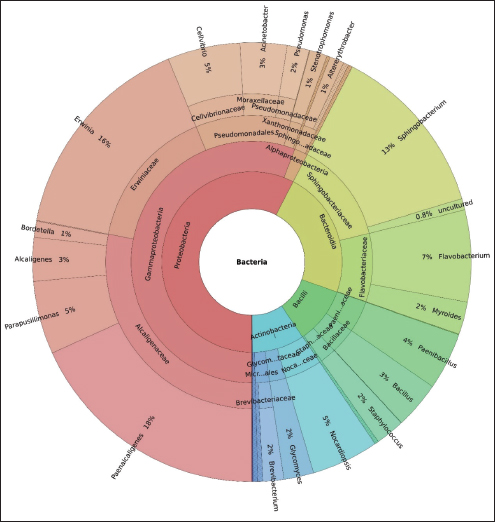
Gut microbiota composition of Bombyx mori fed with M. nigraFigure 3 shows the KRONA plot based on the relative abundance of different bacterial groups. The gut microbiota of Bombyx mori larvae reared on Morus nigra was dominated by seven major bacterial phyla, including Proteobacteria (31.74%), Verrucomicrobiota (23.35%), Bacteroidota (12.58%), Bdellovibrionota (12.30%), Fusobacteriota (9.23%), Firmicutes (5.59%), and Actinobacteriota (5.20%). At the class level, the most abundant taxa were as follows: Gammaproteobacteria (30.76%), Verrucomicrobiae (23.35%), Bacteroidia (12.58%), Bdellovibrionia (12.30%), Fusobacteriia (9.23%), Bacilli (5.59%), Actinobacteria (5.20%), and Alphaproteobacteria (0.98%). Burkholderiales (29.90%) was the most abundant order, followed by Enterobacterales (16.70%), Sphingobacteriales (14.47%), Pseudomonadales (11.15%), Flavobacteriales (9.72%), Streptosporangiales (5.25%), Paenibacillales (4.50%), Bacillales (3.69%), Glycomycetales (2.34%), and Staphylococcales (2.28%). Bacterial families, including Alcaligenaceae (31.17%), Erwiniaceae (17.41%), Sphingobacteriaceae (15.08%), Flavobacteriaceae (10.14%), Cellvibrionaceae (5.93%), Nocardiopsaceae (5.48%), Paenibacillaceae (4.69%), Bacillaceae (3.85%), Moraxellaceae (3.82%), and Glycomycetaceae (2.44%) were identified. The genus-level identification revealed that Paenalcaligenes (22.67%) was the most dominant, followed by Erwinia (19.58%), Sphingobacterium (15.91%), Flavobacterium (8.43%), Parapusillimonas (6.73%), Cellvibrio (6.66%), Nocardiopsis (6.16%), Paenibacillus (5.27%), and Bacillus (4.29%). The genus Acinetobacter (4.29%) was the least abundant. The most abundant phyla, including Proteobacteria and Gammaproteobacteria, and key genera, such as Paenalcaligenes and Erwinia, indicated a complex bacterial community that plays important roles in nutrient degradation and host-microbe interactions.
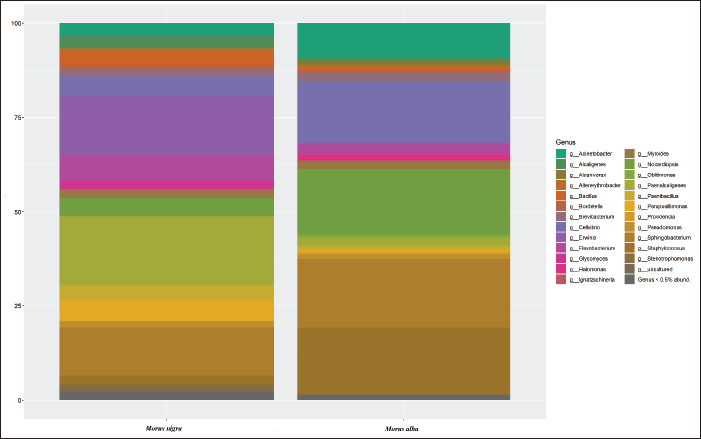
Fig. 3. KRONA plot illustrating the hierarchical taxonomic composition of the gut microbiota in Bombyx mori larvae reared on Morus nigra. Functional role of dominant bacterial groups in the gut of M. nigra-fed silkwormsThe significant presence of the phylum Verrucomicrobiota suggested their role in mucosal health and gut barrier function. These bacteria are mostly involved in the breakdown of complex polysaccharides and may help regulate the gut immune system. They also produce SCFAs, which help maintain an optimal gut pH. The role of Bdellovibrionota bacteria is still under investigation, and they are believed to be predatory and parasitize other bacteria. The presence of these bacteria indicates that there is a natural mechanism of regulating bacterial populations in the gut of silkworms by maintaining a balance between beneficial and potentially harmful bacteria. Members of the phylum Fusobacteriota are mostly involved in protein degradation and are thought to contribute to nitrogen metabolism in the silkworm gut. They carried out fermentation and produced products that support the growth and development of silkworms. Variations at the genus levelThe analysis of the gut microbiota in B. mori fed on M. alba and M. nigra revealed distinct variations in the microbial composition at the genus level, indicating that different mulberry species influence the gut microbiome (fig. 4). The gut microbiota composition of B. mori fed with M. alba was dominated by Proteobacteria, Bacteroidota, Firmicutes, and Actinobacteriota. These bacteria are essential for nitrogen fixation, carbon breakdown, and immune modulation. In contrast, Proteobacteria, Verrucomicrobiota, Bacteroidota, and Bdellovibrionota dominated the microbiota of silkworms fed on M. nigra, making it more complex. The identification of Bdellovibrionota, a predatory bacterium, indicates a more complex microbial association, which may have implications for the health of silkworms.
Fig. 4. Heatmap showing the relative abundance of bacterial genera in the gut microbiota of Bombyx mori larvae reared on M. alba and M. nigra. The rows and columns represent bacterial genera and treatment groups, respectively. Color intensity indicates the relative abundance of each genus, with darker colors corresponding to higher abundance Silkworms fed on M. albaThe most prominent genera include Sphingobacterium (20.33%), which is responsible for cellulose degradation and plant polysaccharide fermentation, thus playing a critical role in plant fiber breakdown into simpler sugars. The relative abundance of Staphylococcus was 19.96%, followed by Nocardiopsis (19.59%). These genera aid in breaking complex organic substances, such as plant proteins, and facilitate digestion and nutrient absorption. Cellvibrio (18.13) and Acinetobacter (10.43) were also dominant genera. Both these genera play a role in gut pH regulation. In general, the microbial composition exhibited by silkworms fed on M. alba is balanced, thereby supporting silkworm growth, digestion, and silk production. Silkworms fed on M. nigraConversely, the genus-level microbiota composition showed a much more complex and less diverse community of microbes when silkworms fed on M. nigra. The relative abundance of Paenalcaligenes was 22.67%, and these bacteria contribute to nutrient degradation and host-microbe associations. Other dominant groups, Erwinia (19.58%) and Sphingobacterium (15.91), were also present but appear to play a different role in this context than they do in the M. alba group. The remaining genera, such as Flavobacterium (8.43) and Cellvibrio (6.66), were also detected, and these bacteria were found to be involved in the breakdown of polysaccharides. The presence of Bdellovibrionota (12.30%) and Fusobacteriota (9.23%) indicated more complex interactions, including potential predatory behavior that might influence the gut ecosystem by regulating the populations of other bacteria. These variations could be attributed to the higher concentrations of secondary metabolites in M. nigra, which may inhibit the growth of certain bacterial species, thus altering the gut microbiota balance and possibly affecting the digestion and health of silkworms. DiscussionThe gut microbiota of B. mori is one of the key factors affecting growth and development. In the present study, the gut microbiota of silkworms was characterized using 16S rRNA amplicon sequencing. Silkworms were fed with M. alba and M. nigra to determine the variation in the gut microbiota and how this affected cocoon growth and production. The larvae were surface sterilized with 70% ethanol and dissected under sterile conditions to extract the gut tissues to make a pooled sample. Pooling is a common practice in microbiome research, especially when dealing with model organisms, such as insects. Several studies have successfully used pooled samples to assess the gut microbiota, including studies on silkworms. The pooled silkworm gut samples have been used to obtain sufficient microbial DNA for downstream sequencing (Li et al., 2020). The pooling of multiple individuals has been found to be a good representation of the microbial composition and is often said to be an applicable method when the aim is to compare microbial compositions with different treatment groups rather than focusing on variation between individuals (Zhang et al., 2022). However, pooling does reduce the potential to detect biological variation within groups, which is potentially important in studies that aim to determine the impact of diet on microbial diversity at the individual level. In addition to pooled samples, individual replicates could be included in future studies to obtain both population-level information and individual variability analysis (Schloss et al., 2011). The microbial diversity found in the silkworm gut is probably affected by several factors, such as the diet’s chemical makeup. The gut microbiota of B. mori is altered by different aspects of diet, particularly the mulberry species provided as food. During the present study, it was noticed that M. alba and M. nigra have different effects on the microbiota of the silkworm gut. Silkworms fed with M. alba exhibited a diverse gut microbiota consisting of Proteobacteria, Firmicutes, and Actinobacteriota. These bacterial phyla are important for carbohydrate degradation, nitrogen fixation, and immune modulation, all of which are crucial for the efficient digestion and absorption of nutrients and overall silkworm health (Li et al., 2020). Stable gut microbiota promotes metabolic processes and immune defense mechanisms for optimal growth and silk yield. The microbial community of silkworms fed with M. nigra was more complex and changed to Proteobacteria, Verrucomicrobiota, and Bdellovibrionota, and the overall community richness was lower than that in the M. alba group. However, some of these bacteria, such as those from the Bdellovibrionota group, have been reported to contribute to mucosal health and immunoregulation. Morus nigra has a higher concentration of secondary metabolites, such as flavonoids, alkaloids, and phenolic compounds, which possess antimicrobial properties (Shu et al., 2023). The levels of secondary metabolites in M. nigra may provide an unfavorable environment for some beneficial microbes associated with digestion and nutrient absorption (Shu et al., 2023). This disruption in gut microbiota could have negative implications for silk production.These substances may make living conditions less favorable for some microbial species and thus have an impact on gut flora composition (Wang et al., 2017). Secondary metabolites may have an inhibitory effect on the growth of specific bacteria, leading to an imbalance in the gut microbiota. Moreover, high concentrations of polyphenolic compounds in M. nigra may disturb the microbial communities that help in the digestion of complicated carbohydrates, which may consequently affect silkworm development and health. Similar results have been found in other herbivorous insects, as dietary secondary metabolites alter the gut microbiota, but often in a negative way in terms of reducing microbial diversity and decreasing growth (Li et al., 2022).). Similarly, Zhang et al. (2022) reported that the nutritional and chemical composition of mulberry leaves can affect silkworm gut microbial diversity and function. Diet composition has an important impact on the gut microbiota of B. mori, which can affect its growth, development, and silk production. Nutritional quality plays an important role in shaping the composition and diversity of gut microbes, which affects silkworms’ metabolic efficiency (Li et al., 2022). As reported by Yuan et al. (2023), silkworms raised on artificial or low-quality diets display depleted gut microbiota, correlating with slow growth and low performance. Li et al. (2022) and Xu et al. (2022) reported that silkworms fed on M. alba have better growth and development. This is because M. alba provides better gut conditions for bacteria, which in turn helps in nutrient absorption and growth. In contrast, Xu et al. (2015) documented that silkworms raised on M. nigra showed overall lower diversity of gut microbiota, mostly dominated by Firmicutes. These findings agree with our results, as we found a similar pattern of diversity in the microbiota. One possible reason is that M. nigra leaves have higher concentrations of secondary metabolites than M. alba leaves, which potentially inhibit the growth of specific bacteria. However, M. alba fed on the silkworm’s gut maintained the relative abundance of those bacteria that allowed for better growth. Wang et al. (2024) reported similar findings. A healthy and varied gut microbiotais important for the digestion and absorption of nutrients and the health of silkworms. Diverse microbiota with a predominance of Proteobacteria and Firmicutes were found in the gut of silkworms fed on M. alba. These bacteria are involved in nutrient breakdown, carbohydrate fermentation, and nitrogen fixation. They are important for the digestion and immune health of silkworms, leading to optimal growth and silk production (Li et al., 2020). In contrast, silkworms fed on M. nigra exhibited a more complex microbiota, including Bdellovibrionota and Verrucomicrobiota. These bacteria might play a role in immune modulation and mucosal health but might also disturb the gut microbiota balance (Zhang et al., 2022). Morus alba leaves provide a more balanced diet, and they are low in toxicity, which in turn promotes bacteria that aid in the digestion and metabolism of nutrients. However, when M. nigra is rich in secondary metabolites, it can have a negative impact on silkworms due to inefficient digestion (Wang et al., 2017). Hence, these compounds may act as inhibitors of some bacteria, as reported in the current study, which showed decreased levels of Proteobacteria and Bacteroidetes in the gut microbiota of silk moths fed on M. nigra. The effect of secondary metabolites in plants on gut microbiota has been widely examined in many other herbivorous insects. Wang et al. (2017) and Tao et al. (2022) reported that a few polyphenolic compounds present in the leaves can control the growth of gut symbiotic bacteria, thereby influencing the overall fitness of the insect. Proteobacteria in M. alba-fed silkworms are efficient in degrading plant secondary metabolites. The reduction of toxicity by Proteobacteria can enhance the growth and silk production of silkworms fed with M. alba (Shu et al., 2023). Similar results were noted during the present study. However, studies have found that the gut microbiota of B. mori is essential for the detoxification of plant-derived toxins via biotransformation processes such as glucosylation and hydrolysis (Rajan et al., 2020). In the present study, when reared on M. nigra, a mulberry type associated with increased toxins, silkworms expressed significantly higher microbial activity related to detoxification of purportedly harmful secondary metabolites. Li et al. (2022) and Xu et al. (2022) reported that the detoxification capacity of the gut microbiota can be improved using probiotics to reduce the adverse effects of M. nigra. The role of the gut microbiota in detoxification is not limited to B. mori. The detoxification process has been reported for many herbivorous insects, such as D. saccharalis (Li et al., 2023). Silkworm growth is a dynamic process that depends on the absorption rate of nutrients, the availability of essential nutrients, and the metabolic capability of the gut microbiota. Such results were also consistently observed in other studies as M. alba-fed silkworms grew faster and developed more robustly than M. nigra-fed silkworms (Dong et al., 2018; Shu et al., 2023; Yuan et al., 2023). Although the exact mechanisms by which the gut microbiota alters silk production remain elusive, they may include multiple processes, such as nutrient metabolism, immune modulation, and detoxification. As discussed previously, microbial-specific groups such as Lactobacillus and Weissella primarily perform a fundamental function in the digestion of feed that has been formulated into a protein- or carbohydrate-rich diet to release nutrients available for animal absorption (Yuan et al., 2023). A rich gut microbiota is required to degrade polysaccharides, fats, and proteins into nutrient-utilizable components. Morus alba is nutritious and contains the required proteins, carbohydrates, vitamins, and minerals to facilitate a broad and healthy microbiota. A diet containing these ingredients will promote the growth of beneficial microbial species, including but not limited to Lactobacillus, Weissella, and Rhodococcus, which can improve digestion, increase nutrient absorption, and improve overall health (Liang et al., 2014). Morus nigra is a very closely related species, but its secondary metabolites, such as flavonoids, alkaloids, and other phenolic compounds, are significantly higher (Shu et al., 2023). Such compounds prevent many bacteria from growing, leading to a depletion of diversity in gut microbiota, making digestion ineffective (Li et al., 2022). Probiotics can be used to enhance growth and silk production, as highlighted by Shu et al. (2023.). Bacillus subtilis probiotics can be supplemented into the diet to promote better microbial diversity, nutrient assimilation, and immune function efficacy. Silkworms fed on M. nigra show reduced microbial diversity; therefore, supplementation with probiotics could restore gut microbial flora homeostasis to increase digestion and silk production. Several limitations must be considered when interpreting this study’s findings. First, the small sample size of 10 larvae per treatment group limits the generalizability of the results. Although pooling samples allows for microbial diversity analysis at the group level, it reduces the ability to detect individual-level variability, which could influence the results. Increasing the sample size in future studies will help better capture the biological variation within the treatment groups. The lack of biological replication (i.e., pooling samples from larvae) is another limitation that affects the reliability of the findings. Future studies should consider using multiple biological replicates per treatment group to ensure that the results are more robust and allow for statistical comparisons between individual samples. ConclusionThis study characterized the gut microbiota of B. mori larvae fed on M. alba and M. nigra. Notable bacterial phyla, such as Proteobacteria, Bacteroidota, Firmicutes, and Actinobacteriota, have been reported in the gut of silkworms fed on M. alba. These bacterial phyla are involved in nitrogen fixation, immunity, carbohydrate breakdown, digestion, and nutrient absorption, thus providing optimal gut health. However, Proteobacteria, Verrucomicrobiota, Bacteroidota, and Bdellovibrionota dominated the gut microbiota of silkworms fed on M. nigra, indicating a more complex bacterial interaction. Bdellovibrionota is a predatory bacterium that helps in mucosal health and immune regulation. These bacteria are believed to disrupt the gut ecosystem balance, affecting silkworm health and growth. The nutritional analysis revealed significant differences between the two mulberry species, with M. alba containing higher levels of crude protein and fat, which likely supported the superior growth observed in larvae fed with M. alba. Future research should investigate the effects of probiotic supplementation, such as Lactobacillus or B. subtilis, on microbial diversity, growth performance, and cocoon production. Future studies should increase the sample size and incorporate biological replicates, which will provide more reliable results and allow for the detection of individual-level variations in microbiota composition. These insights could ultimately contribute to more effective sericulture management strategies by optimizing silkworm growth and silk yield. AcknowledgmentWe extend our heartfelt thanks to the Punjab Forestry, Wildlife, and Fisheries Department for providing us with silkworm seeds. Special thanks to the BS Zoology students for their assistance in collecting and processing mulberry leaves during the rearing period. Conflict of interestThe authors declare no conflicts of interest. Funding This study received no specific grant. Authors’ contributionsMuhammad Shafique: Methodology, Investigation, Writing-Original Draft, Waqas Ali: Conceptualization, Supervision, and Formal analysis, Syed Mohsin Bukhari: Writing – Review and Editing, and Sahid Mehmood: Formal analysis and Data Curation. Data availabilityData supporting the findings of this study are not openly available due to sensitivity reasons. However, they are available upon reasonable request from the corresponding author. ReferencesAjmal, R., Imran, F., Ayub, A., Bilal, A., Bilal, M., Ali, K. and Masood, M. 2025. Preliminary assessment and metagenomic analysis of oral microbiota of the small Indian mongoose Herpestes auropunctatus through next-generation sequencing. Mech. Biosyst. 16(1), e25010. Anand, A.A., Vennison, S.J., Sankar, S.G., Gilwax Prabhu, D.I., Vasan, P.T., Raghuraman, T., Jerome Geoffrey, C. and Vendan, S.E. 2010. Isolation and characterization of bacteria from the gut of Bombyx mori that degrade cellulose, xylan, pectin and starch and their impact on digestion. J. Biol. Chem. Sci. Eng. Insect Sci. 10, 107. Cappellozza, S., Saviane, A., Tettamanti, G., Squadrin, M., Vendramin, E., Paolucci, P., Franzetti, E. and Squartini, A. 2011. Identification of Enterococcus mundtii as a pathogenic agent involved in the “flacherie” disease in Bombyx mori L. larvae reared on artificial diet. J. Invertebr. Pathol. 106(3), 386–393. Chen, B., Du, K., Sun, C., Vimalanathan, A., Liang, X., Li, Y., Wang, B., Lu, X., Li, L. and Shao, Y. 2018. Gut bacterial and fungal communities of domesticated silkworm (Bombyx mori) and wild mulberry-feeding relatives. ISME. J. 12(9), 2252–2262. Chen, B., Zhang, N., Xie, S., Zhang, X., He, J., Muhammad, A., Sun, C., Lu, X. and Shao, Y. 2020. Gut bacteria of the silkworm Bombyx mori facilitate host resistance to the toxic effects of organophosphate insecticides. Environ. Int. 143, 105886. Dong, H.L., Zhang, S.X., Chen, Z.H., Tao, H., Li, X., Qiu, J.F., Cui, W.Z., Sima, Y.H., Cui, W.Z. and Xu, S.Q. 2018. Differences in gut microbiota between silkworms (Bombyx mori) reared on fresh mulberry (Morus alba var. multicaulis) leaves or an artificial diet. RSC. Adv. 8(46), 26188–26200. Feng, N., Guo, X., Liang, S., Zhu, Y. and Liu, J. 2011. Biosorption of heavy metals from aqueous solutions by chemically modified orange peel. J. Hazard. Mater. 185(1), 49–54. Feng, W., Wang, X.Q., Zhou, W., Liu, G.Y. and Wan, Y.J. 2013. Isolation and characterization of lipase-producing bacteria in the intestine of the silkworm, Bombyx mori, reared on different forage. J. Agricult. Agricult. Res. Insect Sci. 11, 135. Fometu, S.S., Ma, Q., Wang, J., Guo, J., Ma, L. and Wu, G. 2022. Biological effect evaluation of different sized titanium dioxide nanoparticles using Bombyx mori (silkworm) as a model animal. Trace Elem. Res. 200(12), 5260–5272. Gudda, F.O., Waigi, M.G., Odinga, E.S., Yang, B., Carter, L. and Gao, Y. 2020. Antibiotic-contaminated wastewater irrigated vegetables pose resistance selection risks to the gut microbiome. Environ. Pollut. 264, 114752. Heng, J., Liu, H., Xu, J., Huang, X., Sun, X., Yang, R., Xia, Q. and Zhao, P. 2022. KPI5 regulates the expression of antibacterial peptide genes and hemolymph melanization in the silkworm Bombyx mori. Front. Immunol. 13, 907427. Hussain, M., Ali, W., Meyer, C.F.J., Javid, A. and Imran, M. 2025. Characterization and metagenomics analysis of the oral microbiome of Pteropus medius: insights from next-generation sequencing. Mech. Biosyst. 16(1), 25016. Li, C., Xu, S., Xiang, C., Xu, S., Zhou, Q. and Zhang, J. 2022. The gut microbiota of silkworm are altered by antibiotic exposure. FEBS Open Biol. 12, 2203–2212. Li, F., Li, M., Mao, T., Wang, H., Chen, J., Lu, Z., Qu, J., Fang, Y., Gu, Z. and Li, B. 2020. Effects of phoxim exposure on gut microbial composition in the silkworm, Bombyx mori. Ecotoxicol. Environ. Saf. 189, 110011. Li, G.N., Xia, X.J., Parfait, S., Zhao, H.H., Long, Y.H. and Zhu, Y. 2015. Effects of fluoride on the intestinal microflora of Bombyx mori. Acta Microbiol. Sin. 55, 926–934. Li, J., Deng, J., Deng, X., Liu, L. and Zha, X. 2023. A metabonomic analysis of the silkworm midgut revealed differences between the physiological effects of an artificial and mulberry leaf diet. Insects 14(4), 347; doi: 10.3390/insects14040347 Liang, X., Fu, Y., Tong, L. and Liu, H. 2014. Microbial shifts of the silkworm larval gut in response to lettuce leaf feeding. Appl. Microbiol. Biotechnol. 98(8), 3769–3776. Ma, W., Zheng, X., Li, L., Shen, J., Li, W. and Gao, Y. 2020. Changes in the gut microbiota of honeybees associated with jujube flower disease. Ecotoxicol. Environ. Saf. 198, 110616. Martinson, V.G., Douglas, A.E. and Jaenike, J. 2017. Community structure of the gut microbiota in sympatric species of wild Drosophila. Ecol. Lett. 20(5), 629–639. Msango Soko, K., Gandotra, S., Chandel, R.K., Sharma, K., Ramakrishinan, B. and Subramanian, S. 2020. Composition and diversity of gut bacteria associated with the eri silk moth, Samia ricini, (Lepidoptera: saturniidae) as revealed by culture-dependent and metagenomics analysis. J. Microbiol. Biotechnol. 30(9), 1367–1378; doi:10.1016/j.jmbio.2013.09.010 Nishida, S., Ono, Y. and Sekimizu, K. 2016. Lactic acid bacteria activating innate immunity improve survival in a silkworm model of bacterial infection. Drug. Discov. Ther. 10, 49–56. Pickard, J.M., Zeng, M.Y., Caruso, R. and Núñez, G. 2017. Gut microbiota: role in pathogen colonization, immune responses, and inflammatory disease. Immunol. Rev. 279, 70–89. Qin, L., Qi, J., Shen, G., Qin, D., Wu, J., Song, Y., Cao, Y., Zhao, P. and Xia, Q. 2022. Effects of microbial transfer during food-gut-feces circulation on the health of Bombyx mori. Microbiol. Spectr. 10(6), 2357. Rajan, R., Chunduri, A.R., Lima, A. and Mamillapalli, A. 2020. 16S rRNA sequence data of Bombyx mori gut bacteriome after spermidine supplementation. BMC. Res. Notes. 13(1), 94. Schloss, P.D., Gevers, D. and Westcott, S.L. 2011. Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PLos One. 6(12), e27310. Shu, Q., Wang, Y., Gu, H., Zhu, Q., Liu, W., Dai, Y., Li, F. and Li, B. 2023. Effects of artificial diet breeding on the intestinal microbial populations of silkworm (Bombyx mori) at the young stage. Arch. Insect Biochem. Physiol. 113(3), e22019. Sree, L. and Vijayalakshmi. 2018. Proximate composition, nutritional evaluation and mineral analysis in the leaves of an indigenous medicinal plant Alternanthera sessilis. Int. J. Health Sc. 8, 55–62. Steel, R.G.D., Torrie, J.H. and Dinkey, D.A. 1996. Principles and procedures of statistics, 2nd ed. Singapore: McGraw Hill Book Co. Tao, S., Wang, J., Liu, M., Sun, F., Li, B. and Ye, C. 2022. Hemolymph metabolomic differences in silkworms (Bombyx mori L.) under mulberry leaf and two artificial diet rearing methods. Arch. Physiol. Biol. Insect. Biochem. Physiol. 109, e21851. Wang, D., Dong, Z., Zhang, Y., Guo, K., Guo, P., Zhao, P. and Xia, Q. 2017. Proteomics provides insight into the interaction between mulberry and silkworm. Proteome Res. 16, 2472–2480. Wang, G., Jiang, G., Peng, R., Wang, Y., Li, J., Sima, Y. and Xu, S. 2024. Multi-omics integrative analysis revealed characteristic changes in blood cell immunity and amino acid metabolism in a silkworm model of hyperproteinemia. Int. J. Biol. Macromol. 258, 128809; doi:10.1016/j.ijbm.2012.12.8809 Wei, J., Zhang, Y. and Xu, L. 2024. Uncovering the immune evasion mechanism of microbial pathogens: lessons from anti-encapsulation strategies adopted by a fungus. Integr. Zool. 19(6), 1229–1232; doi:10.1016/j.izz.2019.12.029 Xu, G., Sun, Z.L., Hu, X.L., Xue, R.Y., Cao, G.L. and Gong, C.L. 2015. Analyze the diversity of silkworm intestinal bacteria based on 16S rRNA gene sequence. Sci. Seric. 41, 641–649. Xu, K., Lan, H., He, C., Wei, Y., Lu, Q., Cai, K., Yu, D., Yin, X., Li, Y. and Lv, J. 2022. Toxicological effects of trace amounts of pyriproxyfen on the midgut of silkworms. Pestic. Biochem. Physiol. 188, 105266. Yuan, S., Sun, Y., Chang, W., Zhang, J., Sang, J., Zhao, J., Song, M., Qiao, Y., Zhang, C., Zhu, M., Tang, Y. and Lou, H. 2023. The silkworm (Bombyx mori) gut microbiota is involved in metabolic detoxification by glucosylation of plant toxins. Commun. Biol. 6(1), 790. Zhang, X., Zhang, F. and Lu, X. 2022. Diversity and functional roles of the gut microbiota in Lepidopteran insects. Microorganisms 10(6), 1234. | ||
| How to Cite this Article |
| Pubmed Style Shafique M, Ali W, Bukhari SM, Mehmood S. Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Vet. J.. 2026; 16(1): 275-286. doi:10.5455/OVJ.2026.v16.i1.25 Web Style Shafique M, Ali W, Bukhari SM, Mehmood S. Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. https://www.openveterinaryjournal.com/?mno=279251 [Access: January 31, 2026]. doi:10.5455/OVJ.2026.v16.i1.25 AMA (American Medical Association) Style Shafique M, Ali W, Bukhari SM, Mehmood S. Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Vet. J.. 2026; 16(1): 275-286. doi:10.5455/OVJ.2026.v16.i1.25 Vancouver/ICMJE Style Shafique M, Ali W, Bukhari SM, Mehmood S. Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Vet. J.. (2026), [cited January 31, 2026]; 16(1): 275-286. doi:10.5455/OVJ.2026.v16.i1.25 Harvard Style Shafique, M., Ali, . W., Bukhari, . S. M. & Mehmood, . S. (2026) Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Vet. J., 16 (1), 275-286. doi:10.5455/OVJ.2026.v16.i1.25 Turabian Style Shafique, Muhammad, Waqas Ali, Syed Mohsin Bukhari, and Shahid Mehmood. 2026. Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Veterinary Journal, 16 (1), 275-286. doi:10.5455/OVJ.2026.v16.i1.25 Chicago Style Shafique, Muhammad, Waqas Ali, Syed Mohsin Bukhari, and Shahid Mehmood. "Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing." Open Veterinary Journal 16 (2026), 275-286. doi:10.5455/OVJ.2026.v16.i1.25 MLA (The Modern Language Association) Style Shafique, Muhammad, Waqas Ali, Syed Mohsin Bukhari, and Shahid Mehmood. "Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing." Open Veterinary Journal 16.1 (2026), 275-286. Print. doi:10.5455/OVJ.2026.v16.i1.25 APA (American Psychological Association) Style Shafique, M., Ali, . W., Bukhari, . S. M. & Mehmood, . S. (2026) Comparative analysis of gut microbiota in Bombyx mori fed on M. alba and M. nigra using 16S rRNA amplicon sequencing. Open Veterinary Journal, 16 (1), 275-286. doi:10.5455/OVJ.2026.v16.i1.25 |